Abstract
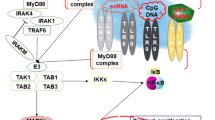
The innate immune system provides the first line of host defence against invading pathogens. Key to upregulation of the innate immune response are Toll-like receptors (TLRs), which recognize pathogen-associated molecular patterns (PAMPs) and trigger a signaling pathway culminating in the production of inflammatory mediators. Central to this TLR signaling pathway are heterotypic protein–protein interactions mediated through Toll/interleukin-1 receptor (TIR) domains found in both the cytoplasmic regions of TLRs and adaptor proteins. Pathogenic bacteria have developed a range of ingenuous strategies to evade the host immune mechanisms. Recent work has identified a potentially novel evasion mechanism involving bacterial TIR domain proteins. Such domains have been identified in a wide range of pathogenic bacteria, and there is evidence to suggest that they interfere directly with the TLR signaling pathway and thus inhibit the activation of NF-κB. The individual TIR domains from the pathogenic bacteria Salmonella enterica serovar Enteritidis, Brucella sp, uropathogenic E. coli and Yersinia pestis have been analyzed in detail. The individual bacterial TIR domains from these pathogenic bacteria seem to differ in their modes of action and their roles in virulence. Here, we review the current state of knowledge on the possible roles and mechanisms of action of the bacterial TIR domains.



Similar content being viewed by others
References
Akira S, Uematsu S, Takeuchi O (2006) Pathogen recognition and innate immunity. Cell 124:783–801
Fritz JH, Le Bourhis L, Sellge G, Magalhaes JG, Fsihi H, Kufer TA, Collins C, Viala J, Ferrero RL, Girardin SE et al (2007) Nod1-mediated innate immune recognition of peptidoglycan contributes to the onset of adaptive immunity. Immunity 26:445–459
Kumar H, Kawai T, Akira S (2009) Pathogen recognition in the innate immune response. Biochem J 420:1–16
Gay NJ, Gangloff M (2007) Structure and function of Toll receptors and their ligands. Annu Rev Biochem 76:141–165
Akira S, Takeda K (2004) Toll-like receptor signalling. Nat Rev Immunol 4:499–511
Hashimoto C, Hudson KL, Anderson KV (1988) The Toll gene of Drosophila, required for dorsal-ventral embryonic polarity, appears to encode a transmembrane protein. Cell 52:269–279
Gay NJ, Keith FJ (1991) Drosophila Toll and IL-1 receptor. Nature 351:355–356
Triantafilou M, Gamper FGJ, Haston RM, Mouratis MA, Morath S, Hartung T, Triantafilou K (2006) Membrane sorting of toll-like receptor (TLR)-2/6 and TLR2/1 heterodimers at the cell surface determines heterotypic associations with CD36 and intracellular targeting. J Biol Chem 281:31002–31011
Latz E, Verma A, Visintin A, Gong M, Sirois CM, Klein DCG, Monks BG, McKnight CJ, Lamphier MS, Duprex WP et al (2007) Ligand-induced conformational changes allosterically activate Toll-like receptor 9. Nat Immunol 8:772–779
O’Neill LAJ, Bowie AG (2007) The family of five: TIR-domain-containing adaptors in Toll-like receptor signalling. Nat Rev Immunol 7:353–364
Núñez Miguel R, Wong J, Westoll JF, Brooks HJ, O’Neill LAJ, Gay NJ, Bryant CE, Monie TP (2007) A dimer of the Toll-like receptor 4 cytoplasmic domain provides a specific scaffold for the recruitment of signalling adaptor proteins. PLoS One 2:e788
Brown V, Brown RA, Ozinsky A, Hesselberth JR, Fields S (2006) Binding specificity of Toll-like receptor cytoplasmic domains. Eur J Immunol 36:742–753
Horng T, Barton GM, Flavell RA, Medzhitov R (2002) The adaptor molecule TIRAP provides signalling specificity for Toll-like receptors. Nature 420:329–333
Rowe DC, McGettrick AF, Latz E, Monks BG, Gay NJ, Yamamoto M, Akira S, O’Neill LA, Fitzgerald KA, Golenbock DT (2006) The myristoylation of TRIF-related adaptor molecule is essential for Toll-like receptor 4 signal transduction. Proc Natl Acad Sci USA 103:6299–6304
Carty M, Goodbody R, Schröder M, Stack J, Moynagh PN, Bowie AG (2006) The human adaptor SARM negatively regulates adaptor protein TRIF-dependent Toll-like receptor signaling. Nat Immunol 7:1074–1081
Khan JA, Brint EK, O’Neill LAJ, Tong L (2004) Crystal structure of the Toll/interleukin-1 receptor domain of human IL-1RAPL. J Biol Chem 279:31664–31670
