Abstract
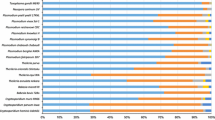
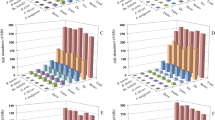
Simple sequence repeats (SSRs) are known to be responsible for genetic complexities and play major roles in gene and genome evolution. To this respect, malaria parasites are known to have rapidly evolving and complex genomes with complicated and differential pathogenic behaviors. Hence, by studying the whole genome comparative SSRs patterns, one can understand genomic complexities and differential evolutionary patterns of these species. We herein utilized the whole genome sequence information of three Plasmodium species, Plasmodium falciparum, Plasmodium vivax, and Plasmodium knowlesi, to comparatively analyze genome-wide distribution of SSRs. The study revealed that despite having the smallest genome size, P. falciparum bears the highest SSR content among the three Plasmodium species. Furthermore, distribution patterns of different SSRs types (e.g., mono, di, tri, tetra, penta, and hexa) in term of relative abundance and relative density provide evidences for greater accumulation of di-repeats and marked decrease of mono-repeats in P. falciparum in comparison to other two species. Overall, the types and distribution of SSRs in P. falciparum genome was found to be different than that of P. vivax and P. knowlesi. The latter two species have quite similar SSR organizations in many aspects of the data. The results were discussed in terms of comparative SSR patterns among the three Plasmodium species, uniqueness of P. falciparum in SSR organization and general pattern of evolution of SSRs in Plasmodium.


Similar content being viewed by others
References
Bell GI, Jurka J (1997) The length distribution of perfect dimer repetitive DNA is consistent with its evolution by an unbiased single-step mutation process. J Mol Evol 44:414–421
Cambareri EB, Jensen BC, Schabtach E, Selker EU (1989) Repeat-induced GC to AT mutations in Neurospora. Science 244:1571–1575
Carlton JM, Adams JH, Silva JC et al (2008) Comparative genomics of the neglected human malaria parasite Plasmodium vivax. Nature 455:757–763
Caserta M, Verdone L (2002) Aspects of nucleosomal positional flexibility and fluidity. Chem Bio Chem 3:1172–1182
Chernin J (2000) Parasite examples grouped according to life cycle. In: Wrigglesworth J (ed) Parasitology. CRC Press, London, p 51
Dieringer D, Schlotterer C (2003) Two distinct modes of microsatellite mutation processes: evidence from the complete genomic sequences of nine species. Genome Res 13:2242–2251
Durand PM, Oelofse AJ, Coetzer TL (2006) An analysis of mobile genetic elements in three Plasmodium species and their potential impact on the nucleotide composition of the P. falciparum genome. BMC Genomics 7:282
Ellegren H (2000a) Heterogeneous mutation processes in human microsatellite DNA sequences. Nat Genet 24:400–402
Ellegren H (2000b) Microsatellite evolution: a battle between replication slippage and point mutation. Trends Genet 18:70
Ellegren H (2004) Microsatellites: simple sequences with complex evolution. Nat Rev Genet 5:435–445
Escalante AA, Ayala FJ (1994) Phylogeny of the malarial genus Plasmodium, derived from rRNA gene sequences. Proc Natl Acad Sci U S A 91:11373–11377
Field D, Wills C (1998) Abundant microsatellite polymorphism in Saccharomyces cerevisiae, and different distributions of microsatellites in eight prokaryotes and S. cerevisiae, result from strong mutation pressures and a variety of selective forces. Proc Natl Acad Sci U S A 95:1647–1652
Fondon JW, Garner HR (2004) Molecular origins of rapid and continuous morphological evolution. Proc Natl Acad Sci U S A 101:18058–18063
Galinski MR, Barnwell JW (2009) Monkey malaria kills four humans. Trends Parasitol 25:200–204
Gardner MJ, Hall N, Fung E et al (2002) Genome sequence of the human malaria parasite Plasmodium falciparum. Nature 419:498–511
Garnica DP, Pinzón AM, Quesada-Ocampo LM, Bernal AJ, Barreto E, Grünwald NJ, Restrepo S (2006) Survey and analysis of microsatellites from transcript sequences in Phytophthora species: frequency, distribution, and potential as markers for the genus. BMC Genomics 7:245
Grayburn WS, Selker EU (1989) A natural case of RIP: degeneration of the DNA sequence in an ancestral tandem duplication. Mol Cell Biol 9:4416–4421
