Abstract
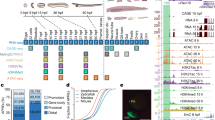
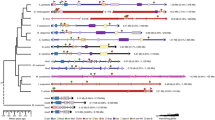
The single amphioxus Hox cluster contains 15 genes and may well resemble the ancestral chordate Hox cluster. We have sequenced the Hox genomic complement of the European amphioxus Branchiostoma lanceolatum and compared it to the American species, Branchiostoma floridae, by phylogenetic footprinting to gain insights into the evolution of Hox gene regulation in chordates. We found that Hox intergenic regions are largely conserved between the two amphioxus species, especially in the case of genes located at the 3′ of the cluster, a trend previously observed in vertebrates. We further compared the amphioxus Hox cluster with the human HoxA, HoxB, HoxC, and HoxD clusters, finding several conserved noncoding regions, both in intergenic and intronic regions. This suggests that the regulation of Hox genes is highly conserved across chordates, consistent with the similar Hox expression patterns in vertebrates and amphioxus.




Similar content being viewed by others
References
Amemiya CT, Prohaska SJ, Hill-Force A, Cook A, Wasserscheid J, Ferrier DEK, Pascual-Anaya J, Garcia-Fernàndez J, Dewar K, Stadler PF (2008) The amphioxus Hox cluster: characterization, comparative genomics, and evolution. J Exp Zoolog (Mol Dev Evol) 310B:n/a
Bray N, Dubchak I, Pachter L (2003) AVID: a global alignment program. Genome Res 13:97–102
Brudno M, Do CB, Cooper GM, Kim MF, Davydov E, Program NCS, Green ED, Sidow A, Batzoglou S (2003) LAGAN and Multi-LAGAN: Efficient tools for large-scale multiple alignment of genomic DNA. Genome Res 13:721–731
Cañestro C, Hjelmqvist L, Albalat R, Garcia-Fernàndez J, González-Duarte R, Jornvall H (2000) Amphioxus alcohol dehydrogenase is a class 3 form of single type and of structural conservation but with unique developmental expression. Eur J Biochem 267:6511–6518
Chiu C-H, Amemiya C, Dewar K, Kim C-B, Ruddle FH, Wagner GP (2002) Molecular evolution of the HoxA cluster in the three major gnathostome lineages. Proc Natl Acad Sci USA 99:5492–5497
Duboule D (1994) Temporal colinearity and the phylotypic progression: a basis for the stability of a vertebrate Bauplan and the evolution of morphologies through heterochrony. Dev Suppl 1994:135–142
Duboule D (2007) The rise and fall of Hox gene clusters. Development 134:2549–2560
Duboule D, Dollé P (1989) The structural and functional organization of the murine HOX gene family resembles that of Drosophila homeotic genes. EMBO J 8:1497–1505
Ewing B, Green P (1998) Base-calling of automated sequencer traces using Phred. II. Error Probabilities. Genome Res 8:186–194
Ewing B, Hillier L, Wendl MC, Green P (1998) Base-calling of automated sequencer traces using Phred. I. Accuracy assessment. Genome Res 8:175–185
Ferrier DEK, Minguillón C, Holland PWH, Garcia-Fernàndez J (2000) The amphioxus Hox cluster: deuterostome posterior flexibility and Hox14. Evol Dev 2:284–293
Force A, Lynch M, Pickett FB, Amores A, Yan YL, Postlethwait J (1999) Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151:1531–1545
Garcia-Fernàndez J, Holland PWH (1994) Archetypal organization of the amphioxus Hox gene cluster. Nature 370:563–566
Gómez-Skarmeta JL, Lenhard B, Becker TS (2006) New technologies, new findings, and new concepts in the study of vertebrate cis-regulatory sequences. Dev Dyn 235:870–885
Gordon D, Abajian C, Green P (1998) Consed: a graphical tool for sequence finishing. Genome Res 8:195–202
Haerry TE, Gehring WJ (1996) Intron of the mouse Hoxa-7 gene contains conserved homeodomain binding sites that can function as an enhancer element in Drosophila. Proc Natl Acad Sci USA 93:13884–13889
Haerry TE, Gehring WJ (1997) A conserved cluster of homeodomain binding sites in the mouse Hoxa-4 intron functions in Drosophila embryos as an enhancer that is directly regulated by Ultrabithorax. Dev Biol 186:1–15
Holland LZ, Albalat R, Azumi K, Benito-Gutiérrez È, Blow MJ, Bronner-Fraser M, Brunet F, Butts T, Candiani S, Dishaw LJ, Ferrier DEK, Garcia-Fernàndez J, Gibson-Brown JJ, Gissi C, Godzik A, Hallböök F, Hirose D, Hosomichi K, Ikuta T, Inoko H, Kasahara M, Kasamatsu J, Kawashima T, Kimura A, Kobayashi M, Kozmik Z, Kubokawa K, Laudet V, Litman GW, McHardy AC, Meulemans D, Nonaka M, Olinski RP, Pancer Z, Pennacchio LA, Pestarino M, Rast JP, Rigoutsos I, Robinson-Rechavi M, Roch G, Saiga H, Sasakura Y, Satake M, Satou Y, Schubert M, Sherwood N, Shiina T, Takatori N, Tello J, Vopalensky P, Wada S, Xu A, Ye Y, Yoshida K, Yoshizaki F, Yu J-K, Zhang Q, Zmasek CM, Putnam NH, Rokhsar DS, Satoh N, Holland PWH (2008) The amphioxus genome illuminates vertebrate origins and cephalochordate biology. Genome Res 18:1100–1111