Slack JL, Schooley K, Bonnert TP, Mitcham JL, Qwarnstrom EE, Sims JE, Dower SK (2000) Identification of two major sites in the type I interleukin-1 receptor cytoplasmic region responsible for coupling to pro-inflammatory signaling pathways. J Biol Chem 275:4670–4678
Poltorak A, He X, Smirnova I, Liu MY, van Huffel C, Du X, Birdwell D, Alejos E, Silva M, Galanos C et al (1998) Defective LPS signaling in C3H/HeJ and C57BL/10ScCr mice: mutations in Tlr4 gene. Science 282:2085–2088
Xu Y, Tao X, Shen B, Horng T, Medzhitov R, Manley JL, Tong L (2000) Structural basis for signal transduction by the Toll/interleukin-1 receptor domains. Nature 408:111–115
Nyman T, Stenmark P, Flodin S, Johansson I, Hammarström M, Nordlund P (2008) The crystal structure of the human toll-like receptor 10 cytoplasmic domain reveals a putative signaling dimer. J Biol Chem 283:11861–11865
Ohnishi H, Tochio H, Kato Z, Orii KE, Li A, Kimura T, Hiroaki H, Kondo N, Shirakawa M (2009) Structural basis for the multiple interactions of the MyD88 TIR domain in TLR4 signaling. Proc Natl Acad Sci USA 106:10260–10265
Valkov E, Stamp A, Dimaio F, Baker D, Verstak B, Roversi P, Kellie S, Sweet MJ, Mansell A, Gay NJ et al (2011) Crystal structure of Toll-like receptor adaptor MAL/TIRAP reveals the molecular basis for signal transduction and disease protection. Proc Natl Acad Sci USA 108:14879–14884
Watters TM, Kenny EF, O’Neill LAJ (2007) Structure, function and regulation of the Toll/IL-1 receptor adaptor proteins. Immunol Cell Biol 85:411–419
Kawahara K, Tsukano H, Watanabe H, Lindner B, Matsuura M (2002) Modification of the structure and activity of lipid A in Yersinia pestis lipopolysaccharide by growth temperature. Infect Immun 70:4092–4098
Nigou J, Zelle-Rieser C, Gilleron M, Thurnher M, Puzo G (2001) Mannosylated lipoarabinomannans inhibit IL-12 production by human dendritic cells: evidence for a negative signal delivered through the mannose receptor. J Immunol 166:7477–7485
Stack J, Haga IR, Schröder M, Bartlett NW, Maloney G, Reading PC, Fitzgerald KA, Smith GL, Bowie AG (2005) Vaccinia virus protein A46R targets multiple Toll-like-interleukin-1 receptor adaptors and contributes to virulence. J Exp Med 201:1007–1018
Bowie A, Kiss-Toth E, Symons JA, Smith GL, Dower SK, O’Neill LA (2000) A46R and A52R from vaccinia virus are antagonists of host IL-1 and toll-like receptor signaling. Proc Natl Acad Sci USA 97:10162–10167
Harte MT, Haga IR, Maloney G, Gray P, Reading PC, Bartlett NW, Smith GL, Bowie A, O’Neill LAJ (2003) The poxvirus protein A52R targets Toll-like receptor signaling complexes to suppress host defense. J Exp Med 197:343–351
Spear AM, Loman NJ, Atkins HS, Pallen MJ (2009) Microbial TIR domains: not necessarily agents of subversion? Trends Microbiol 17:393–398
Lang T, Lo C, Skinner N, Locarnini S, Visvanathan K, Mansell A (2011) The Hepatitis B e antigen (HBeAg) targets and suppresses activation of the Toll-like receptor signaling pathway. J Hepatol. doi:10.1016/j.jhep.2010.12.042
Ahmad H, Gubbels R, Ehlers E, Meyer F, Waterbury T, Lin R, Zhang L (2011) Kaposi sarcoma-associated herpesvirus degrades cellular Toll-interleukin-1 receptor domain-containing adaptor-inducing beta-interferon (TRIF). J Biol Chem 286:7865–7872
Newman RM, Salunkhe P, Godzik A, Reed JC (2006) Identification and characterization of a novel bacterial virulence factor that shares homology with mammalian Toll/interleukin-1 receptor family proteins. Infect Immun 74:594–601
Low LY, Mukasa T, Reed JC, Pascual J (2007) Characterization of a TIR-like protein from Paracoccus denitrificans. Biochem Biophys Res Comm 356:481–486
Cirl C, Wieser A, Yadav M, Duerr S, Schubert S, Fischer H, Stappert D, Wantia N, Rodriguez N, Wagner H et al (2008) Subversion of Toll-like receptor signaling by a unique family of bacterial Toll/interleukin-1 receptor domain-containing proteins. Nat Med 14:399–406
Zhang Q, Zmasek CM, Cai X, Godzik A (2011) TIR domain-containing adaptor SARM is a late addition to the ongoing microbe–host dialog. Develop Comp Immunol 35:461–468
Radhakrishnan GK, Yu Q, Harms JS, Splitter GA (2009) Brucella TIR domain-containing protein mimics properties of the Toll-like receptor adaptor protein TIRAP. J Biol Chem 284:9892–9898
