Abstract
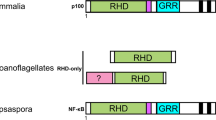
The Rel/nuclear factor-kappa B (NF-κB) and nuclear factor of activated T-cells (NFAT) transcription factors contribute to the regulation of an assortment of biological processes by binding DNA with high specificity using their Rel homology domain (RHD). Recently, it has been shown that members of these gene families are present in the genome of the anthozoan cnidarian Nematostella vectensis, indicating that they predate the evolution of the most recent ancestor to living bilaterians. By identifying a single NF-κB gene in the genome of the demosponge Amphimedon queenslandica, a representative of an even earlier branching metazoan lineage, we demonstrate here that the Rel/NF-κB family originated at the dawn of the Metazoa. There is no evidence of RHDs in fungal and choanoflagellate genomes, supporting the notion that the RHD is a metazoan-specific innovation. The A. queenslandica gene (AmqNF-κB) encodes a protein that is highly similar in structure to the vertebrate NF-κB p50/p52 proteins, possessing both a RHD and ankyrin (ANK) repeats. The intact AmqNF-κB contrasts with the N. vectensis NF-κB, which lacks ANK repeats, and suggests that the ancestral metazoan NF-κB was configured identically to contemporary vertebrate and sponge forms. AmqNF-κB is expressed during A. queenslandica embryogenesis, suggesting a developmental role.






Similar content being viewed by others
References
Adamska M, Degnan SM, Green KM, Adamski M, Craigie A, Larroux C, Degnan BM (2007a) Wnt and TGF-β expression in the sponge Amphimedon queenslandica and the origin of metazoan embryonic patterning. PLoS ONE 2:e1031
Adamska M, Matus DQ, Adamski M, Green KM, Rokhsar DS, Martindale MQ, Degnan BM (2007b) The evolutionary origin of hedgehog proteins. Curr Biol 17:R836–R837
Baeuerle PA, Henkel T (1994) Function and activation of NF-κB in the immune system. Annu Rev Immunol 12:141–179
Baldwin ASJ (1996) The NF-κB and IκB proteins: new discoveries and insights. Annu Rev Immunol 14:649–681
Benedito AB, Lehtinen M, Massol R, Lopes UG, Kirchhausen T, Rao A, Bonni A (2005) The transcription factor NFAT3 mediates neuronal survival. J Biol Chem 280:2818–2825
Borchiellini C, Manuel M, Alivon E, Boury-Esnault N, Vacelet J, Le Parco Y (2001) Sponge paraphyly and the origin of Metazoa. J Evol Biol 14:171–179
Bork P (1993) Hundreds of ankyrin-like repeats in functionally diverse proteins: mobile modules that cross phyla horizontally? Proteins 17:363–374
Brownell E, O’Brien SJ, Nash WG, Rice N (1985) Genetic characterisation of human c-rel sequences. Mol Cell Biol 5:2826–2831
Cavalier-Smith T, Chao EEY (2003) Phylogeny of choanozoa, apusozoa, and other protozoa and early eukaryote megaevolution. J Mol Evol 56:540–563
Chen IS, Mak TW, O’Rear JJ, Temin HM (1981) Characterisation of reticuloendotheliosis virus strain T DNA and isolation of a novel variant of reticuloendotheliosis virus strain T by molecular cloning. J Virol 40:800–811
Chen Z, Hagler J, Palombella VJ, Melandri F, Scherer D, Ballard D, Maniatis T (1995) Signal-induced site-specific phosphorylation targets IκBα to the ubiquitin-proteasome pathway. Genes Dev 9:1586–1597
DiDonato JA, Mercurio F, Karin M (1995) Phosphorylation of IκBα precedes but is not sufficient for its dissociation from NF-κB. Mol Cell Biol 15:1302–1311
Dushay MS, Asling B, Hultmark D (1996) Origins of immunity: Relish, a compound Rel-like gene in the antibacterial defense of Drosophila. Proc Natl Acad Sci U S A 93:10343–10347
Felsenstein J (2003) PHYLIP (Phylogeny Inference Package). Department of Genetics, University of Washington, Seattle (distributed by the author)
Ghosh G, Van Duyne G, Ghosh S, Sigler PB (1995) Structure of NF-κB p50 homodimer bound to a κB site. Nature 373:303–310
