Abstract
Background
The advantages of radiotherapy for head and neck squamous cell carcinoma (HNSCC) depend on the radiation sensitivity of the patient. Here, we established and verified radiological factor-related gene signature and built a prognostic risk model to predict whether radiotherapy would be beneficial.
Methods
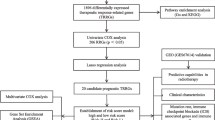
Data from The Cancer Genome Atlas, Gene Expression Omnibus, and RadAtlas databases were subjected to LASSO regression, univariate COX regression, and multivariate COX regression analyses to integrate genomic and clinical information from patients with HNSCC. HNSCC radiation-related prognostic genes were identified, and patients classified into high- and low-risk groups, based on risk scores. Variations in radiation sensitivity according to immunological microenvironment, functional pathways, and immunotherapy response were investigated. Finally, the expression of HNSCC radiation-related genes was verified by qRT-PCR.
Results
We built a clinical risk prediction model comprising a 15-gene signature and used it to divide patients into two groups based on their susceptibility to radiation: radiation-sensitive and radiation-resistant. Overall survival was significantly greater in the radiation-sensitive than the radiation-resistant group. Further, our model was an independent predictor of radiotherapy response, outperforming other clinical parameters, and could be combined with tumor mutational burden, to identify the target population with good predictive value for prognosis at 1, 2, and 3 years. Additionally, the radiation-resistant group was more vulnerable to low levels of immune infiltration, which are significantly associated with DNA damage repair, hypoxia, and cell cycle regulation. Tumor Immune Dysfunction and Exclusion scores also suggested that the resistant group would respond less favorably to immunotherapy.
Conclusions
Our prognostic model based on a radiation-related gene signature has potential for application as a tool for risk stratification of radiation therapy for patients with HNSCC, helping to identify candidates for radiation therapy and overcome radiation resistance.







Similar content being viewed by others
Data availability
The RNA seq data and clinical information in this study are available in the TCGA database (https://portal.gdc.cancer.gov/) and GEO database (https://www.ncbi.nlm.nih.gov/geo/), access number: GSE65858, GSE41613, GSE42743, and the dataset related to radiation exposure can be found in RadAtlas (http://biokb.ncpsb.org.cn/RadAtlas/index.php). All results have been presented in the form of tables and supplementary tables. Details of the data used in this manuscript can be obtained from the corresponding author.
References
Ahn GO, Brown JM (2009) Influence of bone marrow-derived hematopoietic cells on the tumor response to radiotherapy: experimental models and clinical perspectives. Cell Cycle 8(7):970–976
Akervall J, Nandalur S, Zhang J et al (2014) A novel panel of biomarkers predicts radioresistance in patients with squamous cell carcinoma of the head and neck. Eur J Cancer 50(3):570–581
Aran D, Hu Z, Butte AJ (2017) xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol 18(1):1–14
Argiris A, Stenson KM, Brockstein BE et al (2004) Neck dissection in the combined-modality therapy of patients with locoregionally advanced head and neck cancer. Head Neck J Sci Spec Head Neck 26(5):447–455
Ashrafizadeh M, Farhood B, Musa AE, Taeb S, Najafi M (2020) Damage-associated molecular patterns in tumor radiotherapy. Int Immunopharmacol 86:106761
Badoual C, Hans S, Rodriguez J et al (2006) Prognostic value of tumor-infiltrating CD4+ T-cell subpopulations in head and neck cancers. Clin Cancer Res 12(2):465–472
Bai M, Ma X, Li X et al (2015) The accomplices of NF-$κ$B lead to radioresistance. Curr Protein Pept Sci 16(4):279–294
Bartek J, Lukas J, Bartkova J (2007) DNA damage response as an anti-cancer barrier: damage threshold and the concept of’conditional haploinsufficiency’. Cell Cycle 6(19):2344–2347
Becht E, Giraldo NA, Lacroix L et al (2016) Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol 17(1):1–20
Cao W, Shiverick KT, Namiki K et al (2008) Docetaxel and bortezomib downregulate Bcl-2 and sensitize PC-3-Bcl-2 expressing prostate cancer cells to irradiation. World J Urol 26(5):509–516
Chaiswing L, Weiss HL, Jayswal RD, Clair DKS, Kyprianou N (2018) Profiles of radioresistance mechanisms in prostate cancer. Crit Rev Oncog 23(1–2).
