Abstract
Inverted (Schneiderian) sinonasal papilloma (ISP) is a neoplasm derived from mucosa of the sinonasal tract characterized by local aggressive growth, a tendency to recur and an association with sinonasal carcinoma. The etiology of ISP remains unclear. Recently, identical mutations in exons 19 and 20 of the oncogene EGFR were reported in ISP and ISP-associated sinonasal carcinoma. Nevertheless, it remains unclear whether recurring ISPs show identical EGFR mutations at different time points or whether these mutations are identical throughout the respective ISP sample. We used Sanger sequencing to test 60 formalin-fixed paraffin embedded ISP samples from 40 patients regarding mutations in exons 19 and 20 of EGFR—together with exon 15 of BRAF. Overall, 32 samples of 22 patients showed a mutation in EGFR exon 20, whereas 28 samples of 18 patients showed none. No mutation in EGFR exon 19 was found in any sample. Four samples of four patients showed a BRAF exon 15 mutation. Interestingly, samples of four patients exhibited genetic heterogeneity, enabling us to report this in ISP for the first time.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Inverted sinonasal papilloma (ISP) is a neoplastic proliferation arising from the sinonasal tract mucosa with predominantly inverted growth [1]. ISPs are mostly located in the nasal cavity and the maxillary sinus [2]. They often recur and may grow locally destructive [3]. The association with carcinoma is stronger for ISP than for exophytic papillomas [1]. In about 1.9 to 27% of ISPs, sinonasal squamous cell carcinomas (SSCCs) are found, the majority being synchronous tumors [4]. The etiology of ISP is virtually unknown. Exposure to organic solvents has been reported as a risk factor, and the importance of HPV-infection has been discussed, albeit controversially [2, 4].
The epidermal growth factor receptor (EGFR) gene is a prominent oncogene in multiple tumor entities including adenocarcinoma of the lung [5]. Here, specific mutations within exons 18–21, coding for the tyrosine kinase (TK) domain, lead to constitutive activation and thus to aberrant signaling [6]. A subset of patients with tumors showing corresponding mutations might benefit from targeted therapy with TK inhibitors [6].
Recently, EGFR mutations in exons 19 and 20 have been found in ISP and ISP-associated SSCC [3, 7] and their possible consequences for therapy were discussed [8]. Other driver mutations commonly found in other entities seem to be very rare [9, 10].
Nevertheless, BRAF mutations have been shown to occur in several pre-malignant lesions like serrated adenomas of the colon [11], atypical adenomatous hyperplasia in the lung [12], endometrial hyperplasia [13], or benign melanocytic nevi of the skin [14] and the conjunctiva [15]. Therefore, we included the analysis of BRAF mutations in our study.
We screened for mutations in EGFR exon 19 and 20 and BRAF exon 15 in 60 FFPE samples derived of a cohort of 40 ISP patients. For 15 patients, multiple samples were available. Material from four patients showed genetic heterogeneity.
Material and methods
Tumor samples
Formalin-fixed paraffin embedded (FFPE) tumor material from 40 patients diagnosed with ISP was obtained from the archive of the Institute of Pathology, University Medicine, Rostock. For 15 of these patients, multiple samples were available; so altogether 60 ISP samples were analyzed. Additionally, 8 samples from 5 SSCC patients were included.
The hematoxylin and eosin (H&E) sections were examined using a BX41 light microscope (Olympus, Hamburg, Germany) to check tissue sufficiency and quality. If tumor cell ratio did not reach at least 50%, tumor tissue was dissected from slide-mounted 10μm paraffin sections before DNA extraction using a stereoscopic microscope (Technival-2, Zeiss, Jena, Germany). To check the results of each microdissection, tissue remaining on the glass slide was stained with routine H&E stains and examined microscopically.
If tumor content fell below 50% and dissection as described above was not feasible, laser capture microdissection (LCM) was performed. Briefly, 5-μm sections were mounted on Membrane Slides (MembraneSlide 1.0 PEN, Zeiss, Göttingen, Germany) and stained weakly with H&E. LCM was performed with a Zeiss Observer Z1 microscope (Zeiss) combined with a PALM Micro Beam (Zeiss) and RoboMover (Zeiss), allowing precise selection of desired cells, using PALMRobo Software V4.5 (Zeiss).
Specimen collection was conducted in accordance with the ethical guidelines for the use of human material, approved by the Ethics Committee of the University of Rostock (Reference number: A2017-0159).
Mutation analysis
DNA was extracted from 10-μm sections of FFPE samples by deparaffinization and proteinase-K digestion (Roche, Mannheim, Germany), followed by purification with the Wizard DNA cleanup system (Promega, Mannheim, Germany) according to the manufacturer’s protocol.
For LCM obtained tissue fragments, DNA extraction was performed using the QIAamp DNA Micro Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol for isolation of genomic DNA from laser-microdissected tissues.
