Abstract
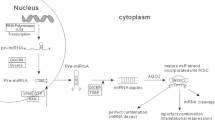
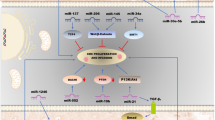
MicroRNAs are small RNAs that regulate gene expression at the post-transcriptional level. After their discovery 15 years ago, a new layer of gene regulation was introduced into every field of human biology and medicine. Considering the strong association between genetic alterations and neoplastic diseases, it is not surprising that there is a special focus on miRNAs and cancer. A multitude of experimental studies on colorectal cancer, the most common cancer site and furthermore the second most common cause of death due to cancer, deliver insight into miRNA-mediated, regulatory links to well-known oncogenic and tumour suppressor signalling pathways. Furthermore, several investigations have described the ability of microRNA expression patterns to predict prognosis in colon cancer and support diagnosis of poorly differentiated tumours. In this short review, we give a comprehensive overview focussed on miRNAs in colorectal cancer research.



Similar content being viewed by others
Abbreviations
- 3′UTR:
-
3′ untranslated regions of mRNAs
- CRC:
-
Colorectal cancer
- CTGF:
-
Connective tissue growth factor
- CUP:
-
Cancer of unknown primary
- EMT:
-
Epithelial-to-mesenchymal transition
- miR/miRNA:
-
microRNA
- miRAGE:
-
miRNA serial analysis of gene expression
- mRNA:
-
messenger RNA
- PDCD4:
-
programmed cell death 4
- PI-3-K:
-
Phosphatidylinositol-3-kinase-AKT pathway
- PTEN:
-
Phosphatase and tensin homolog
- siRNA:
-
Small interfering RNA
- SNP:
-
Single nucleotide polymorphism
- SIRT1:
-
Silent information regulator 1
- TGF-β:
-
Transforming growth factor β
- TNF-α:
-
Tumour necrosis factor α
- Tsp-1:
-
Thrombospondin-1
- UICC:
-
International Union Against Cancer (classification system)
- ZEB1/ZEB2:
-
Zinc finger E-box binding homeobox 1/2
References
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75:843–854
Sassen S, Miska EA, Caldas C (2008) MicroRNA: implications for cancer. Virchows Arch 452:1–10
Esquela-Kerscher A, Slack FJ (2006) Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer 6:259–269
Singh SK, Pal Bhadra M, Girschick HJ et al (2008) MicroRNAs—micro in size but macro in function. Febs J 275:4929–4944
Akao Y, Nakagawa Y, Naoe T (2007) MicroRNA-143 and -145 in colon cancer. DNA Cell Biol 26:311–320
Baek D, Villen J, Shin C et al (2008) The impact of microRNAs on protein output. Nature 455:64–71
Selbach M, Schwanhausser B, Thierfelder N et al (2008) Widespread changes in protein synthesis induced by microRNAs. Nature 455:58–63
Griffiths-Jones S, Grocock RJ, van Dongen S et al (2006) miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 34:D140–D144
Ying SY, Lin SL (2006) Current perspectives in intronic micro RNAs (miRNAs). J Biomed Sci 13:5–15
Lim LP, Lau NC, Garrett-Engele P et al (2005) Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433:769–773
Leslie A, Carey FA, Pratt NR et al (2002) The colorectal adenoma–carcinoma sequence. Br J Surg 89:845–860
Hermeking H (2007) p53 enters the microRNA world. Cancer Cell 12:414–418
Chang TC, Wentzel EA, Kent OA et al (2007) Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell 26:745–752
Yamakuchi M, Ferlito M, Lowenstein CJ (2008) miR-34a repression of SIRT1 regulates apoptosis. Proc Natl Acad Sci USA 105:13421–13426
Welch C, Chen Y, Stallings RL (2007) MicroRNA-34a functions as a potential tumor suppressor by inducing apoptosis in neuroblastoma cells. Oncogene 26:5017–5022
Bommer GT, Gerin I, Feng Y et al (2007) p53-mediated activation of miRNA34 candidate tumor-suppressor genes. Curr Biol 17:1298–1307
Lodygin D, Tarasov V, Epanchintsev A et al (2008) Inactivation of miR-34a by aberrant CpG methylation in multiple types of cancer. Cell Cycle 7:2591–2600
Tazawa H, Tsuchiya N, Izumiya M et al (2007) Tumor-suppressive miR-34a induces senescence-like growth arrest through modulation of the E2F pathway in human colon cancer cells. Proc Natl Acad Sci USA 104:15472–15477
Toyota M, Suzuki H, Sasaki Y et al (2008) Epigenetic silencing of microRNA-34b/c and B-cell translocation gene 4 is associated with CpG island methylation in colorectal cancer. Cancer Res 68:4123–4132
