Abstract
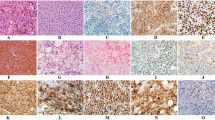
Plasmablastic myeloma (PBM) is a blastic morphologic variant of plasma cell myeloma with less favorable prognosis than those with non-blastic morphology. PBM is rare, without clear-cut definition and detailed clinicopathologic features in the literature. PBM may mimic plasmablastic lymphoma (PBL) as they share nearly identical morphology and immunophenotype. Using the criteria of ≥ 30% plasmablasts in tissue sections, we retrospectively recruited PBM cases and analyzed their clinical, imaging, and pathologic findings, with emphasis on extramedullary involvement. We performed immunohistochemistry, in situ hybridization for Epstein-Barr virus (EBER), and fluorescence in situ hybridization (FISH) for lymphoma- and myeloma-associated genetic alterations. Of the 25 recruited cases, 15 (60%) had extramedullary involvement, which occurred as initial presentation in nine cases. The most common extramedullary sites were soft tissue and/or skin (10/15, 67%), followed by pleural effusion, the lungs, and lymph nodes. Immunohistochemically, tumor cells expressed MYC (74%; 17/23), CD56 (56%; 14/25), and cyclin D1 (16%; 4/25), while CD117 was all negative (n = 25). Of the 20 cases stained with p53, four (20%) cases were diffusely positive, and the remaining 16 cases showed a heterogeneous pattern. EBER was negative in all 24 cases examined. Of the 13 cases examined with FISH, the genetic aberrations identified included del(13q14)(92%; 12/13), gain of chromosome 1q (90%; 9/10), loss of chromosome 1p (60%; 6/10), IGH-FGFR3 reciprocal translocation (23%; 3/13), rearranged MYC (15%; 2/13), and rearranged CCND1 (8%; 1/13), while there were no cases with TP53 deletion (n = 10) or rearrangement of BCL2 (n = 13) or BCL6 (n = 13). The prognosis was dismal regardless of the presence or absence of extramedullary involvement. In conclusion, PBM in Taiwan frequently presented as extramedullary and extranodal lesions, particularly in soft tissue and/or skin, mimicking PBL. FISH for targeted genetic alterations such as del(13q14), gain of chromosome 1q, loss of chromosome 1p, and IGH-FGFR3 might be helpful for the differential diagnoses. Larger studies are warranted to investigate the genetic alterations between PBM and PBL.



Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this article and its supplementary information file.
Code availability
Not applicable.
References
McKenna RW, Kyle RA, Kuehl WM, Harris NL, Coupland RW, Fend F (2017) Plasma cell neoplasms. In: Swerdlow SH, Campo E, Harris NL, Jaffe ES, Pileri SA, Stein H, Thiele J (eds) WHO classification of tumours of haematopoietic and lymphoid tissues revised, 4th edn. International Agency for Research on Cancer (IARC) 69372 Lyon Cedex 08, France, pp 241–258
Rajkumar SV, Dimopoulos MA, Palumbo A, Blade J, Merlini G, Mateos MV et al (2014) International Myeloma Working Group updated criteria for the diagnosis of multiple myeloma. Lancet Oncol 15(12):e538-548. https://doi.org/10.1016/S1470-2045(14)70442-5
Wutke K, Varbiro M, Rudiger KD, Kelenyi G (1981) Cytological and histological classification of multiple myeloma. Haematologia (Budap) 14(3):315–329
Greipp PR, Raymond NM, Kyle RA, O’Fallon WM (1985) Multiple myeloma: significance of plasmablastic subtype in morphological classification. Blood 65(2):305–310
Bartl R, Frisch B, Fateh-Moghadam A, Kettner G, Jaeger K, Sommerfeld W (1987) Histologic classification and staging of multiple myeloma. A retrospective and prospective study of 674 cases. Am J Clin Pathol 87(3):342–355. https://doi.org/10.1093/ajcp/87.3.342