Condit PT, Ridings GR, Coin JW, Williams GR, Mitchell D Jr, Boles GW (1964) Methotrexate and radiation in the treatment of patients with cancer. Cancer Res 24(9):1524–1533
Costa-Silva J, Domingues D, Lopes FM (2017) RNA-Seq differential expression analysis: an extended review and a software tool. PLoS ONE 12(12):e0190152
Dewhirst MW, Cao Y, Moeller B (2008) Cycling hypoxia and free radicals regulate angiogenesis and radiotherapy response. Nat Rev Cancer 8(6):425–437
Donlon NE, Power R, Hayes C, Reynolds JV, Lysaght J (2021) Radiotherapy, immunotherapy, and the tumour microenvironment: turning an immunosuppressive milieu into a therapeutic opportunity. Cancer Lett 502:84–96
Geng J, Zhang Y, Gao Q et al (2021) Switching on prodrugs using radiotherapy. Nat Chem 13(8):805–810
Haimovitz-Friedman A, Kolesnick RN, Fuks Z (1996) Modulation of the apoptotic response: potential for improving the outcome in clinical radiotherapy. Semin Radiat Oncol 6:273–283
Hänzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform 14(1):1–15
Hipp SJ, Goldman S, Kaushal A et al (2020) A phase I trial of lenalidomide and radiotherapy in children with diffuse intrinsic pontine gliomas or high-grade gliomas. J Neurooncol 149(3):437–445
Hodi FS, Wolchok JD, Schadendorf D et al (2021) TMB and inflammatory gene expression associated with clinical outcomes following immunotherapy in advanced melanoma. Cancer Immunol Res 9(10):1202–1213
Huang T, Bi Y, Cui Z, Guan J, Huang Y (2020) MUC1 confers radioresistance in head and neck squamous cell carcinoma (HNSCC) cells. Bioengineered 11(1):769–778
Hutchinson MKND, Mierzwa M, D’Silva NJ (2020) Radiation resistance in head and neck squamous cell carcinoma: dire need for an appropriate sensitizer. Oncogene 39(18):3638–3649
Jeganathan J, Saraf R, Mahmood F et al (2017) Mitochondrial dysfunction in atrial tissue of patients developing postoperative atrial fibrillation. Ann Thorac Surg 104(5):1547–1555
Jiang P, Gu S, Pan D et al (2018) Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med 24(10):1550–1558
Jiang W, He Y, He W et al (2021) Exhausted CD8+ T cells in the tumor immune microenvironment: new pathways to therapy. Front Immunol 11:622509
Kabakov AE, Yakimova AO (2021) Hypoxia-induced cancer cell responses driving radioresistance of hypoxic tumors: approaches to targeting and radiosensitizing. Cancers (basel) 13(5):1102
Kassambara A, Kosinski M, Biecek P, Fabian S (2017) Package ‘survminer.’ Draw Surviv Curves using ‘ggplot2’(R Packag version 03 1)
Kaur S, Nag A, Gangenahalli G, Sharma K (2019) Peroxisome proliferator activated receptor gamma sensitizes non-small cell lung carcinoma to gamma irradiation induced apoptosis. Front Genet 10:554
Ke ZB, Wu YP, Huang P et al (2021) Identification of novel genes in testicular cancer microenvironment based on ESTIMATE algorithm-derived immune scores. J Cell Physiol 236(1):706–713
Kim BM, Hong Y, Lee S et al (2015) Therapeutic implications for overcoming radiation resistance in cancer therapy. Int J Mol Sci 16(11):26880–26913
Li J, Zeng Z, Wu Q et al (2021a) Immunological modulation of the Th1/Th2 shift by ionizing radiation in tumors. Int J Oncol 59(1):1–12
Li R, Che W, Liang N, Deng S, Song Z, Yang L (2021b) Silent FOSL1 enhances the radiosensitivity of glioma stem cells by down-regulating miR-27a-5p. Neurochem Res 46(12):3222–3246
Liberzon A, Birger C, Thorvaldsdóttir H, Ghandi M, Mesirov JP, Tamayo P (2015) The molecular signatures database hallmark gene set collection. Cell Syst 1(6):417–425
Mims J, Bansal N, Bharadwaj MS et al (2015) Energy metabolism in a matched model of radiation resistance for head and neck squamous cell cancer. Radiat Res 183(3):291–304
Müller C, Schillert A, Röthemeier C et al (2016) Removing batch effects from longitudinal gene expression-quantile normalization plus ComBat as best approach for microarray transcriptome data. PLoS ONE 11(6):e0156594
Muzaffar J, Bari S, Kirtane K, Chung CH (2021) Recent advances and future directions in clinical management of head and neck squamous cell carcinoma. Cancers (basel) 13(2):338