DNA content was determined by fluorescent quantification (Quantus, Promega). For PCR amplification of EGFR exons 19 and 20 and BRAF exon 15, 50 ng DNA was used with the following primers: EGFR-19F: 5′-tgccagttaacgtcttccttctctc-3′; EGFR-19R: 5′-ccacacagcaaagcagaaactcac-3′; EGFR-20F: 5′-ccaccatgcgaagccacactga-3′ and EGFR-20R: 5′-tccttatctcccctccccgtatctc-3′; BRAF-15F: 5′-tcataatgcttgctctgatagga-3′; BRAF-15R: 5′-ctttctagtaactcagcagc-3′. PCR was performed using the MyTaq HS polymerase (Bioline, Luckenwalde, Germany) applying the following conditions: 95 °C for 1 min, 35 cycles of 95 °C for 15 s, 61 °C (EGFR) or 60 °C (BRAF) for 15 s, and 72 °C for 10 s. PCR products were checked by agarose gel electrophoresis. Subsequently, 15 μl thereof were purified with 3 μl Fast AP alkaline phosphatase (Thermo Scientific, Dreieich, Germany) and 1.5 μl exonuclease I (Thermo Scientific) with incubation at 37 °C for 15 min and 85 °C for 15 min.
The purified PCR products were used as template for Sanger sequencing with the abovementioned primers and BigDye™ Terminator v1.1 Cycle Sequencing Kit (Applied Biosystems, Darmstadt, Germany), substituted with BDX64 sequencing enhancing buffer (Nimagen, Nijmegen, the Netherlands). Analyses were performed on an ABI 3500 genetic analyzer (Applied Biosystems) with SeqScape software 2.7 (Applied Biosystems).
Results
Clinical and histological data
ISP patients
Sixty FFPE samples from 40 patients were analyzed. For 25 patients, only single samples were available. In all, 35 samples were derived from 15 patients (up to four samples/patient): either taken at the same time point from different (yet adjacent) locations or from the same or adjacent locations at different time points (with time intervals of up to several years) in cases of recurrent disease.
Of the 40 patients, 24 were male and 16 were female; the median age at diagnosis was 59 years (SD ± 12.7 years, range 28–86 years). Detailed clinical information is given in Table 1.
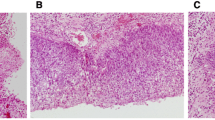
The morphology of ISP was identical in all samples and in accordance with WHO and AFIP descriptions [1, 4]. No special features could be observed in any case.
According to our current knowledge and available clinical data, none of these patients developed an ISP associated SSCC.
We observed no connection between clinical behavior (recurrence) and genetic phenotype (data given in Tables 1 and 2).
SSCC patients
For comparison, 8 samples from 5 SSCC patients were included; one diagnosed as ISP-related SSCC. One of the five SSCC patients was female, the remaining four male. The median age at diagnosis was 59 years (SD 17.4, range 48–83 years). The only patient with an ISP-associated SSCC was male, 59 years old. From this patient, three samples were available, all derived from the nasal cavity (left), at closely matching time points (after 6 or 14 days, respectively). From one the other SSCC patients, two samples were available.
EGFR and BRAF mutation analyses
DNA extraction and Sanger sequencing of EGFR exons 19 and 20 and BRAF exon 15 were successful for all samples. No exon 19 mutation could be detected in any sample.
In 18 patients (45%), no mutations were found in any samples analyzed, while 22 patients (55%) showed a mutation. Here, for 14 patients, only single samples were analyzed and for 8 patients multiple samples. Of these 8 patients, four showed the same mutation patterns in all samples analyzed whereas another four showed genetic heterogeneity. Mutation data overview is given in Table 2.
Most EGFR exon 20 mutations were duplications, insertions, and other complex mutations encompassing the region around amino acid positions 768 to 774. Moreover, few point mutations within this region or at position P794 occurred. All EGFR exon 20 mutations lay within the TK domain. An EGFR exon 20 mutation occurred in at least one sample of all 22 mutation-positive patients.
BRAF exon 15 mutations occurred in only four patients. Only one patient (heterogeneous) showed a V600E mutation (no EGFR mutation in this sample), whereas the other three showed mutations near the V600 hotspot, T589I, H608Y, and Q612*, each accompanying an EGFR mutation.
Of the SSCC patients, the one with ISP-associated SSCC showed an EGFR exon 20 mutation, N771delinsGY, in all three samples. All samples of the other four SSCC patients remained wild type. No BRAF mutation could be detected in any SSCC sample.
Genetic heterogeneity
Of eight mutation-positive patients with multiple samples available, four (#3, #13, #35, and #40) showed an identical mutation pattern, whereas four showed genetic heterogeneity (Fig. 1), described in detail below.