Ayeboa-Sallah B, Qutab S, Grace R, Sharma N (2021) Rare case of plasmablastic myeloma diagnosed on lung biopsy. BMJ Case Rep 14(3):e240998. https://doi.org/10.1136/bcr-2020-240998
Suarez-Londono JA, Rohatgi A, Antoine-Pepeljugoski C, Braunstein MJ (2020) Aggressive presentation of plasmablastic myeloma. BMJ Case Rep 13(4):e234436. https://doi.org/10.1136/bcr-2020-234436
Liu Y, Jelloul F, Zhang Y, Bhavsar T, Ho C, Rao M et al (2020) Genetic basis of extramedullary plasmablastic transformation of multiple myeloma. Am J Surg Pathol 44(6):838–848. https://doi.org/10.1097/PAS.0000000000001459
Licci S (2017) Duodenal localization of plasmablastic myeloma. World J Gastrointest Pathophysiol 8(2):93–95. https://doi.org/10.4291/wjgp.v8.i2.93
Marks E, Shi Y, Wang Y (2017) CD117 (KIT) is a useful marker in the diagnosis of plasmablastic plasma cell myeloma. Histopathology 71(1):81–88. https://doi.org/10.1111/his.13196
Moller HE, Preiss BS, Pedersen P, Kristensen IB, Hansen CT, Frederiksen M et al (2015) Clinicopathological features of plasmablastic multiple myeloma: a population-based cohort. APMIS 123(8):652–658. https://doi.org/10.1111/apm.12411
Geyer JT, Niesvizky R, Jayabalan DS, Mathew S, Subramaniyam S, Geyer AI et al (2014) IgG4 plasma cell myeloma: new insights into the pathogenesis of IgG4-related disease. Mod Pathol 27(3):375–381. https://doi.org/10.1038/modpathol.2013.159
Bohn OL, Hsu K, Hyman DM, Pignataro DS, Giralt S, Teruya-Feldstein J (2014) BRAF V600E mutation and clonal evolution in a patient with relapsed refractory myeloma with plasmablastic differentiation. Clin Lymphoma Myeloma Leuk 14(2):e65-68. https://doi.org/10.1016/j.clml.2013.12.003
Chang ST, Liao YL, Lu CL, Chuang SS, Li CY (2007) Plasmablastic cytomorphologic features in plasma cell neoplasms in immunocompetent patients are significantly associated with EBV. Am J Clin Pathol 128(2):339–344. https://doi.org/10.1309/27H8XJH31F3GUNAT
Greipp PR, Leong T, Bennett JM, Gaillard JP, Klein B, Stewart JA et al (1998) Plasmablastic morphology–an independent prognostic factor with clinical and laboratory correlates: Eastern Cooperative Oncology Group (ECOG) myeloma trial E9486 report by the ECOG Myeloma Laboratory Group. Blood 91(7):2501–2507
Vega F, Chang CC, Medeiros LJ, Udden MM, Cho-Vega JH, Lau CC et al (2005) Plasmablastic lymphomas and plasmablastic plasma cell myelomas have nearly identical immunophenotypic profiles. Mod Pathol 18(6):806–815. https://doi.org/10.1038/modpathol.3800355
Sakakibara A, Kohno K, Eladl AE, Klaisuwan T, Ishikawa E, Suzuki Y et al (2018) Immunohistochemical assessment of the diagnostic utility of PD-L1: a preliminary analysis of anti-PD-L1 antibody (SP142) for lymphoproliferative diseases with tumour and non-malignant Hodgkin-Reed-Sternberg (HRS)-like cells. Histopathology 72(7):1156–1163. https://doi.org/10.1111/his.13475
Campo E, Stein H, Harris NL (2017) Plasmablastic lymphoma. In: Swerdlow SH, Campo E, Harris NL, Jaffe ES, Pileri SA, Stein H, Thiele J (eds) WHO classification of tumours of haematopoietic and lymphoid tissues revised, 4th edn. International Agency for Research on Cancer (IARC) 69372 Lyon Cedex 08, France, pp 321–322
Delecluse HJ, Anagnostopoulos I, Dallenbach F, Hummel M, Marafioti T, Schneider U et al (1997) Plasmablastic lymphomas of the oral cavity: a new entity associated with the human immunodeficiency virus infection. Blood 89(4):1413–1420
