Abstract
Toxoplasmosis is a zoonotic disease caused by the protozoan parasite Toxoplasma gondii. Infection in humans has usually been related to the consumption of raw, undercooked or cured meat. The aim of this study was to develop a droplet digital polymerase chain reaction (ddPCR)-based assay for the detection and quantification of T. gondii in meat samples. To optimize the ddPCR, T.gondii reference DNA aliquots at five known concentrations: 8000 cg/µl, 800 cg/µl, 80 cg/µl, 8 cg/µl were used. Moreover, results obtained by ddPCR and quantitative PCR (qPCR) were compared using 80 known samples (40 positive and 40 negative), as well as 171 unknown diaphragm tissue samples collected at slaughterhouses. The ddPCR showed a sensitivity of 97.5% and a specificity of 100%, with a detection limit of 8 genomic copy/µl of T. gondii. A nearly perfect agreement (κ = 0.85) was found between results obtained by ddPCR and qPCR for both positive and negative known samples analysed. On the 171 diaphragm tissue samples from field, 7.6% resulted positive by ddPCR and only 1.2% by qPCR. Therefore, this innovative method could be very useful for the detection of T. gondii in meat samples, aiming to prevent human infections.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Toxoplasmosis, caused by the intracellular protozoan Toxoplasma gondii, is one of the most common parasitic infection in animals and humans worldwide (Almeira and Dubey, 2021). Domestic and wild felids are the definitive hosts of this parasite and shed in the environment the oocysts that become infective after sporulation, representing a risk for other definitive or intermediate hosts (potentially all the warm-blooded animals, including birds, marine mammals and humans) (Dubey et al., 2020).
As reported by FAO/WHO (2014), toxoplasmosis is considered one of the most important food-, water- and soil-borne diseases. It is estimated that approximately two billion of people are infected with T. gondii (Almeria and Dubey, 2021). In a report on foodborne diseases by European Food Safety Authority (EFSA), T. gondii was ranked third in Europe (EFSA, 2018).
The main routes of infection are ingestion of: (i) food or water contaminated with sporulated oocysts (e.g. vegetables, fruit and molluscan shellfish); (ii) uncooked or undercooked meat containing tissue cysts (Ghozzi et al., 2017; Caradonna et al., 2017; EFSA, 2018). Moreover, tachyzoites excreted in milk could be a source of infection. Indeed, outbreaks of toxoplasmosis associated with consumption of unpasteurized goats’ milk have been reported (FAO/WHO, 2014; EFSA, 2018; Almeria and Dubey, 2021). Tachyzoites can also be transmitted vertically from mother to foetus or via organ transplants (Smith et al., 2021).
Although in immunocompetent people toxoplasmosis can usually be asymptomatic or mild symptomatic, this parasite can cause severe consequences in immunocompromised hosts, i.e. ophthalmitis, encephalitis, pneumonitis and myocarditis which can also be fatal (Smith et al., 2021). Moreover, an association between T. gondii infection and neuropsychiatric disorders, as well as personality changes have been reported in several studies (Chaudhury and Ramana, 2019; Almeria and Dubey, 2021). If women become infected during the pregnancy period, miscarriage, stillbirth or congenital defects may occur that can be immediately visible in the newborn or may develop during their lifetime (e.g. neurological problems, mental retardation, deafness and/or ocular lesions) (Almeria and Dubey, 2021).
In animals, symptoms may vary depending on the species. Tachyzoites can be transmitted vertically especially in sheep and goats. However, as in humans, abortion, stillbirth and neonatal death have been reported, mostly in small ruminants, causing economic losses for farmers (Almeria and Dubey, 2021). No confirmed clinical toxoplasmosis reports have been reported in cattle (Lindsay and Dubey, 2020).
Several direct and indirect techniques have been used to detect T. gondii in intermediate hosts and in food products. Among direct techniques, cat and mouse bioassays are the reference methods for assessing viability of the parasite. However, these tests are not easy to use, considering the long time to obtain results and ethical issues, as well as costs (Guo et al., 2015; EFSA 2018; Almeria and Dubey, 2021). Cell cultures can be a valid alternative to bioassays, but they are limited used and protocols are described mainly for fluid samples, while meat homogenated samples gave variable results (Warnekulasuriya et al., 1998; EFSA 2018, Opsteegh et al., 2020; Almeria and Dubey, 2021).