Illustration of genetic heterogeneity. a Patient #7 showing different mutations at the same time point at adjacent locations. b Patient #14 showing different mutations at the same time point at adjacent locations. c Patient #29 showing different mutations at the same location over time. d Patient #37 showing different mutations at the same location over time
Patient #7
Two samples were available, taken from adjacent locations (os ethmoidale posterior left and os ethmoidale left) at the same time point. One showed an EGFR H773dup, the second the same EGFR mutation and an additional BRAF T589I mutation (Fig. 1a).
Patient #14
Two samples were taken at the same time point from adjacent locations (meatus nasalis right and nasal cavity right). The first sample was wild type for both EGFR and BRAF, while the second harbored an EGFR P794L mutation (Fig. 1b).
Patient #29
Three samples were taken over a longer time period (up to 22 months) from the same location (sphenoid sinus right). Initially, an EGFR H773_V774dup was observed. This mutation could not be found in the second sample taken after 1 year and 7 months: both EGFR and BRAF showed a wild-type sequence. In the third sample, taken 3 months later, a BRAF V600E mutation occurred while EGFR remained wild type (Fig. 1c).
Patient #37
Four samples were taken from the same location (maxillary sinus right) over a time period of almost 9 years. The tumor initially showed a complex EGFR V774_C775insLM mutation.
Two months later, two point mutations in EGFR exon 20 occurred, H773L and V774M, while no insertions were found.
At the third time point (another 2 years later), these EGFR codons were wild type, but a P794L point mutation had occurred. Additionally, the tumor now harbored a BRAF H608Y mutation. Finally, at a fourth time point (another 6.5 years later), the EGFR point mutations H773L and V774M re-occurred, whereas the EGFR P794L and BRAF H608Y mutations could not be detected here (Fig. 1d).
In summary, this study is the first to document the occurrence of genetic heterogeneity in ISP.
Discussion
EGFR mutations
In this study, we analyzed 60 samples taken from 40 ISP patients and found EGFR mutations in tissue of more than half of the patients (22/40; 55%). All EGFR mutations detected lay in the coding region for the TK domain of EGFR [6]. The only case of ISP-associated SSCC showed a N771delinsGY mutation in EGFR exon 20. We could not find an EGFR mutation in any of the four cases of non-ISP-associated SSCC investigated.
In the literature, EGFR mutations have been hitherto demonstrated in 38 to 88% of ISPs [3, 16] and in 50 to 88% of ISP-associated SSCCs [10, 16]: Cabal et al. found EGFR exon 20 mutations in 38% (7/18) of ISPs and in 50% (6/12) of ISP-associated SSCCs, while EGFR exon 20 mutations in ISP-unassociated SSCC occurred in only 5.3% (1/19) [16]. Sasaki et al. reported EGFR mutation in 100% (9/9) of ISPs and 83% (10/12) of ISPs associated with SSCC, the vast majority being exon 20 insertions. EGFR exon 20 mutations were also found in 88% (15/17) of ISP-associated SSCC compared with only 14% (9/63) of non-ISP-associated SSCC. The EGFR mutation was identical in all 12 cases of analyzed ISPs and matched ISP-associated SSCCs [10]. In the work of Udager et al., EGFR mutations were detected in 88% of ISPs (44/50) and in 77% (17/22) of ISP-associated SSCCs. In 83% (10/12) of cases, the identical EGFR mutation could be identified in ISPs and matched ISP-associated SSCCs, the other two cases being EGFR wild type. Most of the EGFR mutations reported here were EGFR 20 insertions [3].
While the complex EGFR V774_C775_insLM is has not been described hitherto, the other complex duplications have been detected in ISP [3, 17]. The point mutations H773L and V774M have yet to be described in ISP, though detected in other tumor entities, including adenocarcinoma of the lung [18, 19] and the point mutation V774M additionally in sebaceous carcinoma of the skin [20], anaplastic astrocytoma grade III [21], and phyllodes tumors of the breast [22]. The P794L and P772L mutations have also not been reported in ISP, albeit respectively discovered in cases of sebaceous adenoma [20] and adenocarcinoma of the lung [23]. The EGFR N771delinsGY mutation has been reported in non-small cell lung cancer [24].
BRAF mutations
In four cases (10% of patients), we detected BRAF mutations. All detected BRAF mutations lay at or near the V600 hotspot. To the best of our knowledge, BRAF mutations have not been detected in ISP before.
Of the four samples identified with BRAF mutations, only one harbored a V600E mutation—well known as oncogenic driver mutation, e.g., in melanocytic tumors [25]. The three remaining samples harbored mutations at different, yet nearby loci. All these mutations have been detected but rarely in other neoplasms: H608Y in single cases of colorectal carcinoma [26, 27] and papillary thyroid carcinoma [28], T589I in a case of colon adenoma [29] and in a case of squamous cell carcinoma of the lung [30], and Q612* in a case of colon carcinoma [31] and in a case of thyroid anaplastic carcinoma [32].
