Abstract
Main conclusion
PpMYB17 positively regulates flavonoid biosynthesis in pear fruit by activating PpCHS, PpCHI, PpF3H, and PpFLS in the flavonoid biosynthesis pathway independently of bHLH or WD40 cofactors in the MBW complex.
Abstract
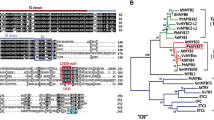
Flavonoids are important secondary metabolites in plants. The flavonoid biosynthesis pathway is regulated by various transcription factors, with MYB transcription factors considered to be the key regulators. However, the regulation of flavonoid biosynthesis in the pear fruit has not been fully characterized. The R2R3-MYB transcription factor PpMYB17 was isolated from ‘Red Zaosu’ pear fruit and functionally characterized. An exposure to light upregulated PpMYB17 expression in the pear fruit. A phylogenetic analysis indicated PpMYB17 is related to the flavonol regulators. A subcellular localization assay suggested that PpMYB17 is a nuclear protein. Overexpression of PpMYB17 increased the flavonoid content of pear calli and Arabidopsis via the upregulated expression of structural genes in the flavonoid biosynthesis pathway, especially FLS. The LC–MS/MS analysis revealed most of the differentially accumulated flavonols, flavanones, flavones, isoflavones, and anthocyanins were significantly more abundant in PpMYB17-overexpressing calli than in wild-type calli. Moreover, PpMYB17 did not interact with PpbHLH3, PpbHLH33, or PpWD40 in a yeast system. Dual-luciferase assays demonstrated that PpMYB17 strongly activates the promoters of PpCHS, PpCHI, PpF3H, PpFLS, and PpUFGT which are key downstream genes in the flavonoid biosynthesis pathway, independently of the PpbHLH3 cofactor. These gene expression changes may enhance flavonoid biosynthesis in pear fruit. The data presented may be useful for further elucidating the flavonoid biosynthesis regulatory network, potentially leading to the development of new pear cultivars that produce fruits with increased flavonoid contents.








Similar content being viewed by others
Abbreviations
- CHS:
-
Chalcone synthase
- CHI:
-
Chalcone isomerase
- F3H:
-
Flavanone 3-hydroxylase
- ANS:
-
Anthocyanin synthase
- FLS:
-
Flavonol synthase
- UFGT:
-
UDP-glucose: flavonoid 3-glucosyltransferase
References
Almeida JRM, D’Amico E, Preuss A, Carbone F, de Vos CHR, Deiml B, Mourgues F, Perrotta G, Fischer TC, Bovy AG, Martens S, Rosati C (2007) Characterization of major enzymes and genes involved in flavonoid and proanthocyanidin biosynthesis during fruit development in strawberry (Fragaria ×ananassa). Arch Biochem Biophys 465(1):61–71. https://doi.org/10.1016/j.abb.2007.04.040
Azuma A, Yakushiji H, Koshita Y, Kobayashi S (2012) Flavonoid biosynthesis-related genes in grape skin are differentially regulated by temperature and light conditions. Planta 236(4):1067–1080
Bai S, Tao R, Tang Y, Yin L, Ma Y, Ni J, Yan X, Yang Q, Wu Z, Zeng Y, Teng Y (2019) BBX16, a B-box protein, positively regulates light-induced anthocyanin accumulation by activating MYB10 in red pear. Plant Biotechnol J 17(10):1985–1997. https://doi.org/10.1111/pbi.13114
Ballester A-R, Molthoff J, de Vos R, te Lintel HB, Orzaez D, Fernández-Moreno J-P, Tripodi P, Grandillo S, Martin C, Heldens J, Ykema M, Granell A, Bovy A (2010) Biochemical and molecular analysis of pink tomatoes: deregulated expression of the gene encoding transcription factor SlMYB12 leads to pink tomato fruit color. Plant Physiol 152(1):71–84. https://doi.org/10.1104/pp.109.147322
Blanco E, Sabetta W, Danzi D, Negro D, Passeri V, De Lisi A, Paolocci F, Sonnante G (2018) Isolation and characterization of the flavonol regulator CcMYB12 from the globe artichoke [Cynara cardunculus var. scolymus (L.) Fiori]. Front Plant Sci 9:941. https://doi.org/10.3389/fpls.2018.00941