Home-made or commercially available indirect serological tests, e.g. immunofluorescence assay (IFAT), enzyme-linked immunosorbent assay (ELISA), latex agglutination tests (LAT), modified agglutination tests (MAT) and hemagglutination assay (HA), are most commonly used to identify T.gondii positive farms and individual animals, especially for epidemiological surveys on a large number of samples (EFSA, 2018; Almeria and Dubey, 2021). Meat juice has been proven to be an excellent matrix for serological studies on T. gondii in different host species, i.e. sheep, pigs, wild boars, cattle and chickens (Meemken and Blaha, 2011; Basso et al., 2013; Meemken et al., 2014; Bacci et al., 2015; Vismarra et al., 2016, 2017; Slany et al., 2016; Felin et al., 2017; Schares et al., 2018; Olsen et al., 2020; Gazzonis et al., 2020).
Moreover, new serological tools have been developed for the detection of T. gondii in chickens, using Luminex technology (Fabian et al., 2020), a luciferase-linked antibody capture assay (LACA) (Duong et al., 2020) or microarrays to detect antibodies in meat juice and serum (Loreck et al., 2020).
The molecular biology methods are the most widely used direct techniques. Several PCR protocols have been described: end-point, nested PCR and real-time PCR (qPCR) to amplify the B1 gene or the 529 bp repeat element that are the most used targets (Reischl et al., 2003; EFSA, 2018; Almeria and Dubey, 2021). To increase the sensitivity of PCR-based diagnostic methods, a magnetic-capture-(MC-) real-time PCR for detection of T. gondii in meat was developed by Opsteegh et al. (2010), applied to sheep and chicken meat by Schares et al. (2018) and improved by Gisbert Algaba et al. (2017) for diagnosis in pork meat. Moreover, some loop-mediated isothermal amplification (LAMP) protocols have also been developed for the early detection of T. gondii as alternative method to the above-mentioned PCR methods, to increase the sensitivity of available diagnostic techniques (Zhang et al., 2009; Lin et al., 2012; Qu et al., 2013; Zhuo et al., 2015). A commercialized LAMP assay is available for the diagnosis of toxoplasmosis in humans (Varlet-Marie et al., 2018). Moreover, an adaptation of the LAMP technique has been combined with a lateral flow dipstick chromatographic detection system for a rapid visualization of results to detect oocysts in vegetable products (Lalle et al., 2018).
However, the development of new more sensitive and specific diagnostic tools is still ongoing. The droplet digital polymerase chain reaction (ddPCR) is a new PCR method that provides absolute and direct quantification of target DNA, without the need of a standard curve like the qPCR, with a higher sensitivity than other PCR methods (Hindson et al., 2011, 2013).
The ddPCR has been successfully used for detection of different parasites, e.g. Cryptosporidium in different animal hosts and in humans, Echinococcus multilocularis in meadow voles and deer mice, Dirofilaria immitis in dogs, Cytauxzoon felis in cats, gastrointestinal nematodes and Trichuris spp., in sheep and cattle, Eimeria spp. Ascaridia galli and Heterakis gallinarum in chickens, Babesia microti, Babesia duncani, Plasmodium spp., Strongyloides stercoralis and Schistosoma japonicum in humans (Yang et al., 2014; Wilson et al., 2015; Weerakoon et al., 2016; Srisutham et al., 2017; Baltrusis et al., 2019; Mahendran et al., 2020; Yu et al., 2020; Shang Kuan and Pichard, 2020; Kao et al., 2021; Tarbiat et al., 2021; Snyder et al., 2021; Iamrod et al., 2021; Massolo et al., 2021).
The aim of this paper is to develop and validate a new ddPCR assay for detection and quantification of T. gondii DNA in meat of intermediate hosts.
Materials and methods
Preparation of DNA samples
To optimize the ddPCR, specific reference strains were obtained from the American Type Culture Collection (ATCC). The stock solution of Toxoplasma gondii ATCC 50174D contained ~ 2 × 105 cg/µl genome equivalent.
Twenty-five grams of 80 negative samples of minced meat from cattle was homogenized by a stomacher, then total DNA was extracted, using a QIAamp DNA Mini kit (Qiagen, Hilden, Germany), according to the manufacturers’ instructions. The negativity of the samples used was evaluated using the qPCR protocol suggested by the National Reference Centre for toxoplasmosis (Palermo, Italy).