Genetic heterogeneity
In ISP samples from four patients, we found genetic heterogeneity: different genetic phenotypes could be detected in multiple samples derived of the same patient, either in samples taken at the same time from different, yet adjacent locations or derived of the same location at different time points.
Intratumoral genetic heterogeneity can develop in different ways: one implies the field cancerization theory [33] which states that different tumors can develop independently from a field of genetically altered pre-neoplastic cells [34]. This mechanism of tumorigenesis has been invoked in carcinoma of the oral cavity [34, 35] or other head and neck cancer [36]. Here, the causative agents might be tobacco smoke or alcohol and similar substances that can be inhaled or swallowed.
In the oral field cancerization model (field effect), the field of precancerous cells might either be derived from cells transformed by multiple events, e.g., after contact with carcinogenic substances (polyclonal field; see Fig. 2a), or by daughter cells of a single genetically altered stem cell which migrate via saliva or in the epithelium (monoclonal field) [34].
Possible causes of intratumoral genetic heterogeneity. a Field cancerization: exposure to noxious agents leads to several mutations (polygons) resulting in different tumor initiating cells (white, gray, and black single circles) causing a tumor mass consisting of different subclones (white, gray, and black bulks). b Linear evolution: in the original tumor (white) additional mutational events (polygons) leads to new subclones (at first gray, later black) which show, over time, growth advantages over the original tumor cells. c Branched evolution: a tumor (white) is hit by different mutational events (polygons) each leading to new subclones (gray and black), which simultaneously develop growth advantages in distinct areas
Tumorigenesis then proceeds in three phases: (1) patch formation, when the progeny of a single transformed stem cell forms a clonal proliferation; (2) clonal expansion, when the normal epithelium is replaced by genetically altered cells due to growth advantages; and (3) transition to tumor, when additional genetic alterations transform the clone into an overt carcinoma with features of invasive and metastatic potential [34]. Indeed, some have applied the theory of field cancerization not only to oral cavity, esophagus, and colon but also to skin, bladder, or vulva [37].
Clonal relationships might give important hints to the way multiple lesions have developed: if multiple lesions share common genetic alterations, they might be derived from the same progenitor cell. On the other hand, if a clonal relationship between multiple lesions is absent, they might derive from independently transformed cells [34].
Another possible cause of intratumoral heterogeneity might be tumor evolution.
In linear evolution, specific subclones may accumulate mutations which offer survival benefits. This leads to outgrowth of distinct clones and therefore temporal heterogeneity (Fig. 2b).
In branched evolution (Fig. 2c), different subclones with distinct mutation patterns exist within a tumor and may outcompete each other leading to genetic heterogeneity in samples from the same time point (reviewed in [38]).
For two of the patients showing genetic heterogeneity (#7 and #14), paired samples were taken each at the same time point from adjacent regions:
In patient #7, an EGFR H773dup was present in both samples, whereas in only one, a BRAF T589I mutation was detected. Therefore, both subclones existed in this tumor simultaneously (Fig. 1a).
In patient #14, one sample was wild type for both genes; the other harbored an EGFR P794L mutation. Again, both subclones existed simultaneously (Fig. 1b).
So, generally speaking, all three models of intratumoral genetic heterogeneity could apply here. However, in patient #7, due to the EGFR H773dup mutation shared by both subclones at the same time point, branched evolution seems more likely.
For patients #29 and #37, multiple samples were taken over longer time courses, of up to 22 months and 9 years, respectively:
Accordingly, ISP tissue in patient #29 initially showed an EGFR H773_V774dup mutation, which could not be detected after 19 months; so the second sample was wild type for both EGFR and BRAF. The third sample, taken 3 months later, showed a BRAF V600E mutation but no EGFR mutation (Fig. 1c).
In patient #37 (Fig. 1d), the situation was even more complex: an EGFR V774_C775insLM mutation was found in the first sample. Two months later, no insertions were observed, but instead, two point mutations in EGFR exon 20, H773L and V774M, occurred. These point mutations could not be detected at the third time point (a further 2 years later), but a P794L point mutation was present. Additionally, the tumor now harbored a de novo BRAF H608Y mutation. The EGFR point mutations H773L and V774M recurred 6.5 years later, whereas EGFR P794L and BRAF H608Y mutations could not be detected (Fig. 1d).
Tumor evolution might be a possible explanation for the loss of mutations in patients #29 and #37. In non-small cell lung cancer (NSCLC), a “loss” of EGFR mutation has been reported under TK inhibitor therapy. This was interpreted as a response to therapeutic pressure, where EGFR wild-type clones outgrew the mutant ones [39].
However, it remains unclear why ISP wild-type clones should outgrow subclones harboring a potentially activating EGFR or BRAF mutation.