International Human Genome Sequence Consortium (2004) Finishing the euchromatic sequence of the human genome. Nature 431:931–945
Kessel M, Gruss P (1991) Homeotic transformations of murine vertebrae and concomitant alteration of Hox codes induced by retinoic acid. Cell 67:89–104
Kim CB, Amemiya C, Bailey W, Kawasaki K, Mezey J, Miller W, Minoshima S, Shimizu N, Wagner G, Ruddle F (2000) Hox cluster genomics in the horn shark, Heterodontus francisci. Proc Natl Acad Sci USA 97:1655–1660
Kon T, Nohara M, Yamanoue Y, Fujiwara Y, Nishida M, Nishikawa T (2007) Phylogenetic position of a whale-fall lancelet (Cephalochordata) inferred from whole mitochondrial genome sequences. BMC Evolutionary Biology 7:127
Lewis EB (1978) A gene complex controlling segmentation in Drosophila. Nature 276:565–570
Mayor C, Brudno M, Schwartz JR, Poliakov A, Rubin EM, Frazer KA, Pachter LS, Dubchak I (2000) VISTA: visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 16:1046–1047
Nohara M, Nishida M, Nishikawa T (2005) New complete mitochondrial DNA sequence of the lancelet Branchiostoma lanceolatum (Cephalochordata) and the Identity of this Species’ Sequences. Zoological Science 22:671–674
Packer AI, Crotty DA, Elwell VA, Wolgemuth DJ (1998) Expression of the murine Hoxa4 gene requires both autoregulation and a conserved retinoic acid response element. Development 125:1991–1998
Podlasek C, Houston J, McKenna KE, McVary KT (2002) Posterior Hox gene expression in developing genitalia. Evol Dev 4:142–163
Prohaska SJ, Fried C, Flamm C, Wagner GP, Stadler PF (2004) Surveying phylogenetic footprints in large gene clusters: applications to Hox cluster duplications. Mol Phylogenet Evol 31:581–604
Richardson MK, Crooijmans RP, Groenen MA (2007) Sequencing and genomic annotation of the chicken (Gallus gallus) Hox clusters, and mapping of evolutionarily conserved regions. Cytogenet Genome Res 117:110–119
Santini S, Boore JL, Meyer A (2003) Evolutionary conservation of regulatory elements in vertebrate Hox gene clusters. Genome Res 13:1111–1122
Schubert M, Holland ND, Laudet V, Holland LZ (2006) A retinoic acid-Hox hierarchy controls both anterior/posterior patterning and neuronal specification in the developing central nervous system of the cephalochordate amphioxus. Developmental Biology 296:190–202
Tanzer A, Amemiya CT, Kim CB, Stadler PF (2005) Evolution of microRNAs located within Hox gene clusters. J Exp Zoolog B Mol Dev Evol 304:75–85
Wada H, Garcia-Fernàndez J, Holland PWH (1999) Colinear and segmental expression of amphioxus Hox genes. Dev Biol 213:131–141
Wagner GP, Lynch VJ (2005) Molecular evolution of evolutionary novelties: the vagina and uterus of therian mammals. J Exp Zoolog B Mol Dev Evol 304B:580–592
Wasserman WW, Sandelin A (2004) Applied bioinformatics for the identification of regulatory elements. Nat Rev Genet 5:276–287
Yamamoto KR, Alberts BM, Benzinger R, Lawhorne L, Treiber G (1970) Rapid bacteriophage sedimentation in the presence of polyethylene glycol and its application to large-scale virus purification. Virology 40:734–744
Zakany J, Duboule D (2007) The role of Hox genes during vertebrate limb development. Curr Opin Genet Dev 17:359–366
Acknowledgments
We wish to thank Nacho Maeso for helping with the Vista browser; Nacho Maeso and Manuel Irimia for critical reading of the manuscript and Senda Jiménez-Delgado for helpful discussions; Ricard Albalat for kindly providing the lambda genomic library of B. lanceolatum; and Jon Permanyer for his help in using the GPS®-1 Genome Priming System and Phred/Phrap/Consed software. This research was supported by grant BFU2005-00252 from Ministerio de Educación y Ciencia, Spain. J.P.-A. holds an FI fellowship of the Generalitat de Catalunya and S.D’A. a “Juan de la Cierva” postdoctoral contract of the Ministerio de Educación y Ciencia, Spain.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. J. Gibson-Brown
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary file
(TXT 189 KB)
Table S1
Similarity between B. floridae and B. lanceolatum Hox genes. (DOC 44.5 KB)
Table S2
CNEs between Hox sequences of B. lanceolatum and Human HoxA cluster. (DOC 31 KB)
Table S3
CNEs between Hox sequences of B. lanceolatum and Human HoxB cluster. (DOC 7 KB)
Table S4
CNEs between Hox sequences of B. lanceolatum and Human HoxC cluster. (DOC 27 KB)
Table S5
CNEs between Hox sequences of B. lanceolatum and Human HoxD cluster. (DOC 29 KB)
Rights and permissions
About this article
Cite this article
Pascual-Anaya, J., D’Aniello, S. & Garcia-Fernàndez, J. Unexpectedly large number of conserved noncoding regions within the ancestral chordate Hox cluster. Dev Genes Evol 218, 591–597 (2008). https://doi.org/10.1007/s00427-008-0246-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-008-0246-8