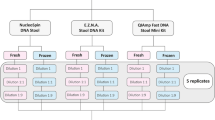
DNA concentration was determined by a Biophotometer (Eppendorf, Hamburg, Germany), then samples were diluted to be analysed by ddPCR at five concentration levels: 8000 cg/µl, 800 cg/µl, 80 cg/µl, 8 cg/µl, to evaluate the limit of detection at 95% of probability (LOD95).
Sensitivity was determined using 40 DNA samples extracted from negative minced meat experimentally contaminated with DNA from the T. gondii ATCC. Ten replicates were prepared for each concentration. Forty samples inoculated only with sterile water were used as negative controls.
Optimization of the ddPCR
Primers and probe (Applied Biosystems, Foster City, CA, USA) used to amplify the region Toxo-529 bp repeat element of the parasite were: forward AF1 CACAGAAGGGACAGAAGT; reverse AF2 TCGCCTTCATCTACAGTC; probe FAM CTCTCCTCCAAGACGGCTGG BHQ (Pepe et al., 2021). Their specificity was evaluated in silico using the NCBI nucleotide BLAST tool and by ddPCR using positive samples for another abortive agent, Neospora caninum.
The annealing temperature for T. gondii was optimized using a thermal gradient (specifically, 56, 56.4, 57.2, 58.4, 59.8, 61, 61.7, and 62 °C temperatures were tested) in a CFX96 (Bio-Rad, Hercules, CA, USA), before developing the ddPCR.
The ddPCR was performed using the QX200 system (Bio-Rad, Hercules, CA, USA). The mastermix was prepared in a total volume of 20 μl, mixing 10 μl of ddPCR Supermix for probes (Bio-Rad, Hercules, CA, USA), 0.5 µM forward primer, 0.5 µM reverse primer, 0.25 µM probe and 35–50 ng for reaction of DNA (Pepe et al., 2021). The reaction mixture was transferred to the DG8 cartridge (Bio-Rad, Hercules, CA, USA). A volume of 70 μl of droplet generation oil was added into the oil well and droplets were formed in the droplet generator (Bio-Rad, Hercules, CA, USA). Then, 40 μl of droplet-partitioned samples were transferred to a 96-well plate and sealed with the specific device. The PCR amplification was carried out on a CFX96 instrument (Bio-Rad, Hercules, CA, USA) with the following thermal profile: 96 °C for 10 min followed by 45 cycles at 98 °C for 30 s, 58.5 °C for 1 min and a final stage at 98 °C for 10 min. After thermal cycling, the 96-well plate was read in the QX200 Droplet Reader, based on positive droplets, according to the Poisson distribution. QuantaSoft software was used to count the PCR-positive and PCR-negative droplets to provide absolute quantification of the target DNA. The quantification measurements of each target were expressed as the number of genomic copies per 1 µl of reaction.
Intra-laboratory repeatability validation was performed by different operators to verify the robustness of the established ddPCR method, calculating the coefficient of variation (CV%) between the assays. Serial dilutions of T. gondii ATCC 50174D genomic DNA from 2 × 105 to 2 gc/µl were analysed.
To evaluate the performance of the ddPCR, the 80 prepared samples (40 negative and 40 positive) were also analysed by qPCR, according to the protocol described in Pepe et al. (2021).
Validation of the developed ddPCR
Overall, 171 diaphragmatic tissue samples (60 cattle, 40 buffaloes, 34 sheep and 37 pigs) collected in slaughterhouses were used for ddPCR validation. An aliquot of 25 g of each sample was weighted and subjected to DNA extraction with the QIAamp DNA Mini commercial kit (Qiagen, Hilden, Germany).
DNA samples were analysed by qPCR (Pepe et al., 2021) and ddPCR (as described above) to compare the results obtained.
The Standards for Reporting of Diagnostic Accuracy Studies (STARD) checklist (https://www.equator-network.org/reporting-guidelines/stard/) was used to report results on performances of techniques (Cohen et al., 2016).
Statistical analysis
The intra-assay CVs for each dilution level (8000, 800, 80 and 8 cg/ml) and overall were calculated by dividing the standard deviation/the arithmetic mean concentration value [(CV% = standard deviation [SD]/mean value for each level) × 100].