The disappearance of these mutations might therefore raise the question whether these really offer a survival advantage for distinct subclones and, accordingly, if these mutations have clinical significance. However, since EGFR mutations have been reported in 38 to 88% of ISP by various groups (see above), random coincidence does not seem a likely explanation.
On the other hand, EGFR mutations occur in only a fraction of ISPs and ISP-associated SSCCs. So one might speculate that either another genetic alteration might be the original transforming driving event or that EGFR mutations and another type of driver mutation are mutually exclusive events.
Genetic heterogeneity has been increasingly identified in a variety of cancers where it may serve to promote tumor survival in response to therapy [40, 41].
The occurrence of genetic heterogeneity in benign neoplasms and precancerous lesions has received less attention. Maley et al. [42] showed that clonal diversity occurs in Barrett’s esophagus and, moreover, is linked to the risk of progression to esophageal adenocarcinoma. Others have reported subclones with different driver mutations even in small colonic adenomas [43].
Until recently, little genetic research has been performed on ISP [2]. In 2000, Califano et al. [44] investigated the random X chromosome inactivation patterns in ISP in four female patients and concluded that ISP is a monoclonal lesion. However, in one case of ISP being associated with SSCC, the inactivated X-chromosomal allele was different in the ISP and the carcinoma investigated.
Otherwise, Udager et al. [3] could demonstrate the same EGFR mutations in ISP and associated SSCCs in most cases and Sasaki et al. [10] in all cases analyzed.
Yakusawa et al. have investigated the genetic variants of ISP and associated SSCC in the same patients. Interestingly, they found differences: they detected one mutation in ISP alone (GNAQ) and two mutations in SSCC alone (MSH6, PIK3CA), with the majority of mutations being identical. They did not compare different samples of ISP or ISP-associated SSCC derived from the same patient [9].
The development of ISP has been linked with exposure to organic solvents and other occupational factors [45, 46], and the lesion may arise multifocally [1]. This may support the idea of a field effect in the genesis of ISP (de novo genesis of ISP)—as well as some examples of genetic heterogeneity in ISP presented here.
The question whether multiple foci of ISP or ISP-associated SSCC might share the same origin or arise independently is not purely theoretical: Udager demonstrated the sensitivity of cell lines established from ISP-associated SSCC with EGFR mutations to some of the new irreversible TK inhibitors [8]. Moreover, poziotinib, a third-generation EGFR inhibitor, showed initially promising results in an ongoing phase II trial of non-small cell lung cancer patients with EGFR exon 20 insertions [47]. In this setting, it might very well matter whether subclones with different EGFR mutations arise in ISP and/or in ISP-associated SSCC.
In summary, our results add to the body of knowledge regarding EGFR and BRAF mutations in ISP and show that the genetic landscape of ISP might be more complex than hitherto anticipated. Moreover, this is the first study to address testing for genetic heterogeneity in ISP samples derived from the same patient. Additionally, some of the data might be taken to support the idea of the cancer field effect in ISP. Finally, this is the first study to demonstrate BRAF mutations in ISP.
Change history
19 July 2021
A Correction to this paper has been published: https://doi.org/10.1007/s00428-021-03150-1
References
Mills SE, Stelow EB, Hunt JL (2012) Inverted Schneiderian papillomas. In: Mills SE, Stelow EB, Hunt JL (eds) Tumors of the upper aerodigestive tract and ear. ARP Press, Silver Spring, pp 23–25
Re M, Gioacchini FM, Bajraktari A, Tomasetti M, Kaleci S, Rubini C, Bertini A, Magliulo G, Pasquini E (2017) Malignant transformation of sinonasal inverted papilloma and related genetic alterations: a systematic review. Eur Arch Otorhinolaryngol 274:2991–3000. https://doi.org/10.1007/s00405-017-4571-2