Parikh RA, Appleman LJ, Bauman JE et al (2014) Upregulation of the ATR-CHEK1 pathway in oral squamous cell carcinomas. Genes Chromosomes Cancer 53(1):25–37
Plattner C, Finotello F, Rieder D (2020) Deconvoluting tumor-infiltrating immune cells from RNA-seq data using quanTIseq. Methods Enzymol 636:261–285
Qiu Y, Chen T, Hu R et al (2021) Next frontier in tumor immunotherapy: macrophage-mediated immune evasion. Biomark Res 9(1):1–19
Reits EA, Hodge JW, Herberts CA et al (2006) Radiation modulates the peptide repertoire, enhances MHC class I expression, and induces successful antitumor immunotherapy. J Exp Med 203(5):1259–1271
Ritchie ME, Phipson B, Wu DI et al (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43(7):e47–e47
Seidel JA, Otsuka A, Kabashima K (2018) Anti-PD-1 and anti-CTLA-4 therapies in cancer: mechanisms of action, efficacy, and limitations. Front Oncol 8:86
Sheridan MT, O’Dwyer T, Seymour CB, Mothersill CE (1997) Potential indicators of radiosensitivity in squamous cell carcinoma of the head and neck. Radiat Oncol Investig Clin Basic Res 5(4):180–186
Shintani S, Mihara M, Li C et al (2003) Up-regulation of DNA-dependent protein kinase correlates with radiation resistance in oral squamous cell carcinoma. Cancer Sci 94(10):894–900
Sørensen BS, Busk M, Olthof N et al (2013) Radiosensitivity and effect of hypoxia in HPV positive head and neck cancer cells. Radiother Oncol 108(3):500–505
Srinivas US, Tan BWQ, Vellayappan BA, Jeyasekharan AD (2019) ROS and the DNA damage response in cancer. Redox Biol 25:101084
Stanzani E, Martínez-Soler F, Mateos TM et al (2017) Radioresistance of mesenchymal glioblastoma initiating cells correlates with patient outcome and is associated with activation of inflammatory program. Oncotarget 8(43):73640
Sun J, Chen Y, Li M, Ge Z (1998) Role of antioxidant enzymes on ionizing radiation resistance. Free Radic Biol Med 24(4):586–593
Suwa T, Kobayashi M, Nam JM, Harada H (2021) Tumor microenvironment and radioresistance. Exp Mol Med 53(6):1029–1035
Tang L, Wei F, Wu Y et al (2018) Role of metabolism in cancer cell radioresistance and radiosensitization methods. J Exp Clin Cancer Res 37(1):1–15
Templin T, Paul S, Amundson SA et al (2011) Radiation-induced micro-RNA expression changes in peripheral blood cells of radiotherapy patients. Int J Radiat Oncol Biol Phys 80(2):549–557
Tibshirani R (1997) The lasso method for variable selection in the Cox model. Stat Med 16(4):385–395
Ubago-Guisado E, Rodríguez-Barranco M, Ching-López A et al (2021) Evidence update on the relationship between diet and the most common cancers from the European prospective Investigation into Cancer and Nutrition (EPIC) study: a systematic review. Nutrients 13(10):3582
Vaes RDW, Hendriks LEL, Vooijs M, De Ruysscher D (2021) Biomarkers of radiotherapy-induced immunogenic cell death. Cells 10(4):930
Vitale M, Cantoni C, Pietra G, Mingari MC, Moretta L (2014) Effect of tumor cells and tumor microenvironment on NK-cell function. Eur J Immunol 44(6):1582–1592
Wang H, Sethi G, Loke WK, Sim MK (2015) Des-aspartate-angiotensin I attenuates mortality of mice exposed to gamma radiation via a novel mechanism of action. PLoS ONE 10(9):e0138009
Wang Y, Deng W, Li N et al (2018) Combining immunotherapy and radiotherapy for cancer treatment: current challenges and future directions. Front Pharmacol 9:185
Wang Y, Tu W, Tang Y, Zhang S (2020) Prevention and treatment for radiation-induced skin injury during radiotherapy. Radiat Med Prot 1(02):60–68
Widodo SS, Hutchinson RA, Fang Y et al (2021) Toward precision immunotherapy using multiplex immunohistochemistry and in silico methods to define the tumor immune microenvironment. Cancer Immunol Immunother 70(7):1811–1820
Wong KCW, Johnson D, Hui EP, Lam RCT, Ma BBY, Chan ATC (2022) Opportunities and challenges in combining immunotherapy and radiotherapy in head and neck cancers. Cancer Treat Rev 105:102361