Sensitivity, specificity, negative and positive predictive values (NPV and PPV) were calculated for ddPCR, considering the Rt-PCR as gold standard. The agreement between qPCR and ddPCR was calculated using Cohen’s κ statistic (Thrusfield, 2007).
The κ measure was interpreted as follows: 0, no agreement; 0.01–0.20, poor agreement; 0.21–0.40, fair agreement; 0.41–0.60, moderate agreement; 0.61–0.80, substantial agreement; and 0.81–1.0, nearly perfect agreement (Thrusfield, 2007).
The 95% confidence interval (95% CI) was calculated using the free online software “Sample Size Calculator” (Creative Research Systems, CA, USA).
Results
Optimization of the ddPCR
The optimal annealing temperature for ddPCR was 58.5 °C. This temperature was chosen considering the best differentiation of fluorescence between positive and negative samples and avoiding unspecific amplification. No cross-reactions with other parasites, as N. caninum were found in silico and ddPCR assays.
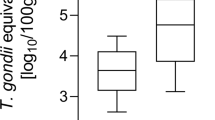
The LoD95 obtained for ddPCR was 8 gc/µl in ddPCR (Fig. 1). The sample was considered positive if showed ≥ two droplets. No positive droplets were detected in the samples used as negative controls. The number of droplets generated for reaction ranged from 8985 to 13,940, with an average of 11,384 droplets. Reactions with more than 8000 accepted droplets for well were used for analysis. The ddPCR data revealed good separation between negative and positive droplets with few interface droplets supporting a high primer specificity and reaction efficiency. At high concentrations (> 10,000 gc/µl), droplets were positively saturated, making the Poisson algorithm invalid and resulting in a relative narrower dynamic range than qPCR.
The sensitivity of ddPCR was 97.5% (95% CI = 85.3–99.9) and specificity 100%. The performance of ddPCR is reported in Table 1. An overall CV% = 9.4 was calculated for all the ddPCR positive replicates (8000 gc/µl CV% = 3.9, 7800 cg/µl CV% = 2.7, 80 cg/µl CV% = 7.4, 8 cg/µl CV% = 23.6). No significant intra-laboratory variation in results was reported (CV% < 0.1).
A nearly perfect agreement (κ = 0.85; p < 0.0001) was found between results obtained by ddPCR and qPCR for positive and negative samples analysed in the development phase. In Table 2, there are reported concentration values obtained by ddPCR and Ct values obtained by qPCR for each level tested.
Validation of ddPCR
Of the 171 samples examined, the qPCR reference method detected T. gondii in only two samples (1.2%; 95% CI = 0.2–4.6) while ddPCR detected 13 positive samples (7.6%; 95% CI = 4.3–12.9). The positive samples not detected by qPCR showed concentrations ranging from 0.3 to 17.1 gc/µl. None of the samples examined, showed an inhibitory effect on PCR, as evidenced by the results of the Internal Amplification Control (IAC).
Discussion
The European Food Safety Authority (EFSA) has suggested that meat-borne transmission accounts for around 60% of human T. gondii infections (EFSA, 2018; Almeira and Dubey, 2021). The main sources of contaminated meat are pork and mutton (Almeira and Dubey, 2021). Although different PCR protocols (i.e. PCR end-point, nested, semi-nested and qPCR) have been developed to detect T. gondii DNA in meat, many published studies have shown that this molecular approach is not very sensitive, due to the inhomogeneous distribution of T. gondii tissue cysts and the small size of the sample used for the analysis (Opsheegh et al., 2010; EFSA, 2018).
For these reasons, more innovative and alternative methods have been developed to increase sensitivity, e.g. the magnetic capture-PCR (mcPCR) and LAMP (Herrmann et al., 2012; Dubey et al., 2021).
In this study, promising results were obtained by ddPCR (7.6% of positive samples to T. gondii vs 1.2% obtained by qPCR). This innovative approach, like qPCR, does not require sequencing of amplified products, because fluorescent-labelled, target-specific probes are used to recognize a desired target.