Udager AM, Rolland DCM, McHugh JB, Betz BL, Murga-Zamalloa C, Carey TE et al (2015) High-frequency targetable EGFR mutations in sinonasal squamous cell carcinomas arising from inverted sinonasal papilloma. Cancer Res 75:2600–2606. https://doi.org/10.1158/0008-5472.CAN-15-0340
Hunt JL, Bell D, Sarioglu S (2017) Sinonasal papilloma, inverted type. In: El-Naggar AK, Chan JKC, Grandis JR et al (eds) WHO classification of head and neck tumours, 4th edn. International Agency for Research on Cancer, Lyon, pp 26–29
Kumar V, Abbas AK, Aster JC (2015) Robbins and Cotran pathologic basis of disease, 9th. Student Consult. Elsevier Saunders, Philadelphia
Sharma SV, Bell DW, Settleman J, Haber DA (2007) Epidermal growth factor receptor mutations in lung cancer. Nat Rev Cancer 7:169–181. https://doi.org/10.1038/nrc2088
Sahnane N, Ottini G, Turri-Zanoni M, Furlan D, Battaglia P, Karligkiotis A, Albeni C, Cerutti R, Mura E, Chiaravalli AM, Castelnuovo P, Sessa F, Facco C (2019) Comprehensive analysis of HPV infection, EGFR exon 20 mutations and LINE1 hypomethylation as risk factors for malignant transformation of sinonasal-inverted papilloma to squamous cell carcinoma. Int J Cancer 144:1313–1320. https://doi.org/10.1002/ijc.31971
Udager AM, McHugh JB, Elenitoba-Johnson KSJ, Brown NA (2015) EGFR mutations in sinonasal squamous tumors: oncogenic and therapeutic implications. Oncoscience 2:908–909. https://doi.org/10.18632/oncoscience.268
Yasukawa S, Kano S, Hatakeyama H, Nakamaru Y, Takagi D, Mizumachi T, Suzuki M, Suzuki T, Nakazono A, Tanaka S, Nishihara H, Homma A (2018) Genetic mutation analysis of the malignant transformation of sinonasal inverted papilloma by targeted amplicon sequencing. Int J Clin Oncol 23:835–843. https://doi.org/10.1007/s10147-018-1296-1
Sasaki E, Nishikawa D, Hanai N, Hasegawa Y, Yatabe Y (2018) Sinonasal squamous cell carcinoma and EGFR mutations: a molecular footprint of a benign lesion. Histopathology 73:953–962. https://doi.org/10.1111/his.13732
Kim SY, Kim TI (2018) Serrated neoplasia pathway as an alternative route of colorectal cancer carcinogenesis. Intest Res 16:358–365. https://doi.org/10.5217/ir.2018.16.3.358
Sivakumar S, Lucas FAS, McDowell TL, Lang W, Xu L, Fujimoto J et al (2017) Genomic landscape of atypical adenomatous hyperplasia reveals divergent modes to lung adenocarcinoma. Cancer Res 77:6119–6130. https://doi.org/10.1158/0008-5472.CAN-17-1605
Feng Y-Z, Shiozawa T, Miyamoto T, Kashima H, Kurai M, Suzuki A, Konishi I (2005) BRAF mutation in endometrial carcinoma and hyperplasia: correlation with KRAS and p53 mutations and mismatch repair protein expression. Clin Cancer Res 11:6133–6138. https://doi.org/10.1158/1078-0432.CCR-04-2670
Price HN (2016) Congenital melanocytic nevi: update in genetics and management. Curr Opin Pediatr 28:476–482. https://doi.org/10.1097/MOP.0000000000000384
Larsen A-C, Dahl C, Dahmcke CM, Lade-Keller J, Siersma VD, Toft PB, Coupland SE, Prause JU, Guldberg P, Heegaard S (2016) BRAF mutations in conjunctival melanoma: investigation of incidence, clinicopathological features, prognosis and paired premalignant lesions. Acta Ophthalmol 94:463–470. https://doi.org/10.1111/aos.13007
Cabal VN, Menendez M, Vivanco B, Potes-Ares S, Riobello C, Suarez-Fernandez L, Garcia-Marin R, Blanco-Lorenzo V, Lopez F, Alvarez-Marcos C, Llorente JL, Hermsen MA (2020) EGFR mutation and HPV infection in sinonasal inverted papilloma and squamous cell carcinoma. Rhinology 0:0. https://doi.org/10.4193/Rhin19.371
Udager AM, McHugh JB, Goudsmit CM, Weigelin HC, Lim MS, Elenitoba-Johnson KSJ et al (2018) Human papillomavirus (HPV) and somatic EGFR mutations are essential, mutually exclusive oncogenic mechanisms for inverted sinonasal papillomas and associated sinonasal squamous cell carcinomas. Ann Oncol 29:466–471. https://doi.org/10.1093/annonc/mdx736
Yang SH, Mechanic LE, Yang P, Landi MT, Bowman ED, Wampfler J, Meerzaman D, Hong KM, Mann F, Dracheva T, Fukuoka J, Travis W, Caporaso NE, Harris CC, Jen J (2005) Mutations in the tyrosine kinase domain of the epidermal growth factor receptor in non-small cell lung cancer. Clin Cancer Res 11:2106–2110. https://doi.org/10.1158/1078-0432.CCR-04-1853