Morscio J, Dierickx D, Nijs J, Verhoef G, Bittoun E, Vanoeteren X et al (2014) Clinicopathologic comparison of plasmablastic lymphoma in HIV-positive, immunocompetent, and posttransplant patients: single-center series of 25 cases and meta-analysis of 277 reported cases. Am J Surg Pathol 38(7):875–886. https://doi.org/10.1097/PAS.0000000000000234
Martinez D, Valera A, Perez NS, Sua Villegas LF, Gonzalez-Farre B, Sole C et al (2013) Plasmablastic transformation of low-grade B-cell lymphomas: report on 6 cases. Am J Surg Pathol 37(2):272–281. https://doi.org/10.1097/PAS.0b013e31826cb1d1
Teruya-Feldstein J, Chiao E, Filippa DA, Lin O, Comenzo R, Coleman M et al (2004) CD20-negative large-cell lymphoma with plasmablastic features: a clinically heterogenous spectrum in both HIV-positive and -negative patients. Ann Oncol 15(11):1673–1679. https://doi.org/10.1093/annonc/mdh399
Colomo L, Loong F, Rives S, Pittaluga S, Martinez A, Lopez-Guillermo A et al (2004) Diffuse large B-cell lymphomas with plasmablastic differentiation represent a heterogeneous group of disease entities. Am J Surg Pathol 28(6):736–747. https://doi.org/10.1097/01.pas.0000126781.87158.e3
Chen BJ, Chuang SS (2020) Lymphoid neoplasms with plasmablastic differentiation: a comprehensive review and diagnostic approaches. Adv Anat Pathol 27(2):61–74. https://doi.org/10.1097/PAP.0000000000000253
Simonitsch-Klupp I, Hauser I, Ott G, Drach J, Ackermann J, Kaufmann J et al (2004) Diffuse large B-cell lymphomas with plasmablastic/plasmacytoid features are associated with TP53 deletions and poor clinical outcome. Leukemia 18(1):146–155. https://doi.org/10.1038/sj.leu.2403206
Sailer M, Vykoupil KF, Peest D, Coldewey R, Deicher H, Georgii A (1995) Prognostic relevance of a histologic classification system applied in bone marrow biopsies from patients with multiple myeloma: a histopathological evaluation of biopsies from 153 untreated patients. Eur J Haematol 54(3):137–146. https://doi.org/10.1111/j.1600-0609.1995.tb00204.x
Köbel M, Piskorz AM, Lee S, Lui S, LePage C, Marass F et al (2016) Optimized p53 immunohistochemistry is an accurate predictor of TP53 mutation in ovarian carcinoma. J Pathol Clin Res 2(4):247–258. https://doi.org/10.1002/cjp2.53
Herbst RS, Baas P, Kim DW, Felip E, Perez-Gracia JL, Han JY et al (2016) Pembrolizumab versus docetaxel for previously treated, PD-L1-positive, advanced non-small-cell lung cancer (KEYNOTE-010): a randomised controlled trial. Lancet 387(10027):1540–1550. https://doi.org/10.1016/S0140-6736(15)01281-7
Herbst RS, Garon EB, Kim DW, Cho BC, Gervais R, Perez-Gracia JL et al (2021) Five year survival update from KEYNOTE-010: pembrolizumab versus docetaxel for previously treated, programmed death-ligand 1-positive advanced NSCLC. J Thorac Oncol 16(10):1718–1732. https://doi.org/10.1016/j.jtho.2021.05.001
Liu CY, Chuang SS (2021) A simple and practical guide for triaging lymphocyte-rich effusions for ancillary studies. Adv Anat Pathol 28(2):94–104. https://doi.org/10.1097/PAP.0000000000000290
Chang ST, Hsieh YC, Kuo CC, Chuang SS (2018) Colonic CD30 positive plasmablastic plasmacytoma masquerading as anaplastic large cell lymphoma. Pathology 50(6):668–670. https://doi.org/10.1016/j.pathol.2018.03.011
Fernandez de Larrea C, Kyle RA, Durie BG, Ludwig H, Usmani S, Vesole DH et al (2013) Plasma cell leukemia: consensus statement on diagnostic requirements, response criteria and treatment recommendations by the International Myeloma Working Group. Leukemia 27(4):780–791. https://doi.org/10.1038/leu.2012.336