The main advantage of the ddPCR technique is based on water–oil droplet emulsion technology to distribute the sample solution into ~ 20,000 partitions, increasing the sensitivity, accuracy and precision to detect and quantify also small quantity of the target to amplify (Kao et al., 2021). Moreover, no standard curve is necessary to quantify DNA, indeed droplets contain more target copies and the absolute count can be calculated using Poisson statistics, permitting to evaluate also small differences in target DNA copy numbers among samples (Hindson et al., 2011). Therefore, the ddPCR is more reproducible than qPCR, indeed it is less operator or laboratory dependent, because there is a lower possibility of errors related to pipetting steps to prepare serial dilutions or misinterpretation due to incorrect preparation of standard curve, so a reliable comparison of quantification of target DNA copies can be performed in different laboratories (Huggett et al., 2008). Moreover, the ddPCR is relatively insensitive to potential PCR inhibitors, such as providing lower variation between replicates; therefore, ddPCR is more repeatable than qPCR (Campomenosi et al., 2016).
Further studies will be needed to confirm our preliminary results and to analyse other matrices. Indeed, a limitation of this study is the small number of the positive samples found during the validation phase under “field conditions”.
However, this innovative approach could be very useful for a rapid detection of small amounts of T. gondii in meat, as well as in other food matrices, e.g. milk, cheese, vegetables and molluscs, so to perform valid control strategies, aimed to reduce the risks of toxoplasmosis infection in animals and humans, according to the One Health approach.
Change history
17 July 2022
Missing Open Access funding information has been added in the Funding Note.
References
Almeria S, Dubey JP (2021) Foodborne transmission of Toxoplasma gondii infection in the last decade. An Overview Res Vet Sci 135:371–385
Bacci C, Vismarra A, Mangia C, Bonardi S, Bruini I, Genchi M, Kramer L, Brindani F (2015)
Detection of Toxoplasma gondii in free-range, organic pigs in Italy using serological and molecular methods. Int J Food Microbiol 202:54–6.
Baltrušis P, Halvarsson P, Höglund J (2019) Molecular detection of two major gastrointestinal parasite genera in cattle using a novel droplet digital PCR approach. Parasitol Res 118(10):2901–2907
Basso W, Hartnack S, Pardini L, Maksimov P, Koudela B, Venturini MC, Schares G, Sidler X, Lewis FI, Deplazes P (2013) Assessment of diagnostic accuracy of a commercial ELISA for the detection of Toxoplasma gondii infection in pigs compared with IFAT, TgSAG1-ELISA and Western blot, using a Bayesian latent class approach. Int J Parasitol 43(7):565–570
Campomenosi P, Gini E, Noonan DM, Poli A, D’Antona P, Rotolo N, Dominioni L, Imperatori A (2016) A comparison between quantitative PCR and droplet digital PCR technologies for circulating microRNA quantification in human lung cancer. BMC Biotechnol 16:60
Caradonna T, Marangi M, Del Chierico F, Ferrari N, Reddel S, Bracaglia G, Normanno G, Putignani L, Giangaspero A (2017) Detection and prevalence of protozoan parasites in ready-to-eat packaged salads on sale in Italy. Food Microbiol 67:67–75
Chaudhury A, Ramana BV (2019) Schizophrenia and bipolar disorders: the Toxoplasma connection. Trop Parasitol 9(2):71–76. https://doi.org/10.4103/tp.TP_28_19 (Epub 2019 Sep 18)
Cohen JF, Korevaar DA, Altman DG, Bruns DE, Gatsonis CA, Hooft L, Irwig L, Levine D, Reitsma JB, de Vet HCW, Bossuyt PMM (2016) STARD 2015 guidelines for reporting diagnostic accuracy studies: explanation and elaboration. BMJ Open 6:e012799
Dubey JP, Murata FHA, Cerqueira-Cézar CK, Kwok OCH, Grigg ME (2020) Recent epidemiologic and clinical importance of Toxoplasma gondii infections in marine mammals: 2009–2020. Vet Parasitol 288:109296
Dubey JP, Murata FHA, Cerqueira-Cézar CK, Kwok OCH (2021) Recent epidemiologic and clinical Toxoplasma gondii infections in wild canids and other carnivores: 2009–2020. Vet Parasitol 209:109337
Duong HD, Appiah-Kwarteng C, Takashima Y, Aye KM, Nagayasu E, Yoshida A (2020) A novel luciferase-linked antibody capture assay (LACA) for the diagnosis of Toxoplasma gondii infection in chickens. Parasitol Int 77:102125
EFSA Panel on Biological Hazards (2018) Public health risks associated with food-borne parasites. EFSA Journal, Scientific Opinion 16(12):5495
FAO/WHO (2014). Multicriteria-based ranking for risk management of food-borne parasites. Report of a Joint FAO/WHO Expert Meeting, 3–7 September 2012, FAO Headquarters, Rome, Italy.