Yadav M, Zhang J, Fischer H, Huang W, Lutay N, Cirl C, Lum J, Miethke T, Svanborg C (2010) Inhibition of TIR domain signaling by TcpC: MyD88-dependent and independent effects on Escherichia coli virulence. PLoS Pathog 6:e1001120
Sengupta D, Koblansky A, Gaines J, Brown T, West AP, Zhang D, Nishikawa T, Park S-G, Roop RM, Ghosh S (2010) Subversion of innate immune responses by Brucella through the targeted degradation of the TLR signaling adapter, MAL. J Immunol 184:956–964
Salcedo SP, Marchesini MI, Lelouard H, Fugier E, Jolly G, Balor S, Muller A, Lapaque N, Demaria O, Alexopoulou L et al (2008) Brucella control of dendritic cell maturation is dependent on the TIR-containing protein Btp1. PLoS Pathog 4:e21
Spear AM, Rana R, Jenner DC, Flick-Smith HC, Oyston PCF, Simpson P, Matthews S, Byrne B, Atkins HS (2012) A TIR domain protein from Yersinia pestis interacts with mammalian IL-1/TLR pathways but does not play a central role in the virulence of Y. pestis in a mouse model of bubonic plague. Microbiology 158:1593–1606
Zhang Y, Ting AT, Marcu KB, Bliska JB (2005) Inhibition of MAPK and NF-kappa B pathways is necessary for rapid apoptosis in macrophages infected with Yersinia. J Immunol 174:7939–7949
Chan SL, Low LY, Hsu S, Li S, Liu T, Santelli E, Le Negrate G, Reed JC, Woods VL, Pascual J (2009) Molecular mimicry in innate immunity: crystal structure of a bacterial TIR domain. J Biol Chem 284:21386–21392
Radhakrishnan GK, Splitter GA (2010) Biochemical and functional analysis of TIR domain containing protein from Brucella melitensis. Biochem Biophys Res Comm 397:59–63
Dunne A, Ejdeback M, Ludidi PL, O’Neill LAJ, Gay NJ (2003) Structural complementarity of Toll/interleukin-1 receptor domains in Toll-like receptors and the adaptors Mal and MyD88. J Biol Chem 278:41443–41451
Rana RR, Simpson P, Zhang M, Jennions M, Ukegbu C, Spear AM, Alguel Y, Matthews SJ, Atkins HS, Byrne B (2011) Yersinia pestis TIR-domain protein forms dimers that interact with the human adaptor protein MyD88. Microb Pathog 51:89–95
Radhakrishnan G, Harms J, Splitter G (2011) Modulation of microtubule dynamics by a TIR domain protein from the intracellular pathogen, Brucella melitensis. Biochem J 439:79–83
Zheng Y, Lilo S, Brodsky IE, Zhang Y, Medzhitov R, Marcu KB, Bliska JB (2011) A Yersinia effector with enhanced inhibitory activity on the NF-κB pathway activates the NLRP3/ASC/caspase-1 inflammasome in macrophages. PLoS Pathog 7:e1002026
Beutler B (2004) Inferences, questions and possibilities in Toll-like receptor signalling. Nature 430:257–263
Ngo VN, Young RM, Schmitz R, Jhavar S, Xiao W, Lim K-H, Kohlhammer H, Xu W, Yang Y, Zhao H et al (2011) Oncogenically active MYD88 mutations in human lymphoma. Nature 470:115–119
Lysakova-Devine T, Keogh B, Harrington B, Nagpal K, Halle A, Golenbock DT, Monie T, Bowie AG (2010) Viral inhibitory peptide of TLR4, a peptide derived from vaccinia protein A46, specifically inhibits TLR4 by directly targeting MyD88 adaptor-like and TRIF-related adaptor molecule. J Immunol 185:4261–4271
Toshchakov VY, Fenton MJ, Vogel SN (2007) Cutting edge: differential inhibition of TLR signaling pathways by cell-permeable peptides representing BB loops of TLRs. J Immunol 178:2655–2660
Chan SL, Mukasa T, Santelli E, Low LY, Pascual J (2010) The crystal structure of a TIR domain from Arabidopsis thaliana reveals a conserved helical region unique to plants. Protein Sci 19:155–161
Bernoux M, Ve T, Williams S, Warren C, Hatters D, Valkov E, Zhang X et al (2011) Structural and functional analysis of a plant resistance protein TIR domain reveals interfaces for self-association, signaling, and autoregulation. Cell Host Microbe 9:200–211
Acknowledgments
This work was funded by the UK Ministry of Defence.
Author information
Authors and Affiliations
Corresponding author
Additional information
Crown Copyright 2011. Published with the permission of the Defence Science and Technology Laboratory on behalf of the Controller of HMSO.
Rights and permissions
About this article
Cite this article
Rana, R.R., Zhang, M., Spear, A.M. et al. Bacterial TIR-containing proteins and host innate immune system evasion. Med Microbiol Immunol 202, 1–10 (2013). https://doi.org/10.1007/s00430-012-0253-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00430-012-0253-2