Xu T, Zhang Y, Chang P, Gong S, Shao L, Dong L (2018) Mesenchymal stem cell-based therapy for radiation-induced lung injury. Stem Cell Res Ther 9(1):1–7
Xu H, Liu Y, Li Y et al (2020) RadAtlas 1.0: a knowledgebase focusing on radiation-associated genes. Int J Radiat Biol 96(8):980–987
Yang C, Jiang L, Zhang H, Shimoda LA, DeBerardinis RJ, Semenza GL (2014) Analysis of hypoxia-induced metabolic reprogramming. Methods Enzymol 542:425–455
Yang QX, Wang YX, Li FC et al (2019) Identification of the gene signature reflecting schizophrenia’s etiology by constructing artificial intelligence-based method of enhanced reproducibility. CNS Neurosci Ther 25(9):1054–1063
Zeng D, Ye Z, Shen R et al (2021) IOBR: multi-omics Immuno-oncology biological research to decode tumor microenvironment and signatures. Front Immunol 12:687975
Zhan Y, Fan S (2020) Multiple mechanisms involving in radioresistance of nasopharyngeal carcinoma. J Cancer 11(14):4193
Zhang L, Li B, Peng Y et al (2020) The prognostic value of TMB and the relationship between TMB and immune infiltration in head and neck squamous cell carcinoma: a gene expression-based study. Oral Oncol 110:104943
Acknowledgements
We appreciate the data obtained from TCGA, GEO and RadAtlas. We sincerely thank all the people involved.
Funding
This study was funded by the Natural Science Foundation of Guangdong (Grant Number 2021A1515010838), Science and Technology Program of Guangzhou (Grant Number 201903010028), The Project Supported by Natural Science Foundation of Jiangxi (20192BAB205065), The Excellent Young Scientists Fund of Jiangxi Cancer Hospital (2021EYS04), Guangdong Provincial People’s Hospital Intermural Program (Grant Number KJ012019447), The Nation Cancer Center Climb Plan (NCC201914B05).
Author information
Authors and Affiliations
Contributions
PY and SL, QL contributed to conceptualization, data curation, formal analysis, data curation, software and project administration and writing manuscript. HQ, YG contributed to software, visualization and manuscript editing. CG, ZG and TW contributed to the methodology and resources. DX and LY contributed to manuscript review. HZ and JL contributed to project administration and supervision.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Ethics approval and consent to participate
The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
432_2023_5304_MOESM1_ESM.tif
Supplementary file1 Supplementary Figure 1. (A) Survival analysis and ROC curves of the complete TCGA cohort. (B)- (I) Survival analysis of different clinical subgroups (treatment mode, age, sex, TNM stage, stage, grade) (TIF 5841 KB)
432_2023_5304_MOESM2_ESM.tif
Supplementary file2 Supplementary Figure 2. (A) 15 genes biological function enrichment. (B) Immunotherapy response in radiosensitive and radioresistant groups. (C) Expression of hypoxia factors in HPV+ patients. (D) Risk Score and Immune Cell Correlation. (E)Top 10 genes with mutation frequency among 15 genes. (F) TMB score of radiation sensitive group and radiation resistant group. (G) The 15 genes of this model are expressed in HNSCC. (H) The 15 genes of this model are expressed in different HNSCC cell lines from CCLE database. (I) Results of qRT-PCR of five genes expressed in cell lines (TIF 10014 KB)
432_2023_5304_MOESM3_ESM.tif
Supplementary file3 Supplementary Figure 3. (A) The performance of each model on all validation datasets was calculated using machine learning combination algorithms. (B)GSE41613 validation cohort and (C) GSE42743 validation cohort KM survival curve, and 1-year, 2-year, 3-year ROC curve.(D) Manage test calculates risk scores, immune microenvironment scores, immune checkpoints(E)and signature correlation heatmaps (TIF 17616 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
You, P., Liu, S., Li, Q. et al. Radiation-sensitive genetic prognostic model identifies individuals at risk for radiation resistance in head and neck squamous cell carcinoma. J Cancer Res Clin Oncol 149, 15623–15640 (2023). https://doi.org/10.1007/s00432-023-05304-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-05304-x