Felin E, Näreaho A, Fredriksson-Ahomaa M (2017) Comparison of commercial ELISA tests for the detection of toxoplasma antibodies in the meat juice of naturally infected pigs. Vet Parasitol 238:30–34
Gazzonis AL, Marino AMF, Garippa G, Rossi L, Mignone W, Dini V, Giunta RP, Luini M, Villa L, Zanzani SA, Manfredi MT (2020) Toxoplasma gondii seroprevalence in beef cattle raised in Italy: a multicenter study. Parasitol Res 119(11):3893–3898
Ghozzi K, Marangi M, Papini R, Lahmar I, Challouf R, Houas N, Dhiab RB, Normanno G, Babba H, Giangaspero A (2017) First report of Tunisian coastal water contamination by protozoan parasites using mollusk bivalves as biological indicators. Mar Pollut Bull 117:197–202
Gisbert Algaba I, Geerts M, Jennes M, Coucke W, Opsteegh M, Cox E, Dorny P, Dierick K, De Craeye S (2017) A more sensitive, efficient and ISO 17025 validated magnetic capture real time PCR method for the detection of archetypal Toxoplasma gondii strains in meat. Int J Parasitol 47(13):875–884
Guo M, Buchanan RL, Dubey JP, Hill DE, Lambertini E, Ying Y, Gamble HR, Jones JL, Pradhan AK (2015) Qualitative assessment for Toxoplasma gondii exposure risk associated with meat products in the United States. J Food Prot 78:2207–2219
Herrmann DC, Maksimov P, Maksimov A, Sutor A, Schwarz S, Jaschke W, Schliephake A, Denzin N, Conraths FJ, Schares G (2012) Toxoplasma gondii in foxes and rodents from the German Federal States of Brandenburg and Saxony- Anhalt: seroprevalence and genotypes. Vet Parasitol 185:78–85
Hindson CM, Chevillet JR, Briggs HA, Gallichotte EN, Ruf IK, Hindson BJ, Vessella RL, Tewari M (2013) Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat Methods 10(10):1003–1005
Hindson BJ, Ness KD, Masquelier DA, Belgrader P, Heredia NJ, Makarewicz AJ, Bright IJ, Lucero MY, Hiddessen AL, Legler TC, Kitano TK, Hodel MR, Petersen JF, Wyatt PW, Steenblock ER, Shah PH, Bousse LJ, Troup CB, Mellen JC, Wittmann DK, Erndt NG, Cauley TH, Koehler RT, So AP, Dube S, Rose KA, Montesclaros L, Wang S, Stumbo DP, Hodges SP, Romine S, Milanovich FP, White HE, Regan JF, Karlin-Neumann GA, Hindson CM, Saxonov S, Colston BW (2011) High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal Chem 83(22):8604–8610
Huggett JF, Novak T, Garson JA, Green C, Morris-Jones SD, Miller RF, Zumla A (2008) Differential susceptibility of PCR reactions to inhibitors: an important and unrecognised phenomenon. BMC Res Notes 1:70
Iamrod K, Chaidee A, Rucksaken R, Kopolrat KY, Worasith C, Wongphutorn P, Intuyod K, Pinlaor S, Sithithaworn J, Sithithaworn P, Hongsrichan N (2021). Development and efficacy of droplet digital PCR for detection of Strongyloides stercoralis in stool. Am J Trop Med Hyg, tpmd210729.
Kao YF, Peake B, Madden R, Cowan SR, Scimeca RC, Thomas JE, Reichard MV, Ramachandran A, Miller CA (2021) A probe-based droplet digital polymerase chain reaction assay for early detection of feline acute cytauxzoonosis. Vet Parasitol 292:109413
Lalle M, Possenti A, Dubey JP, Pozio E (2018) Loop-mediated isothermal amplification-lateral-flow dipstick (LAMP-LFD) to detect Toxoplasma gondii oocyst in ready-to-eat salad. Food Microbiol 70:137–142
Lin Z, Zhang Y, Zhang H, Zhou Y, Cao J, Zhou J (2012)
Comparison of loop-mediated isothermal amplification (LAMP) and real-time PCR method targeting a 529-bp repeat element for diagnosis of toxoplasmosis. Vet Parasitol 185(2–4):296–300.