Tuononen K, Mäki-Nevala S, Sarhadi VK, Wirtanen A, Rönty M, Salmenkivi K, Andrews JM, Telaranta-Keerie AI, Hannula S, Lagström S, Ellonen P, Knuuttila A, Knuutila S (2013) Comparison of targeted next-generation sequencing (NGS) and real-time PCR in the detection of EGFR, KRAS, and BRAF mutations on formalin-fixed, paraffin-embedded tumor material of non-small cell lung carcinoma-superiority of NGS. Genes Chromosom Cancer 52:503–511. https://doi.org/10.1002/gcc.22047
Harvey NT, Tabone T, Erber W, Wood BA (2016) Circumscribed sebaceous neoplasms: a morphological, immunohistochemical and molecular analysis. Pathology 48:454–462. https://doi.org/10.1016/j.pathol.2016.05.004
Cryan JB, Haidar S, Ramkissoon LA, Bi WL, Knoff DS, Schultz N et al (2014) Clinical multiplexed exome sequencing distinguishes adult oligodendroglial neoplasms from astrocytic and mixed lineage gliomas. Oncotarget 5:8083–8092. https://doi.org/10.18632/oncotarget.2342
Piscuoglio S, Ng CK, Murray M, Burke KA, Edelweiss M, Geyer FC et al (2016) Massively parallel sequencing of phyllodes tumours of the breast reveals actionable mutations, and TERT promoter hotspot mutations and TERT gene amplification as likely drivers of progression. J Pathol 238:508–518. https://doi.org/10.1002/path.4672
van Kempen LC, Wang H, Aguirre ML, Spatz A, Kasymjanova G, Vilacha JF, Groves MR, Agulnik J, Small D (2018) Afatinib in osimertinib-resistant EGFR ex19del/T790M/P794L mutated NSCLC. J Thorac Oncol 13:e161–e163. https://doi.org/10.1016/j.jtho.2018.04.020
Harada T, Lopez-Chavez A, Xi L, Raffeld M, Wang Y, Giaccone G (2011) Characterization of epidermal growth factor receptor mutations in non-small-cell lung cancer patients of African-American ancestry. Oncogene 30:1744–1752. https://doi.org/10.1038/onc.2010.545
Kudchadkar R, Paraiso KHT, Smalley KSM (2012) Targeting mutant BRAF in melanoma: current status and future development of combination therapy strategies. Cancer J 18:124–131. https://doi.org/10.1097/PPO.0b013e31824b436e
Carbonell P, Turpin MC, Torres-Moreno D, Molina-Martínez I, García-Solano J, Perez-Guillermo M, Conesa-Zamora P (2011) Comparison of allelic discrimination by dHPLC, HRM, and TaqMan in the detection of BRAF mutation V600E. J Mol Diagn 13:467–473. https://doi.org/10.1016/j.jmoldx.2011.03.009
Karadima ML, Saetta AA, Chatziandreou I, Lazaris AC, Patsouris E, Tsavaris N (2016) The prognostic influence of BRAF mutation and other molecular, clinical and laboratory parameters in stage IV colorectal cancer. Pathol Oncol Res 22:707–714. https://doi.org/10.1007/s12253-016-0056-7
Kowalik A, Kowalska A, Kopczynski J, Walczyk A, Nowak E, Wypiorkiewicz E, Chodurska R, Pieciak L, Gozdz S (2014) Occurrence other than V600E mutation in the BRAF gene in papillary thyroid carcinoma. EJEA. https://doi.org/10.1530/endoabs.35.P1147
Konishi K, Takimoto M, Kaneko K, Makino R, Hirayama Y, Nozawa H, Kurahashi T, Kumekawa Y, Yamamoto T, Ito H, Yoshikawa N, Kusano M, Nakayama K, Rembacken BJ, Ota H, Imawari M (2006) BRAF mutations and phosphorylation status of mitogen-activated protein kinases in the development of flat and depressed-type colorectal neoplasias. Br J Cancer 94:311–317. https://doi.org/10.1038/sj.bjc.6602911
Melo ACD, Inada HKP, Soares FA, Barros M, Terra SBSP, Small IA, Sternberg C, Ferreira CGM (2011) Concomitant mutations in EGFR, KRAS, and BRAF genes in patients with radically resected non-small cell lung cancer (NSCLC) in Brazil. JCO 29:e21046–e21046. https://doi.org/10.1200/jco.2011.29.15_suppl.e21046
Tie J, Gibbs P, Lipton L, Christie M, Jorissen RN, Burgess AW, Croxford M, Jones I, Langland R, Kosmider S, McKay D, Bollag G, Nolop K, Sieber OM, Desai J (2011) Optimizing targeted therapeutic development: analysis of a colorectal cancer patient population with the BRAF(V600E) mutation. Int J Cancer 128:2075–2084. https://doi.org/10.1002/ijc.25555
Pita JM, Figueiredo IF, Moura MM, Leite V, Cavaco BM (2014) Cell cycle deregulation and TP53 and RAS mutations are major events in poorly differentiated and undifferentiated thyroid carcinomas. J Clin Endocrinol Metab 99:E497–E507. https://doi.org/10.1210/jc.2013-1512
Curtius K, Wright NA, Graham TA (2018) An evolutionary perspective on field cancerization. Nat Rev Cancer 18:19–32. https://doi.org/10.1038/nrc.2017.102
Mohan M, Jagannathan N (2014) Oral field cancerization: an update on current concepts. Oncol Rev 8:244. https://doi.org/10.4081/oncol.2014.244