Graef IA, Gastier JM, Francke U, Crabtree GR (2001) Evolutionary relationships among Rel domains indicate functional diversification by recombination. Proc Natl Acad Sci U S A 98:5740–5745
Hayden MS, Ghosh S (2004) Signaling to NF-κB. Genes Dev 18:2195–2224
Hogan PG, Chen L, Nardone J, Rao A (2003) Transcriptional regulation by calcium, calcineurin, and NFAT. Genes Dev 17:2205–2232
Huguet C, Crepieux P, Laudet V (1997) Rel/NF-κB transcription factors and IκB inhibitors: evolution from a unique common ancestor. Oncogene 15:2965–2974
Ip YT, Reach M, Engstrom Y, Kadalayil L, Cai H, González-Crespo S, Tatei K, Levine M (1993) Dif, a dorsal-related gene that mediates an immune response in Drosophila. Cell 75:753–763
Keyser P, Borge-Renberga K, Hultmark D (2007) The Drosophila NFAT homolog is involved in salt stress tolerance. Insect Biochem Mol Biol 37:356–362
Larroux C, Fahey B, Liubicich D, Hinman V, Gauthier M, Gongora M, Green K, Worheide G, Leys S, Degnan BM (2006) Developmental expression of transcription factor genes in a demosponge: insights into the origin of metazoan multicellularity. Evol Dev 8:150–173
Larroux C, Fahey B, Degnan SM, Adamski M, Rokhsar DS, Degnan BM (2007) The NK homeobox gene cluster predates the origin of Hox genes. Curr Biol 17:706–710
Leys S, Degnan BM (2001) Cytological basis of photoresponsive behaviour in a sponge larva. Biol Bull 201:323–338
Leys S, Degnan BM (2002) Embryogenesis and metamorphosis in a haplosclerid demosponge: gastrulation and transdifferentiation of larval ciliated cells to choanocytes. Invert Biol 121:171–189
Loh C, Shaw KT, Carew J, Viola JP, Luo C, Perrino BA, Rao A (1996) Calcineurin binds the transcription factor NFAT1 and reversibly regulates its activity. J Biol Chem 271:10884–10891
Macian F (2005) NFAT proteins: key regulators of T-cell development and function. Nat Rev Immunol 5:472–484
Matthews JR, Wakasugi N, Virelizier JL, Yodoi J, Hay RT (1992) Thioredoxin regulates the DNA binding activity of NF-κB by reduction of a disulphide bond involving cysteine 62. Nucleic Acids Res 20:3821–3830
Matthews JR, Kaszubska W, Turcatti G, Wells TN, Hay RT (1993) Role of cysteine 62 in DNA recognition by the P50 subunit of NF-κB. Nucleic Acids Res 21:1727–1734
Medina M, Collins AG, Silberman JD, Sogin ML (2001) Evaluating hypotheses of basal animal phylogeny using complete sequences of large and small subunit rRNA. Proc Natl Acad Sci U S A 98:9707–9712
Mercurio F, DiDonato JA, Rosette C, Karin M (1992) Molecular cloning and characterisation of a novel Rel/NF-κB family member displaying structural and functional homology to NF-κB p50/p105. DNA Cell Biol 11:523–537
Miller DJ, Hemmrich G, Ball EE, Hayward DC, Khalturin K, Funayama N, Agata K, Bosch TC (2007) The innate immune repertoire in cnidaria—ancestral complexity and stochastic gene loss. Genome Biol 8:R59
Müller CW, Rey FA, Sodeoka M, Verdine GL, Harrison SC (1995) Structure of the NF-κB p50 homodimer bound to DNA. Nature 373:311–317
Müller WEG, Blumbach B, Müller IM (1999) Evolution of the innate and adaptive immune systems: relationships between potential immune molecules in the lowest metazoan phylum (Porifera) and those in vertebrates. Transplantation 68:1215–1227
Müller WEG, Schröder HC, Skorokhod A, Bünz C, Müller IM, Grebenjuk VA (2001) Contribution of sponge genes to unravel the genome of the hypothetical ancestor of Metazoa (Urmetazoa). Gene 276:161–173
Ruben SM, Dillon PJ, Schreck R, Henkel T, Chen CH, Maher M, Baeuerle PA, Rosen CA (1991) Isolation of a rel-related human cDNA that potentially encodes the 65-kD subunit of NF-κB. Science 251:1490–1493
Ruben S, Klement J, Coleman T, Maher M, Chen C, Rosen C (1992) I-Rel: a novel rel-related protein that inhibits NF-κB transcriptional activity. Genes Dev 6:745–760
Sakarya O, Armstrong KA, Adamska M, Adamski M, Wang IF, Tidor B, Degnan BM, Oakley TH, Kosik KS (2007) A post-synaptic scaffold at the origin of the animal kingdom. PLoS ONE 2:e506