Takamizawa J, Konishi H, Yanagisawa K et al (2004) Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 64:3753–3756
Akao Y, Nakagawa Y, Naoe T (2006) let-7 microRNA functions as a potential growth suppressor in human colon cancer cells. Biol Pharm Bull 29:903–906
Chen X, Guo X, Zhang H et al. (2009) Role of miR-143 targeting KRAS in colorectal tumorigenesis. Oncogene. doi:10.1038/onc.2008.474
Michael MZ, OC SM, van Holst Pellekaan NG et al (2003) Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res 1:882–891
Powell SM, Zilz N, Beazer-Barclay Y et al (1992) APC mutations occur early during colorectal tumorigenesis. Nature 359:235–237
Nagel R, le Sage C, Diosdado B et al (2008) Regulation of the adenomatous polyposis coli gene by the miR-135 family in colorectal cancer. Cancer Res 68:5795–5802
Shell S, Park SM, Radjabi AR et al (2007) Let-7 expression defines two differentiation stages of cancer. Proc Natl Acad Sci USA 104:11400–11405
Spaderna S, Schmalhofer O, Hlubek F et al (2006) A transient, EMT-linked loss of basement membranes indicates metastasis and poor survival in colorectal cancer. Gastroenterology 131:830–840
Burk U, Schubert J, Wellner U et al (2008) A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep 9:582–589
Spaderna S, Schmalhofer O, Wahlbuhl M et al (2008) The transcriptional repressor ZEB1 promotes metastasis and loss of cell polarity in cancer. Cancer Res 68:537–544
Xi Y, Formentini A, Chien M et al (2006) Prognostic values of microRNAs in colorectal cancer. Biomark Insights 2:113–121
Nakajima G, Hayashi K, Xi Y et al (2006) Non-coding MicroRNAs hsa-let-7 g and hsa-miR-181b are associated with chemoresponse to S-1 in colon cancer. Cancer Genomics Proteomics 3:317–324
Philp AJ, Campbell IG, Leet C et al (2001) The phosphatidylinositol 3′-kinase p85alpha gene is an oncogene in human ovarian and colon tumors. Cancer Res 61:7426–7429
Guo C, Sah JF, Beard L et al (2008) The noncoding RNA, miR-126, suppresses the growth of neoplastic cells by targeting phosphatidylinositol 3-kinase signaling and is frequently lost in colon cancers. Genes Chromosomes Cancer 47:939–946
Vivanco I, Sawyers CL (2002) The phosphatidylinositol 3-Kinase AKT pathway in human cancer. Nat Rev Cancer 2:489–501
Meng F, Henson R, Wehbe-Janek H et al (2007) MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 133:647–658
Schetter AJ, Leung SY, Sohn JJ et al (2008) MicroRNA expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. JAMA 299:425–436
Slaby O, Svoboda M, Fabian P et al (2007) Altered expression of miR-21, miR-31, miR-143 and miR-145 is related to clinicopathologic features of colorectal cancer. Oncology 72:397–402
Mudduluru G, Medved F, Grobholz R et al (2007) Loss of programmed cell death 4 expression marks adenoma-carcinoma transition, correlates inversely with phosphorylated protein kinase B, and is an independent prognostic factor in resected colorectal cancer. Cancer 110:1697–1707
Asangani IA, Rasheed SA, Nikolova DA et al (2008) MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 27:2128–2136
Mendell JT (2008) miRiad roles for the miR-17–92 cluster in development and disease. Cell 133:217–222
Dews M, Homayouni A, Yu D et al (2006) Augmentation of tumor angiogenesis by a Myc-activated microRNA cluster. Nat Genet 38:1060–1065
Zhao HY, Ooyama A, Yamamoto M et al (2008) Down regulation of c-Myc and induction of an angiogenesis inhibitor, thrombospondin-1, by 5-FU in human colon cancer KM12C cells. Cancer Lett 270:156–163
Landi D, Gemignani F, Naccarati A et al (2008) Polymorphisms within micro-RNA-binding sites and risk of sporadic colorectal cancer. Carcinogenesis 29:579–584
Iorio MV, Ferracin M, Liu CG et al (2005) MicroRNA gene expression deregulation in human breast cancer. Cancer Res 65:7065–7070
Porkka KP, Pfeiffer MJ, Waltering KK et al (2007) MicroRNA expression profiling in prostate cancer. Cancer Res 67:6130–6135
Calin GA, Ferracin M, Cimmino A et al (2005) A MicroRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med 353:1793–1801