Brahem M, Renard CMGC, Eder S, Loonis M, Ouni R, Mars M, Le Bourvellec C (2017) Characterization and quantification of fruit phenolic compounds of European and Tunisian pear cultivars. Food Res Int 95:125–133. https://doi.org/10.1016/j.foodres
Cao Z-H, Zhang S-Z, Wang R-K, Zhang R-F, Hao Y-J (2013) Genome wide analysis of the apple MYB transcription factor family allows the identification of MdMYB121 gene confering abiotic stress tolerance in plants. PLoS ONE 8(7):e69955. https://doi.org/10.1371/journal.pone.0069955
Chen Y, Yang X, He K, Liu M, Li J, Gao Z, Lin Z, Zhang Y, Wang X, Qiu X, Shen Y, Zhang L, Deng X, Ji L, Deng X-W, Chen Z, Gu H, Qu L (2006) The MYB transcription factor superfamily of Arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family. Plant Mol Biol 60(1):107–124
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16(6):735–743
Czemmel S, Stracke R, Weisshaar B, Cordon N, Harris N, Walker A, Robinson S, Bogs J (2009) The grapevine R2R3-MYB transcription factor VvMYBF1 regulates flavonol synthesis in developing grape berries. Plant Physiol 151(3):1513–1530
Delgado L, Zuniga PE, Zi P, Figueroa NE et al (2018) Application of a JA-Ile biosynthesis inhibitor to methyl jasmonate-treated strawberry fruit induces upregulation of specific MBW complex-related genes and accumulation of proanthocyanidins. Molecules 23(6):1433. https://doi.org/10.3390/molecules23061433
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15(10):573–581. https://doi.org/10.1016/j.tplants.2010.06.005
Feng S, Wang Y, Yang S, Xu Y, Chen X (2010) Anthocyanin biosynthesis in pears is regulated by a R2R3-MYB transcription factor PyMYB10. Planta 232(1):245–255
Ferreyra MLF, Rius SP, Casati P (2012) Flavonoids: biosynthesis, biological functions and biotechnological applications. Front Plant Sci 3(222):222. https://doi.org/10.3389/fpls.2012.00222
Fischer TC, Gosch C, Pfeiffer J, Halbwirth H, Halle C, Stich K, Forkmann G (2007) Flavonoid genes of pear (Pyrus communis). Trees 21:521–529
Forbes AM, Meier GP, Haendiges S, Taylor LP (2014) Structure–activity relationship studies of flavonol analogues on pollen germination. J Agric Food Chem 62(10):2175–2181. https://doi.org/10.1021/jf405688d
Gonzalez A, Zhao M, Leavitt JM, Lloyd AM (2008) Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J 53(5):814–827. https://doi.org/10.1111/j.1365-313X.2007.03373.x
Grotewold E (2006) The genetics and biochemistry of floral pigments. Annu Rev Plant Biol 57:761–780. https://doi.org/10.1146/annurev.arplant.57.032905.105248
Grotewold E, Drummond BJ, Bowen B, Peterson T (1994) The myb-homologous P gene controls phlobaphene pigmentation in maize floral organs by directly activating a flavonoid biosynthetic gene subset. Cell 76(3):543–553. https://doi.org/10.1016/0092-8674(94)90117-1
Havsteen BH (2002) The biochemistry and medical significance of the flavonoids. Pharmacol Ther 96(2–3):67–202. https://doi.org/10.1016/S0163-7258(02)00298-X
Huang W, Khaldun A, Chen J, Zhang C, Lv H, Yuan L, Wang Y (2016) A R2R3-MYB transcription factor regulates the flavonol biosynthetic pathway in a traditional Chinese medicinal plant Epimedium sagittatum. Front Plant Sci 7:1089. https://doi.org/10.3389/fpls.2016.01089
Kadomura-Ishikawa Y, Miyawaki K, Noji S, Takahashi A (2013) Phototropin 2 is involved in blue light-induced anthocyanin accumulation in Fragaria ×ananassa fruits. J Plant Res 126(6):847–857
Kobayashi S, Gotoyamamoto N, Hirochika H (2004) Retrotransposon-induced mutations in grape skin color. Science 304(5673):982–982
Lepiniec L, Debeaujon I, Routaboul J-M, Baudry A, Pourcel L, Nesi N, Caboche M (2006) Genetics and biochemistry of seed flavonoids. Annu Rev Plant Biol 57(1):405–430
Li L, Ban Z-J, Li X-H, Wu M-Y, Wang A-L, Jiang Y-Q, Jiang Y-H (2012) Differential expression of anthocyanin biosynthetic genes and transcription factor PcMYB10 in pears (Pyrus communis L.). PLoS ONE 7(9):e46070. https://doi.org/10.1371/journal.pone.0046070
