Abstract
Purpose
In non-small cell lung cancer (NSCLC), success of targeted therapy has promoted researches explicitly orientated based on genetic background. Although PTEN deficiency is common in NSCLC, carcinogenesis about such genetic type has not been fully explored. Here, we have found that classical tumor suppressor P53 could be modulated by deacetylase sirtuin-3 (SIRT3) depending on the PTEN condition in NSCLC, which may be a novel breakpoint for handling PTEN deficiency NSCLC.
Methods
First, we examined SIRT3 and P53 expression files in PTEN-deficient NSCLC clinical samples and investigated their correlation. Second, we built SIRT3 high or low expression models in different PTEN conditions by plasmid overexpression or si-RNA interference in NSCLC cell lines and explored the effect of SIRT3 upon P53. Furthermore, we investigated the influence of SIRT3 upon the ubiquitin–proteasome dependent degradation pathway of P53 in PTEN-deficient NSCLC cell lines. Finally, we probed into the deacetylation modification of P53 via SIRT3.
Results
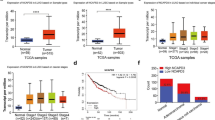
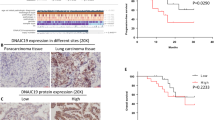
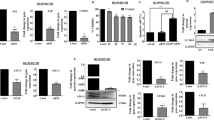
We found that SIRT3 expression was strongly positive and P53 expression was almost negative in PTEN-deficient NSCLC clinical samples. Further, we demonstrated that SIRT3 promoted degradation of P53 in PTEN-deficient NSCLC cell lines via the ubiquitin–proteasome pathway. Finally, we demonstrated that SIRT3 could deacetylate P53 at lysines 320 and 382, which may account for the observed degradation of P53 in PTEN-deficient tumor cells.
Conclusions
We have identified a novel mechanism by which P53 was inactivated via SIRT3 in PTEN-deficient cells. This may shed light on the mechanisms underlying the malignancy of PTEN-deficient NSCLC.






Similar content being viewed by others
References
Bepler G et al (2004) RRM1 and PTEN as prognostic parameters for overall and disease-free survival in patients with non-small-cell lung cancer. J Clin Oncol Off J Am Soc Clin Oncol 22:1878–1885. https://doi.org/10.1200/jco.2004.12.002
Chen Z, Fillmore CM, Hammerman PS, Kim CF, Wong KK (2014a) Non-small-cell lung cancers: a heterogeneous set of diseases. Nat Rev Cancer 14:535–546. https://doi.org/10.1038/nrc3775
Chen Z et al (2014b) Role of the stem cell-associated intermediate filament nestin in malignant proliferation of non-small cell lung cancer. PLoS One 9:e85584. https://doi.org/10.1371/journal.pone.0085584
Choudhary C et al (2009) Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science (New York, NY) 325:834–840. https://doi.org/10.1126/science.1175371
Deal RA, Tang Y, Fletcher R, Torquati A, Omotosho P (2017) Understanding intestinal glucose transporter expression in obese compared to non-obese subjects. Surg Endosc. https://doi.org/10.1007/s00464-017-5858-5
Goldstraw P, Ball D, Jett JR, Le Chevalier T, Lim E, Nicholson AG, Shepherd FA (2011) Non-small-cell lung cancer. Lancet (Lond Engl) 378:1727–1740. https://doi.org/10.1016/s0140-6736(10)62101-0
Guan KL, Xiong Y (2011) Regulation of intermediary metabolism by protein acetylation. Trends Biochem Sci 36:108–116. https://doi.org/10.1016/j.tibs.2010.09.003
Hao HH et al (2013) Valproic acid reduces autophagy and promotes functional recovery after spinal cord injury in rats. Neurosci Bull 29:484–492. https://doi.org/10.1007/s12264-013-1355-6
Honda R, Tanaka H, Yasuda H (1997) Oncoprotein MDM2 is a ubiquitin ligase E3 for tumor suppressor p53. FEBS Lett 420:25–27
Ito A, Lai CH, Zhao X, Saito S, Hamilton MH, Appella E, Yao TP (2001) p300/CBP-mediated p53 acetylation is commonly induced by p53-activating agents and inhibited by MDM2. EMBO J 20:1331–1340. https://doi.org/10.1093/emboj/20.6.1331
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D (2011) Global cancer statistics. CA Cancer J Clin 61:69–90. https://doi.org/10.3322/caac.20107
Kim SC et al (2006) Substrate and functional diversity of lysine acetylation revealed by a proteomics survey. Mol Cell 23:607–618. https://doi.org/10.1016/j.molcel.2006.06.026
Li S, Banck M, Mujtaba S, Zhou M-M, Sugrue MM, Walsh MJ (2010) p53-induced growth arrest is regulated by the mitochondrial SirT3 deacetylase. PLoS One. https://doi.org/10.1371/journal.pone.0010486
Luo J, Su F, Chen D, Shiloh A, Gu W (2000) Deacetylation of p53 modulates its effect on cell growth and apoptosis. Nature 408:377–381. https://doi.org/10.1038/35042612
Marsit CJ, Zheng S, Aldape K, Hinds PW, Nelson HH, Wiencke JK, Kelsey KT (2005) PTEN expression in non-small-cell lung cancer: evaluating its relation to tumor characteristics, allelic loss, and epigenetic alteration. Hum Pathol 36:768–776. https://doi.org/10.1016/j.humpath.2005.05.006
Mitsudomi T et al (1992) p53 gene mutations in non-small-cell lung cancer cell lines and their correlation with the presence of ras mutations and clinical features. Oncogene 7:171–180