Bandres E, Cubedo E, Agirre X et al (2006) Identification by Real-time PCR of 13 mature microRNAs differentially expressed in colorectal cancer and non-tumoral tissues. Mol Cancer 5:29
Schepeler T, Reinert JT, Ostenfeld MS et al (2008) Diagnostic and prognostic microRNAs in stage II colon cancer. Cancer Res 68:6416–6424
Popat S, Hubner R, Houlston RS (2005) Systematic review of microsatellite instability and colorectal cancer prognosis. J Clin Oncol 23:609–618
Ribic CM, Sargent DJ, Moore MJ et al (2003) Tumor microsatellite-instability status as a predictor of benefit from fluorouracil-based adjuvant chemotherapy for colon cancer. N Engl J Med 349:247–257
Lanza G, Ferracin M, Gafa R et al (2007) mRNA/microRNA gene expression profile in microsatellite unstable colorectal cancer. Mol Cancer 6:54
Linsley PS, Schelter J, Burchard J et al (2007) Transcripts targeted by the microRNA-16 family cooperatively regulate cell cycle progression. Mol Cell Biol 27:2240–2252
Lu J, Getz G, Miska EA et al (2005) MicroRNA expression profiles classify human cancers. Nature 435:834–838
Ramaswamy S, Tamayo P, Rifkin R et al (2001) Multiclass cancer diagnosis using tumor gene expression signatures. Proc Natl Acad Sci USA 98:15149–15154
Volinia S, Calin GA, Liu CG et al (2006) A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA 103:2257–2261
Rosenfeld N, Aharonov R, Meiri E et al (2008) MicroRNAs accurately identify cancer tissue origin. Nat Biotechnol 26:462–469
Pavlidis N, Briasoulis E, Hainsworth J et al (2003) Diagnostic and therapeutic management of cancer of an unknown primary. Eur J Cancer 39:1990–2005
Pentheroudakis G, Golfinopoulos V, Pavlidis N (2007) Switching benchmarks in cancer of unknown primary: from autopsy to microarray. Eur J Cancer 43:2026–2036
Lujambio A, Calin GA, Villanueva A et al (2008) A microRNA DNA methylation signature for human cancer metastasis. Proc Natl Acad Sci USA 105:13556–13561
Gaur A, Jewell DA, Liang Y et al (2007) Characterization of microRNA expression levels and their biological correlates in human cancer cell lines. Cancer Res 67:2456–2468
Kumar MS, Lu J, Mercer KL et al (2007) Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nat Genet 39:673–677
Nakajima N, Takahashi T, Kitamura R et al (2006) MicroRNA-1 facilitates skeletal myogenic differentiation without affecting osteoblastic and adipogenic differentiation. Biochem Biophys Res Commun 350:1006–1012
Chen X, Ba Y, Ma L et al (2008) Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 18:997–1006
Srinivasan M, Sedmak D, Jewell S (2002) Effect of fixatives and tissue processing on the content and integrity of nucleic acids. Am J Pathol 161:1961–1971
Bresters D, Schipper ME, Reesink HW et al (1994) The duration of fixation influences the yield of HCV cDNA-PCR products from formalin-fixed, paraffin-embedded liver tissue. J Virol Methods 48:267–272
Xi Y, Nakajima G, Gavin E et al (2007) Systematic analysis of microRNA expression of RNA extracted from fresh frozen and formalin-fixed paraffin-embedded samples. RNA 13:1668–1674
Nuovo GJ (2008) In situ detection of precursor and mature microRNAs in paraffin embedded, formalin fixed tissues and cell preparations. Methods 44:39–46
Silahtaroglu AN, Nolting D, Dyrskjot L et al (2007) Detection of microRNAs in frozen tissue sections by fluorescence in situ hybridization using locked nucleic acid probes and tyramide signal amplification. Nat Protoc 2:2520–2528
Vasudevan S, Tong Y, Steitz JA (2007) Switching from repression to activation: microRNAs can up-regulate translation. Science 318:1931–1934
Orom UA, Nielsen FC, Lund AH (2008) MicroRNA-10a binds the 5'UTR of ribosomal protein mRNAs and enhances their translation. Mol Cell 30:460–471
Cummins JM, He Y, Leary RJ et al (2006) The colorectal microRNAome. Proc Natl Acad Sci USA 103:3687–3692
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Faber, C., Kirchner, T. & Hlubek, F. The impact of microRNAs on colorectal cancer. Virchows Arch 454, 359–367 (2009). https://doi.org/10.1007/s00428-009-0751-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-009-0751-9