Lindsay DS, Dubey JP (2020) Neosporosis, Toxoplasmosis, and Sarcocystosis in ruminants: an update. Vet Clin North Am Food Anim Pract 36(1):205–222
Loreck K, Mitrenga S, Heinze R, Ehricht R, Engemann C, Lueken C, Ploetz M, Greiner M, Meemken D (2020) Use of meat juice and blood serum with a miniaturised protein microarray assay to develop a multi-parameter IgG screening test with high sample throughput potential for slaughtering pigs. BMC Vet Res 16:106
Mahendran P, Kent Liew JW, Amir A, Ching X, Lau Y (2020) Droplet digital polymerase chain reaction (ddPCR) for the detection of Plasmodium knowlesi and Plasmodium vivax. Malar J 19(1):241
Massolo A, Gerber A, Umhang G, Nicholas C, Liccioli S, Mori K, Klein C (2021) Droplet digital PCR as a sensitive tool to assess exposure pressure from Echinococcusmultilocularis in intermediate hosts. Acta Trop 223:106078
Meemken D, Blaha T (2011) “Meat juice multi-serology” - a tool for the continuous improvement of herd health and food safety in the framework of the risk based meat inspection of slaughter pigs. Arch Leb 62(6):192–199
Meemken D, Tangemann AH, Meermeier D, Gundlach S, Mischok D, Greiner M, Klein G, Blaha T (2014) Establishment of serological herd profiles for zoonoses and production diseases in pigs by “meat juice multi-serology.” Prev Vet Med 113(4):589–598
Olsen A, Sandberg M, Houe H, Nielsen HV, Denwood M, Jensen TB, Alban L (2020) Seroprevalence of Toxoplasma gondii infection in sows and finishers from conventional and organic herds in Denmark: implications for potential future serological surveillance. Prev Vet Med 185:105149
Opsteegh M, Dam-Deisz C, de Boer P, DeCraeye S, Faré A, Hengeveld P, Luiten R, Schares G, van Solt-Smits C, Verhaegen B, Verkleij T, van der Giessen J, Wisselink HJ (2020) Methods to assess the effect of meat processing on viability of Toxoplasma gondii: towards replacement of mouse bioassay by in vitro testing. Int J Parasitol 50(5):357–369
Opsteegh M, Langelaar M, Sprong H, Den Hartog L, De Craeye S, Bokken G, Ajzenberg D, Kijlstra A, Van Der Giessen J (2010) Direct detection and genotyping of Toxoplasma gondii in meat samples using magnetic capture and PCR. Int J Food Microbiol 139:193–201
Pepe P, Bosco A, Capuano F, Baldi L, Giordano A, Mancusi A, Buonanno M, Morena L, Pinto R, Sarnelli P, Cingoli G, Rinaldi L (2021) Towards an integrated approach for monitoring Toxoplasmosis in Southern Italy. Animals 11(7):1949–1961
Qu D, Zhou H, Han J, Tao S, Zheng B, Chi N, Su C, Du A (2013) Development of reverse transcription loop-mediated isothermal amplification (RT-LAMP) as a diagnostic tool of Toxoplasma gondii in pork. Vet Parasitol 192(1–3):98–103
Reischl U, Bretagne S, Krüger D, Ernault P, Costa JM (2003) Comparison of two DNA targets for the diagnosis of Toxoplasmosis by real-time PCR using fluorescence resonance energy transfer hybridization probes. BMC Infect Dis 3:7
Schares G, Koethe M, Bangoura B, Geuthner AC, Randau F, Ludewig M, Maksimov P, Sens M, Bärwald A, Conraths FJ, Villena I, Aubert D, Opsteegh M, Van der Giessen J (2018) Toxoplasma gondii infections in chickens - performance of various antibody detection techniques in serum and meat juice relative to bioassay and DNA detection methods. Int J Parasitol 48(9–10):751–762
Shang Kuan T, Prichard RK (2020) Developmental regulation of Dirofilaria immitis microfilariae and evaluation of ecdysone signaling pathway transcript level using droplet digital PCR. Parasit Vectors 13(1):614