Sedgwick SG, Smerdon SJ (1999) The ankyrin repeat: a diversity of interactions on a common structural framework. Trends Biochem Sci 24:311–316
Sen R, Baltimore D (1986) Inducibility of kappa immunoglobulin enhancer-binding protein NF-κB by a posttranslational mechanism. Cell 47:921–928
Siebenlist U, Franzoso G, Brown K (1994) Structure, regulation and function of NF-κB. Annu Rev Cell Biol 10:405–455
Simionato E, Ledent V, Richards G, Thomas-Chollier M, Kerner P, Coornaert D, Degnan BM, Vervoort M (2007) Origin and diversification of the basic helix–loop–helix gene family in metazoans: insights from comparative genomics. BMC Evol Biol 7:33
Steward R (1987) Dorsal, an embryonic polarity gene in Drosophila, is homologous to the vertebrate proto-oncogene, c-rel. Science 238:692–694
Sullivan JC, Kalaitzidis D, Gilmore TD, Finnerty JR (2007) Rel homology domain-containing transcription factors in the cnidarian Nematostella vectensis. Dev Genes Evol 217:63–72
Thompson JD, Higgins DG, Gibson TJ (1994) Clustal W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Traenckner E, Pahl H, Henkel T, Schmidt K, Wilk S, Baeuerle P (1995) Phosphorylation of human IκBα on serines 32 and 36 controls IκBα proteolysis and NF-κB activation in response to diverse stimuli. EMBO J 14:2876–2883
Wiens M, Korzhev M, Krasko A, Thakur NL, Perovic-Ottstadt S, Breter HJ, Ushijima H, Diehl-Siefert B, Müller IM, Müller WEG (2005) Innate immune defence of the sponge Suberites domuncula against bacteria involves a MyD88-dependent signalling pathway: induction of a perforin-like molecule. J Biol Chem 280:27949–27959
Wiens M, Korzhev M, Perovic-Ottstadt S, Luthringer B, Brandt D, Klein S, Müller WEG (2007) Toll-like receptors are part of the innate immune defense system of sponges (Demospongiae: Porifera). Mol Biol Evol 24:792–804
Acknowledgement
This work was supported by Australian Research Council grants to B.M.D. We gratefully acknowledge the contribution of The United States Department of Energy Joint Genome Institute in the production of Amphimedon (Reniera) genomic and EST sequences used in this study through the Community Sequencing Program. We thank the Director and staff of the University of Queensland Heron Island Research Station for field assistance; the Great Barrier Reef Marine Park Authority for granting permission to carry out this research; and Maja Adamska, Sandie Degnan, Claire Larroux, and Gemma Richards for their valuable advice.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Q. Martindale
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S2
The nucleotide and deduced amino acid sequence of A. queenslandica NF-κB cDNA. Exons 1 to 24 are delimited in the text with alternating blue and black text. The cDNA is 3,601-bp long and encodes a protein of 1,095 amino acids (DOC 33.0 KB)
Fig. S3
Comparison of the amino acid sequence of AmqNF-κB ANK domains with other genes containing the ANK-repeat motif. The intron boundaries are shown in red for known sequences (the genomic predictions for NvBcl3 and NvIkB presented gaps and therefore did not permit determination of the intron exon boundaries of the ANK-repeat motif) (DOC 34.5 KB)
Table S1
Sequence ID for genes used for phylogenetic analyses with corresponding Uniprot primary accession numbers (available on the web site http://www.ebi.uniprot.org/index.shtml) and species name (DOC 51.0 KB)
Rights and permissions
About this article
Cite this article
Gauthier, M., Degnan, B.M. The transcription factor NF-κB in the demosponge Amphimedon queenslandica: insights on the evolutionary origin of the Rel homology domain. Dev Genes Evol 218, 23–32 (2008). https://doi.org/10.1007/s00427-007-0197-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-007-0197-5