Slaughter DP, Southwick HW, Smejkal W (1953) Field cancerization in oral stratified squamous epithelium; clinical implications of multicentric origin. Cancer 6:963–968. https://doi.org/10.1002/1097-0142(195309)6:5<963:aid-cncr2820060515>3.0.co;2-q
Califano J, van der Riet P, Westra W, Nawroz H, Clayman G, Piantadosi S, Corio R, Lee D, Greenberg B, Koch W, Sidransky D (1996) Genetic progression model for head and neck cancer: implications for field cancerization. Cancer Res 56(11):2488–2492
Sabharwal R, Mahendra A, Moon NJ, Gupta P, Jain A, Gupta S (2014) Genetically altered fields in head and neck cancer and second field tumor. South Asian J Cancer 3:151–153. https://doi.org/10.4103/2278-330X.136766
Hiley C, de Bruin EC, McGranahan N, Swanton C (2014) Deciphering intratumor heterogeneity and temporal acquisition of driver events to refine precision medicine. Genome Biol 15:453. https://doi.org/10.1186/s13059-014-0453-8
Xiong L, Cui S, Ding J, Sun Y, Zhang L, Zhao Y et al (2017) Dynamics of EGFR mutations in plasma recapitulates the clinical response to EGFR-TKIs in NSCLC patients. Oncotarget 8:63846–63856. https://doi.org/10.18632/oncotarget.19139
McGranahan N, Swanton C (2017) Clonal heterogeneity and tumor evolution: past, present, and the future. Cell 168:613–628. https://doi.org/10.1016/j.cell.2017.01.018
Johnson BE, Mazor T, Hong C, Barnes M, Aihara K, McLean CY et al (2014) Mutational analysis reveals the origin and therapy-driven evolution of recurrent glioma. Science 343:189–193. https://doi.org/10.1126/science.1239947
Maley CC, Galipeau PC, Finley JC, Wongsurawat VJ, Li X, Sanchez CA, Paulson TG, Blount PL, Risques RA, Rabinovitch PS, Reid BJ (2006) Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nat Genet 38:468–473. https://doi.org/10.1038/ng1768
Sievers CK, Zou LS, Pickhardt PJ, Matkowskyj KA, Albrecht DM, Clipson L, Bacher JW, Pooler BD, Moawad FJ, Cash BD, Reichelderfer M, Vo TN, Newton MA, Larget BR, Halberg RB (2017) Subclonal diversity arises early even in small colorectal tumours and contributes to differential growth fates. Gut 66:2132–2140. https://doi.org/10.1136/gutjnl-2016-312232
Califano J, Koch W, Sidransky D, Westra WH (2000) Inverted sinonasal papilloma : a molecular genetic appraisal of its putative status as a precursor to squamous cell carcinoma. Am J Pathol 156:333–337. https://doi.org/10.1016/S0002-9440(10)64734-7
d'Errico A, Zajacova J, Cacciatore A, Baratti A, Zanelli R, Alfonzo S, Beatrice F (2013) Occupational risk factors for sinonasal inverted papilloma: a case-control study. Occup Environ Med 70:703–708. https://doi.org/10.1136/oemed-2013-101384
Wang M-J, Noel JE (2017) Etiology of sinonasal inverted papilloma: a narrative review. World J Otorhinolaryngol Head Neck Surg 3:54–58. https://doi.org/10.1016/j.wjorl.2016.11.004
Vyse S, Huang PH (2019) Targeting EGFR exon 20 insertion mutations in non-small cell lung cancer. Signal Transduct Target Ther 4:5. https://doi.org/10.1038/s41392-019-0038-9
Acknowledgments
We thank the technical assistants of the Institute of Pathology for their support.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
SZ collected the clinical data, chose the cases for analysis, revised the H&E sections to decide on suitability, and wrote the manuscript; AE provided the FFPE material, critically supervised the project, and contributed to the manuscript; BS performed molecular analysis and wrote the manuscript. All authors contributed to the conception of the project.
Corresponding author
Ethics declarations
Specimen collection was conducted with informed consent in accordance with the ethics guidelines for the use of human material, approved by the Ethics Committee of the University of Rostock (reference number: A2017-0159).
Disclaimer
All authors approved the final version of the manuscript and agreed to be accountable for all aspects of the work.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: Retrospective Open Access.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zonnur, S., Erbersdobler, A. & Schneider, B. EGFR and BRAF mutations in inverted sinonasal papilloma — a more complex landscape?. Virchows Arch 478, 915–924 (2021). https://doi.org/10.1007/s00428-020-02945-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-020-02945-y