Molina JR, Yang P, Cassivi SD, Schild SE, Adjei AA (2008) Non-small cell lung cancer: epidemiology, risk factors, treatment, and survivorship. Mayo Clin Proc 83:584–594. https://doi.org/10.4065/83.5.584
Nakamura S, Roth JA, Mukhopadhyay T (2000) Multiple lysine mutations in the C-terminal domain of p53 interfere with MDM2-dependent protein degradation and ubiquitination. Mol Cell Biol 20:9391–9398
Oren M et al (2002) Regulation of p53: intricate loops and delicate balances. Ann N Y Acad Sci 973:374–383
Pao W, Girard N (2011) New driver mutations in non-small-cell lung cancer. Lancet Oncol 12:175–180. https://doi.org/10.1016/s1470-2045(10)70087-5
Park SJ, More S, Murtuza A, Woodward BD, Husain H (2017) New targets in non-small cell lung cancer. Hematol Oncol Clin N Am 31:113–129. https://doi.org/10.1016/j.hoc.2016.08.010
Robles AI, Linke SP, Harris CC (2002) The p53 network in lung carcinogenesis. Oncogene 21:6898–6907. https://doi.org/10.1038/sj.onc.1205563
Sakaguchi K et al (1998) DNA damage activates p53 through a phosphorylation-acetylation cascade. Genes Dev 12:2831–2841
Salmena L, Carracedo A, Pandolfi PP (2008) Tenets of PTEN tumor suppression. Cell 133:403–414. https://doi.org/10.1016/j.cell.2008.04.013
Schiller JH et al (2002) Comparison of four chemotherapy regimens for advanced non-small-cell lung cancer. N Engl J Med 346:92–98. https://doi.org/10.1056/NEJMoa011954
Selivanova G (2004) p53: fighting cancer. Curr Cancer Drug Targets 4:385–402
Song MS, Salmena L, Pandolfi PP (2012) The functions and regulation of the PTEN tumour suppressor. Nat Rev Mol Cell Biol 13:283–296. https://doi.org/10.1038/nrm3330
Soria JC et al (2002) Lack of PTEN expression in non-small cell lung cancer could be related to promoter methylation. Clin Cancer Res Off J Am Assoc Can Res 8:1178–1184
Spoerke JM et al (2012) Phosphoinositide 3-kinase (PI3 K) pathway alterations are associated with histologic subtypes and are predictive of sensitivity to PI3K inhibitors in lung cancer preclinical models. Clin Cancer Res Off J Am Assoc Can Res 18:6771–6783. https://doi.org/10.1158/1078-0432.ccr-12-2347
Takahashi T et al (1992) Wild-type but not mutant p53 suppresses the growth of human lung cancer cells bearing multiple genetic lesions. Can Res 52:2340–2343
Tang JM, He QY, Guo RX, Chang XJ (2006) Phosphorylated Akt overexpression and loss of PTEN expression in non-small cell lung cancer confers poor prognosis. Lung Cancer 51:181–191. https://doi.org/10.1016/j.lungcan.2005.10.003
Thomas A, Liu SV, Subramaniam DS, Giaccone G (2015) Refining the treatment of NSCLC according to histological and molecular subtypes Nature reviews. Clin Oncol 12:511–526. https://doi.org/10.1038/nrclinonc.2015.90
Wade M, Li YC, Wahl GM (2013) MDM2, MDMX and p53 in oncogenesis and cancer therapy. Nat Rev Cancer 13:83–96. https://doi.org/10.1038/nrc3430
Wang D et al (2016) Acetylation-regulated interaction between p53 and SET reveals a widespread regulatory mode. Nature 538:118–122. https://doi.org/10.1038/nature19759
Woods DB, Vousden KH (2001) Regulation of p53 function. Exp Cell Res 264:56–66. https://doi.org/10.1006/excr.2000.5141
Xiao K et al (2013) Sirt3 is a tumor suppressor in lung adenocarcinoma cells. Oncol Rep 30:1323–1328. https://doi.org/10.3892/or.2013.2604
Xiong Y et al (2017) SIRT3 is correlated with the malignancy of non-small cell lung cancer. Int J Oncol. https://doi.org/10.3892/ijo.2017.3868
Zhang Y-Y, Zhou L-M (2012) Sirt3 inhibits hepatocellular carcinoma cell growth through reducing Mdm2-mediated p53 degradation. Biochem Biophys Res Commun 423:26–31. https://doi.org/10.1016/j.bbrc.2012.05.053
Zhao K et al (2015) Oroxylin A promotes PTEN-mediated negative regulation of MDM2 transcription via SIRT3-mediated deacetylation to stabilize p53 and inhibit glycolysis in wt-p53 cancer cells. J Hematol Oncol. https://doi.org/10.1186/s13045-015-0137-1
Acknowledgements
We thank the staff at the Department of Thoracic Surgery, Tangdu Hospital, Fourth Military Medical University, Xi’an, China, and the State Key Laboratory of Cancer Biology, Department of Biochemistry and Molecular Biology, Fourth Military Medical University, Xi’an, China, for technical support.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Funding
This study was funded by the Special Clinical Research Fund from Wu JiePing Medical Foundation (320.6750.15095), and the Natural Science and Technology Foundation Research Key Project of Shaanxi Province (2015JZ 024), China.
Additional information
Dr. Yong Han was considered to be the corresponding author and Dr. Lintao Jia was considered to be the co-corresponding author.
Rights and permissions
About this article
Cite this article
Xiong, Y., Wang, L., Wang, S. et al. SIRT3 deacetylates and promotes degradation of P53 in PTEN-defective non-small cell lung cancer. J Cancer Res Clin Oncol 144, 189–198 (2018). https://doi.org/10.1007/s00432-017-2537-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-017-2537-9