Slany M, Reslova N, Babak V, Lorencova A (2016) Molecular characterization of Toxoplasma gondii in pork meat from different production systems in the Czech Republic. Int J Food Microbiol 238:252–255
Snyder RP, Guerin MT, Hargis BM, Imai R, Kruth PS, Page G, Rejman E, Barta JR (2021) Exploiting digital droplet PCR and next generation sequencing technologies to determine the relative abundance of individual Eimeria species in a DNA sample. Vet Parasitol 296:109443
Srisutham S, Saralamba N, Malleret B, Rénia L, Dondorp AM, Imwong M (2017) Four human Plasmodium species quantification using droplet digital PCR. PLoS One 12(4):e0175771
Smith NC, Goulart C, Hayward JA, Kupz A, Miller CM, van Dooren GG (2021) Control of human toxoplasmosis. Int J Parasitol 51:95–121
Tarbiat B, Enweji N, Baltrusis P, Halvarsson P, Osterman-Lind E, Jansson DS, Höglund J (2021) A novel duplex ddPCR assay for detection and differential diagnosis of Ascaridiagalli and Heterakisgallinarum eggs from chickens feces. Vet Parasitol 296:109499
Varlet-Marie E, Sterkers Y, Perrotte M, Bastien P (2018) ‘Molecular Biology’ working group of the French National Reference Centre for Toxoplasmosis. Int J Parasitol 48(6):457–462
Vismarra A, Barilli E, Miceli M, Mangia C, Bacci C, Brindani F, Kramer L (2017) Toxoplasma gondii and pre-treatment protocols for polymerase chain reaction analysis of milk samples: a field trial in sheep from Southern Italy. Ital J Food Saf 6:6501
Vismarra A, Mangia C, Barilli E, Brindani F, Bacci C, Kramer L (2016)
Meat Juice Serology for Toxoplasma Gondii Infection in Chickens. Ital J Food Saf., 5(1):5586.
Warnekulasuriya MR, Johnson JD, Holliman RE (1998) Detection of Toxoplasma gondii in cured meats. Int J Food Microbiol 45(3):211–215
Weerakoon KG, Gordon CA, Gobert GN, Cai P, McManus DP (2016) Optimisation of a droplet digital PCR assay for the diagnosis of Schistosoma japonicum infection: a duplex approach with DNA binding dye chemistry. J Microbiol Methods 125:19–27
Wilson M, Glaser KC, Adams-Fish D, Boley M, Mayda M, Molestina RE (2015) Development of droplet digital PCR for the detection of Babesia microti and Babesia duncani. Exp Parasitol 149:24–31
Yang R, Paparini A, Monis P, Ryan U (2014) Comparison of next-generation droplet digital PCR (ddPCR) with quantitative PCR (qPCR) for enumeration of Cryptosporidium oocysts in faecal samples. Int J Parasitol 44(14):1105–1113
Yu Z, Zhao Z, Chen L, Li J, Ju X (2020) Development of a droplet digital PCR for detection of Trichuriasis in sheep. J Parasitol 106(5):603–610
Zhang H, Thekisoe OM, Aboge GO, Kyan H, Yamagishi J, Inoue N, Nishikawa Y, Zakimi S, Xuan X (2009) Toxoplasma gondii: sensitive and rapid detection of infection by loop-mediated isothermal amplification (LAMP) method. Exp Parasitol 122(1):47–50
Zhuo X, Huang B, Luo J, Yu H, Yan B, Yang Y, Du A (2015) Development and application of loop-mediated isothermal amplification assays based on ITS-1 for rapid detection of Toxoplasma gondii in pork. Vet Parasitol 208(3–4):246–249
Funding
Open access funding provided by Università degli Studi di Napoli Federico II within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Section Editor: Xing-Quan ZHU
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mancusi, A., Giordano, A., Bosco, A. et al. Development of a droplet digital polymerase chain reaction tool for the detection of Toxoplasma gondii in meat samples. Parasitol Res 121, 1467–1473 (2022). https://doi.org/10.1007/s00436-022-07477-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-022-07477-9