Valera A, Balague O, Colomo L, Martinez A, Delabie J, Taddesse-Heath L et al (2010) IG/MYC rearrangements are the main cytogenetic alteration in plasmablastic lymphomas. Am J Surg Pathol 34(11):1686–1694. https://doi.org/10.1097/PAS.0b013e3181f3e29f
Taddesse-Heath L, Meloni-Ehrig A, Scheerle J, Kelly JC, Jaffe ES (2010) Plasmablastic lymphoma with MYC translocation: evidence for a common pathway in the generation of plasmablastic features. Mod Pathol 23(7):991–999. https://doi.org/10.1038/modpathol.2010.72
Montes-Moreno S, Martinez-Magunacelaya N, Zecchini-Barrese T, Villambrosia SG, Linares E, Ranchal T et al (2017) Plasmablastic lymphoma phenotype is determined by genetic alterations in MYC and PRDM1. Mod Pathol 30(1):85–94. https://doi.org/10.1038/modpathol.2016.162
Liu Z, Filip I, Gomez K, Engelbrecht D, Meer S, Lalloo PN et al (2020) Genomic characterization of HIV-associated plasmablastic lymphoma identifies pervasive mutations in the JAK-STAT pathway. Blood Cancer Discov 1(1):112–125. https://doi.org/10.1158/2643-3230.BCD-20-0051
Ramis-Zaldivar JE, Gonzalez-Farre B, Nicolae A, Pack S, Clot G, Nadeu F et al (2021) MAPK and JAK-STAT pathways dysregulation in plasmablastic lymphoma. Haematologica 106(10):2682–2693. https://doi.org/10.3324/haematol.2020.271957
Garcia-Reyero J, Martinez Magunacelaya N, Gonzalez de Villambrosia S, Loghavi S, Gomez Mediavilla A, Tonda R et al (2021) Genetic lesions in MYC and STAT3 drive oncogenic transcription factor overexpression in plasmablastic lymphoma. Haematologica 106(4):1120–1128. https://doi.org/10.3324/haematol.2020.251579
Avet-Loiseau H, Attal M, Moreau P, Charbonnel C, Garban F, Hulin C et al (2007) Genetic abnormalities and survival in multiple myeloma: the experience of the Intergroupe Francophone du Myelome. Blood 109(8):3489–3495. https://doi.org/10.1182/blood-2006-08-040410
Cook JR, Hsi ED, Worley S, Tubbs RR, Hussein M (2006) Immunohistochemical analysis identifies two cyclin D1+ subsets of plasma cell myeloma, each associated with favorable survival. Am J Clin Pathol 125(4):615–624. https://doi.org/10.1309/BDR9-59TT-4JU6-388C
Loghavi S, Khoury JD, Medeiros LJ (2015) Epstein-Barr virus-positive plasmacytoma in immunocompetent patients. Histopathology 67(2):225–234. https://doi.org/10.1111/his.12640
Funding
This study was funded by Taipei Medical University-Shuang Ho Hospital, grant no. 109TMU-SHH-15, and the Ministry of Science and Technology, Taiwan (MOST 108–2314-B-384–004).
Author information
Authors and Affiliations
Contributions
HCC and SSC initiated the study; BJC, CTU, CFY, CHH, YKL, and SSC analyzed the data; YZS did the experiments; BJC and SSC wrote the manuscript; HCC and SSC revised the manuscript; all authors approved the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This study was approved by the Internal Review Board of Chi-Mei Medical Center, Tainan, Taiwan.
Consent to participate
Not applicable.
Consent for publication
The consent for publication will be provided after acceptance.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Hsiu-Chu Chou and Shih-Sung Chuang contributed equally, and both are considered corresponding authors.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, BJ., Yuan, CT., Yang, CF. et al. Plasmablastic myeloma in Taiwan frequently presents with extramedullary and extranodal mass mimicking plasmablastic lymphoma. Virchows Arch 481, 283–293 (2022). https://doi.org/10.1007/s00428-022-03342-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-022-03342-3