Li X, Xue C, Li J, Qiao X, Li L, La Yu, Huang Y, Wu J (2016) Genome-wide identification, evolution and functional divergence of MYB transcription factors in Chinese white pear (Pyrus bretschneideri). Plant Cell Physiol 57(4):824–847. https://doi.org/10.1093/pcp/pcw029
Matsui K, Oshima Y, Mitsuda N, Sakamoto S, Nishiba Y, Walker AR, Ohme-Takagi M, Robinson SP, Yasui Y, Mori M (2018) Buckwheat R2R3 MYB transcription factor FeMYBF1 regulates flavonol biosynthesis. Plant Sci 274:466–475. https://doi.org/10.1016/j.plantsci.2018.06.025
Mehrtens F, Kranz H, Bednarek P, Weisshaar B (2005) The Arabidopsis transcription factor MYB12 is a flavonol-specific regulator of phenylpropanoid biosynthesis. Plant Physiol 138(2):1083–1096. https://doi.org/10.1104/pp.104.058032
Meissner RC, Jin H, Cominelli E, Denekamp M, Fuertes A, Greco R, Kranz HD, Penfield S, Petroni K, Urzainqui A, Martin C, Paz-Ares J, Smeekens S, Tonelli C, Weisshaar B, Baumann E, Klimyuk V, Marillonnet S, Patel K, Speulman E, Tissier AF, Bouchez D, Jones JJD, Pereira A, Wisman E, Bevan M (1999) Function search in a large transcription factor gene family in Arabidopsis: assessing the potential of reverse genetics to identify insertional mutations in R2R3 MYB genes. Plant Cell 11(10):1827–1840. https://doi.org/10.1105/tpc.11.10.1827
Nan W, Xu H, Jiang S, Zhang Z, Lu N, Qiu H, Qu C, Wang Y, Wu S, Chen X (2017) MYB12 and MYB22 play essential roles in proanthocyanidin and flavonol synthesis in red-fleshed apple (Malus sieversii f. niedzwetzkyana). Plant J 90(2):276–292. https://doi.org/10.1111/tpj.13487
Ni J, Bai S, Zhao Y, Qian M, Tao R, Yin L, Gao L, Teng Y (2019) Ethylene response factors Pp4ERF24 and Pp12ERF96 regulate blue light-induced anthocyanin biosynthesis in ‘Red Zaosu’ pear fruits by interacting with MYB114. Plant Mol Biol 99(1–2):67–78
Ni J, Zhao Y, Tao R, Yin L, Gao L, Strid Å, Qian M, Li J, Li Y, Shen J, Teng Y, Bai S (2020) Ethylene mediates the branching of the jasmonate light-induced anthocyanin biosynthhway by suppressing anthocyanin biosynthesis in red Chinese pear fruits. Plant Biotechnol J 18:1223–1240. https://doi.org/10.1111/pbi.13287
Niu Q, Li J, Cai D, Qian M, Jia H, Bai S, Hussain S, Liu G, Teng Y, Zheng X (2016) Dormancy-associated MADS-box genes and microRNAs jointly control dormancy transition in pear (Pyrus pyrifolia white pear group) flower bud. J Exp Bot 67(1):239–257. https://doi.org/10.1093/jxb/erv454
Pollastri S, Tattini M (2011) Flavonols: old compounds for old roles. Ann Bot 108(7):1225–1233. https://doi.org/10.1093/aob/mcr234
Qian M, Sun Y, Allan AC, Teng Y, Zhang D (2014) The red sport of ‘Zaosu’ pear and its red-striped pigmentation pattern are associated with demethylation of the PyMYB10 promoter. Phytochemistry 107:16–23. https://doi.org/10.1016/j.phytochem.2014.08.001
Qian M, Ni J, Niu Q, Bai S, Bao L, Li J, Sun Y, Zhang D, Teng Y (2017) Response of miR156-SPL module during the red peel coloration of bagging-treated Chinese sand pear (Pyrus pyrifolia Nakai). Front Physiol 8:550. https://doi.org/10.3389/fphys.2017.00550
Quattrocchio F, Verweij W, Koes R (2005) Flavonoids: a colorful model for the regulation and evolution of biochemical pathways. Trends Plant Sci 10(5):236–242. https://doi.org/10.1016/j.tplants.2005.03.002
Ravaglia D, Espley RV, Henry-Kirk RA, Andreotti C, Ziosi V, Hellens RP, Costa G, Allan AC (2013) Transcriptional regulation of flavonoid biosynthesis in nectarine (Prunus persica) by a set of R2R3 MYB transcription factors. BMC Plant Biol 13(1):68
Stracke R, Werber M, Weisshaar B (2001) The R2R3-MYB gene family in Arabidopsis thaliana. Curr Opin Plant Biol 4(5):447–456. https://doi.org/10.1016/S1369-5266(00)00199-0
Stracke R, Ishihara H, Huep G, Barsch A, Mehrtens F, Niehaus K, Weisshaar B (2007) Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. Plant J 50(4):660–677. https://doi.org/10.1111/j.1365-313X.2007.03078.x