Guo X, Mrazek J (2008) Long simple sequence repeats in host-adapted pathogens localize near genes encoding antigens, housekeeping genes, and pseudogenes. J Mol Evol 67:497–509
Gupta A, Mehra P, Nitharwal R, Sharma A, Biswas AK, Dhar SK (2006) Analogous expression pattern of Plasmodium falciparum replication initiation proteins PfMCM4 and PfORC1 during the asexual and sexual stages of intraerythrocytic developmental cycle. FEMS Microbiol Lett 26:12–18
Gur-Arie R, Cohen CJ, Eitan Y, Shelef L, Hallerman EM, Kashi Y (2000) Simple sequence repeats in Escherichia coli: abundance, distribution, composition, and polymorphism. Genome Res 10:62–71
Hall DB, Holmlin RE, Barton JK (1996) Oxidative DNA damage through long-range electron transfer. Nature 382:731–735
Hancock JM (1993) Evolution of sequences repetition and gene duplication in the TATA-binding protein TBP (TFIID). Nucleic Acids Res 21:2823–2830
Hancock JM (1996) Simple sequences and the expanding genome. BioEssays 18:421–425
Hancock JM (1999) Microsatellites and other simple sequences: genomic context and mutational mechanism. In: Goldstein DB, Schlotterer C (eds) Microsatellites: evolution and applications. Oxford University Press, Oxford, pp 1–7
Hefferon TW (2004) A variable dinucleotide repeat in the CFTR gene contributes to phenotype diversity by forming RNA secondary structures that alter splicing. Proc Natl Acad Sci U S A 101:3504–3509
Hurst GDD, Werren JH (2001) The role of selfish genetic element in eukaryotic evolution. Nat Rev Genet 2:597–606
Jayashree B, Punna P, Prasad P, Bantte K, Tom HC, Chandra S, Hoisington DA, Varshney RK (2006) A database of simple sequence repeats from cereal and legume expressed sequence tags mined in silico: survey and evaluation. In Silico Biol 6:607–620
Jongwutiwes S, Putaporntip C, Iwasaki T, Sata T, Kanbara H (2004) Naturally acquired Plasmodium knowlesi malaria in human, Thailand. Emerg Infect Dis 10:2211–2213
Kantety RV, La Rota M, Matthews DE, Sorrells ME (2002) Data mining for simple sequence repeats in expressed sequence tags from barley, maize, rice, sorghum and wheat. Plant Mol Biol 48:501–510
Karaoglu H, Lee CMY, Meyer W (2005) Survey of simple sequence repeats in completed fungal genomes. Mol Biol Evol 22:639–649
Kashi Y, King DG (2006) Simple sequence repeats as advantageous mutators in evolution. Trends Genet 22:253–259
Katti MV, Ranjekar PK, Gupta VS (2001) Differential distribution of simple sequence repeats in eukaryotic genome sequences. Mol Biol Evol 18:1161–1167
Kelkar YD, Tyekucheva S, Chiaromonte F, Makova KD (2008) The genome-wide determinants of human and chimpanzee microsatellite evolution. Genome Res 18:30–38
Kim TS, Booth JG, Gauch HG, Sun Q, Park J, Lee YH, Lee K (2008) Simple sequence repeats in Neurospora crassa: distribution, polymorphism and evolutionary inferences. BMC Genomics 9:31
King DG, Soller M, Kashi Y (1997) Evolutionary tuning knobs. Endeavour 21:36–40
Kruglyak S, Durrett RT, Schug MD, Aquadro CF (1998) Equilibrium distributions of microsatellite repeat length resulting from a balance between slippage events and point mutations. Proc Natl Acad Sci U S A 95:10774–10778
Kuhlmann M, Borisova BE, Kaller M, Larsson P, Stach D, Na J, Eichinger L, Lyko F, Ambros V, Söderbom F, Hammann C, Nellen W (2005) Silencing of retrotransposons in Dictyostelium by DNA methylation and RNAi. Nucleic Acids Res 33:6405–6417
Kumpatla SP, Mukhopadhyay S (2005) Mining and survey of simple sequence repeats in expressed sequence tags of dicotyledonous species. Genome 48:985–998
Lai Y, Sun F (2003) The relationship between microsatellite slippage mutation rate and the number of repeat units. Mol Biol Evol 20:2123–2131
Li YC, Korol AB, Fahima T, Nevo E (2004) Microsatellites within genes: structure, function and evolution. Mol Biol Evol 21:991–1007
Loire E, Praz F, Higuet D, Netter P, Achaz G (2009) Hypermutability of genes in Homo sapiens due to the hosting of long mono-SSR. Mol Biol Evol 26:111–121
Martinsen ES, Perkins SL, Schall JJ (2008) A three-genome phylogeny of malaria parasites (Plasmodium and closely related genera): evolution of life-history traits and host switches. Mol Phylogenet Evol 47:261–273
Messier M, Li SH, Stewart CB (1996) The birth of microsatellites. Nature 381:483
Moran NA, McLaughlin HJ, Sorek R (2009) The dynamics and time scale of ongoing genomic erosion in symbiotic bacteria. Science 323:379–381
Moxon ER, Rainey PB, Nowak MA, Lenski RE (1994) Adaptive evolution of highly mutable loci in pathogenic bacteria. Curr Biol 4:24–33
Pain A, Bohme U, Berry AE et al (2008) The genome of the simian and human malaria parasite Plasmodium knowlesi. Nature 455:799–804
Perkins SL, Schall J (2002) A molecular phylogeny of malarial parasites recovered from cytochrome b gene sequences. J Parasitol 88:972–978
Phimpraphi W, Paul RE, Yimsamran S et al (2008) Longitudinal study of Plasmodium falciparum and Plasmodium vivax in a Karen population in Thailand. Malar J 7:99
Prasad MD, Muthulakshmi M, Madhu M, Archak S, Mita K, Nagaraju J (2005) Survey and analysis of microsatellites in the Silkworm, Bombyx mori: frequency, distribution, mutations, marker potential and their conservation in heterologous species. Genetics 169:197–214
Pupko T, Graur D (1999) Evolution of microsatellites in the yeast Sacchaomyces cerevisiae: role of length and number of repeated units. J Mol Evol 48:313–316
Richard GF, Dujon B (1997) Trinucleotide repeats in yeast. Res Microbiol 148:731–744
Sattabongkot J, Tsuboi T, Zollner GE, Sirichaisinthop J, Cui L (2004) Plasmodium vivax transmission: chances for control? Trends Parasitol 20:192–198
Schunk M, Kumma WP, Miranda IB, Osman ME, Roewer S, Alano A, Löscher T, Bienzle U, Mockenhaupt FP (2006) High prevalence of drug-resistance mutations in Plasmodium falciparum and Plasmodium vivax in southern Ethiopia. Malar J 5:54
Sen D, Gilbert W (1988) Formation of parallel four-stranded complexes by guanine-rich motifs in DNA and its implications for meiosis. Nature 334:364–366
Sharma PC, Grover A, Kahl G (2007) Mining microsatellites in eukaryotic genomes. Trends Biotech 25:490–498
Singh B, Lee KS, Matusop A, Radhakrishnan A, Shamsul SSG, Cox-Singh J, Thomas A, Conway DJ (2004) A large focus of naturally acquired Plasmodium knowlesi infections in human beings. Lancet 363:1017–1024
Singh V, Mishra N, Awasthi G, Dash AP, Das A (2009) Why is it important to study malaria epidemiology in India? Trends Parasitol 25:452–457
Su X, Carlton JM (2000) Genome display and typing of Plasmodium parasites using anchored polyA and polyT oligonucleotides. Exp Parasitol 94:273–278
Subramanian S, Mishra RK, Singh L (2003) Genome-wide analysis of microsatellite repeats in humans: their abundance and density in specific genomic regions. Genome Biol 4:R13
Suter B, Schnappauf G, Thoma F (2002) Poly (dA.dT) sequences exist as rigid DNA structures in nucleosome-free yeast promotors in vivo. Nucleic Acids Res 28:4083–4089
van Belkum A (1999) Short sequence repeats in microbial pathogenesis and evolution. Cell Mol Life Sci 56:729–734
van der Wel AM, Tomas AM, Kocken CHM, Malhotra P, Jase CJ (1997) Transfection of the primate malaria parasite Plasmodium knowlesi using entirely heterologous constructs. J Exp Med 85:1499–1503
Volkman SK, Lozovsky E, Barry AE, Bedford T, Bethke L, Myrick A, Day KP, Hartl DL, Wirth DF, Sawyer SA (2007) Genomic heterogeneity in the density of non coding single nucleotide and microsatellite polymorphisms in Plasmodium falciparum. Gene 387:1–6
Zhu HM, Li J, Zheng H (2006) Human natural infection of Plasmodium knowlesi. Chinese J parasitol parasitic diseases 24:70–71
Acknowledgment
The authors thank Prof. A. P. Dash, former Director, NIMR for facilities and encouragements. ST and MS are supported from the Indian Council of Medical Research (ICMR), New Delhi. AD thanks the ICMR, New Delhi, for intramural funding support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Suchi Tyagi and Meenu Sharma have contributed equally to the work.
Electronic supplementary materials
Below is the link to the electronic supplementary material.
ESM 1
(DOC 29 kb)
Rights and permissions
About this article
Cite this article
Tyagi, S., Sharma, M. & Das, A. Comparative genomic analysis of simple sequence repeats in three Plasmodium species. Parasitol Res 108, 451–458 (2011). https://doi.org/10.1007/s00436-010-2086-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-010-2086-5