Stracke R, Holtgräwe D, Schneider J, Pucker B, Sörensen TR, Weisshaar B (2014) Genome-wide identification and characterisation of R2R3-MYB genes in sugar beet (Beta vulgaris). BMC Plant Biol 14(1):249
Wang Y-R, Lu Y-F, Hao S-X, Zhang M-L, Zhang J, Tian J, Yao Y-C (2015) Different coloration patterns between the red-and white-fleshed fruits of malus crabapples. Sci Hortic 194:26–33. https://doi.org/10.1016/j.scienta.2015.07.041
Wilkins O, Nahal H, Foong J, Provart NJ, Campbell MM (2009) Expansion and diversification of the Populus R2R3-MYB family of transcription factors. Plant Physiol 149(2):981–993. https://doi.org/10.1104/pp.108.132795
Winkel-Shirley B (2002) Biosynthesis of flavonoids and effects of stress. Curr Opin Plant Biol 5(3):218–223. https://doi.org/10.1016/S1369-5266(02)00256-X
Yao G, Ming M, Allan AC, Gu C, Li L, Wu X, Wang R, Chang Y, Qi K, Zhang S, Wu J (2017) Map-based cloning of the pear gene MYB 114 identifies an interaction with other transcription factors to coordinately regulate fruit anthocyanin biosynthesis. Plant J 92:437–451. https://doi.org/10.1111/tpj.13666
Yu BY, Huang ZDZ, C, Qian M, Zheng X, Teng Y, Su J, Shu Q, (2012) Isolation of anthocyanin biosynthetic genes in red Chinese sand pear (Pyrus pyrifolia Nakai) and their expression as affected by organ/tissue, cultivar, bagging and fruit side. Sci Hortic 136(2):29–37. https://doi.org/10.1016/j.scienta.2011.12.026
Zhai R, Liu X-T, Feng W-T, Chen S-S, Xu L-F, Wang Z-G, Zhang J-L, Li P-M, Ma F-W (2014) Different biosynthesis patterns among flavonoid 3-glycosides with distinct effects on accumulation of other flavonoid metabolites in pears (Pyrus bretschneideri Rehd.). PLoS ONE. https://doi.org/10.1371/journal.pone.0091945
Zhai R, Wang Z, Zhang S, Meng G, Song L, Wang Z, Li P, Ma F, Xu L (2015) Two MYB transcription factors regulate flavonoid biosynthesis in pear fruit (Pyrus bretschneideri Rehd.). J Exp Bot 67(5):1275–1284. https://doi.org/10.1093/jxb/erv524
Zhai R, Zhao Y, Wu M, Yang J, Li X, Liu H, Wu T, Liang F, Yang C, Wang Z, Ma F, Xu L (2019) The MYB transcription factor PbMYB12b positively regulates flavonol biosynthesis in pear fruit. BMC Plant Biol 19:85. https://doi.org/10.1186/s12870-019-1687-0
Zhang J, Subramanian S, Stacey G, Yu O (2009) Flavones and flavonols play distinct critical roles during nodulation of Medicago truncatula by Sinorhizobium meliloti. Plant J 57(1):171–183. https://doi.org/10.1111/j.1365-313X.2008.03676.x
Zhang D, Yu B, Bai J, Qian M, Shu Q, Su J, Teng Y (2012) Effects of high temperatures on UV-B/visible irradiation induced postharvest anthocyanin accumulation in ‘Yunhongli No. 1’(Pyrus pyrifolia Nakai) pears. Sci Hortic 134:53–59. https://doi.org/10.1016/j.scienta.2011.10.025
Zoratti L, Karppinen K, Luengo Escobar A, Häggman H, Jaakola L (2014) Light-controlled flavonoid biosynthesis in fruits. Front Plant Sci 5:534. https://doi.org/10.3389/fpls.2014.00534
Acknowledgements
This study was supported by the National Natural Science Foundation of China (Grant Nos. 31901986 to JN and 31772272 to SB) and the Earmarked Fund for China Agriculture Research System (CARS-28).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2020_3473_MOESM1_ESM.tif
Supplementary Figure S1. PpMYB17 was unable to activate PpMYB10 promoter. a PpMYB17 could not activate the promoter of PpMYB10 in dual-luciferase assay. b Yeast one hybrid assay of PpMYB17 and the promoter of PpMYB10 (TIF 9719 kb)
Rights and permissions
About this article
Cite this article
Premathilake, A.T., Ni, J., Bai, S. et al. R2R3-MYB transcription factor PpMYB17 positively regulates flavonoid biosynthesis in pear fruit. Planta 252, 59 (2020). https://doi.org/10.1007/s00425-020-03473-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-020-03473-4