Abstract
Main Conclusion
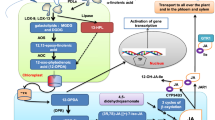
The SmERF6, which recognizes the GCC-box of SmCPS1 and SmKSL1 promoter in nucleus, regulates the tanshinone biosynthesis in Salvia miltiorrhiza hairy roots.
Tanshinone, an important medicinal ingredient in Salvia miltiorrhiza, is best known for its use in medicine. However, the transcription factor regulation of tanshinone biosynthesis is unclear. Here, we isolated and identified a transcription factor in the ERF family of S. miltiorrhiza, SmERF6, which was screened from an S. miltiorrhiza cDNA library by the promoters of two key tanshinone synthesis genes (SmKSL1 and SmCPS1); this factor regulated tanshinone biosynthesis. The gene was highly expressed in the root and responded to ethylene treatment. SmERF6 modulated tanshinone biosynthesis by directly binding to an ethylene-responsive element (GCC-box) of the SmKSL1 and SmCPS1 promoters and activating their transcription. Overexpression of SmERF6 in the hairy roots increased their tanshinone accumulation, and SmERF6 silencing by RNAi led to a lower tanshinone content. Furthermore, tanshinone accumulation maintained homeostasis with the total phenolic acid and flavonoid contents in S. miltiorrhiza. These findings elucidated how SmERF6 directly co-regulates the transcription of SmCPS1 and SmKSL1 and modulates tanshinone synthesis to accelerate the metabolic flux of tanshinone accumulation in S. miltiorrhiza.






Similar content being viewed by others
Abbreviations
- SmCPS1 :
-
Salvia miltiorrhiza copalyl diphosphate synthases 1
- SmKSL1 :
-
Salvia miltiorrhiza kaurene synthase like 1
- CT:
-
Cryptotanshinone
- DT-I:
-
Dihydrotanshinone
- ERF:
-
Ethylene response factor
- T-I (IIA):
-
Tanshinone I (IIA)
References
Bai Z, Xia P, Wang R, Jiao J, Ru M, Liu J, Liang Z (2017) Molecular cloning and characterization of five SmGRAS genes associated with tanshinone biosynthesis in Salvia miltiorrhiza hairy roots. PLoS ONE 12(9):e0185322. https://doi.org/10.1371/journal.pone.0185322
Blouin M, Mathieu J, Leadley PW (2012) Plant homeostasis, growth and development in natural and artificial soils. Ecol Complex 9(2):10–15
Cao ZF, Li J, Chen F, Li YQ, Zhou HM, Liu Q (2001) Effect of two conserved amino acid residues on DREB1A function. Biochem Biokhimiia 66(6):623–627. https://doi.org/10.1023/A:1010251129429
Claeys H, Skirycz A, Maleux K, Inzé D (2012) DELLA signaling mediates stress-induced cell differentiation in Arabidopsis leaves through modulation of anaphase-promoting complex/cyclosome activity. Plant Physiol 159(2):739–747
Cui GH, Duan LX, Jin B, Qian J, Xue Z, Shen G, Snyder JH, Song J, Chen S, Huang L, Peters RJ, Qi XQ (2015) Functional divergence of diterpene syntheses in the medicinal plant Salvia miltiorrhiza. Plant Physiol 169:1607–1618
Devers EA, Teply J, Reinert A, Gaude N, Krajinski F (2013) An endogenous artificial microRNA system for unraveling the function of root endosymbioses related genes in Medicago truncatula. BMC Plant Biol 13:82. https://doi.org/10.1186/1471-2229-13-82
Dong Y, Morris-Natschke SL, Lee KH (2011) Biosynthesis, total syntheses, and antitumor activity of tanshinones and their analogs as potential therapeutic agents. Nat Prod Rep 28(3):529–542
Druege U, Franken P, Hajirezaei MR (2016) Plant hormone homeostasis, signaling, and function during adventitious root formation in cuttings. Front Plant Sci 7(133):381. https://doi.org/10.3389/fpls.2016.00381
Dubois M, Van dBL, Claeys H, Van VK, Matsui M, Inzé D (2015) The ETHYLENE RESPONSE FACTORs ERF6 and ERF11 antagonistically regulate mannitol-induced growth inhibition in Arabidopsis. Plant Physiol 169(1):166–179
Fujimoto SY, Ohta M, Usui A, Shinshi H, Ohme-Takagi M (2000) Arabidopsis ethylene-responsive element binding factors act as transcriptional activators or repressors of GCC box-mediated gene expression. Plant Cell 12(3):393–404
Gao W, Hu TY, Guo J, Lv DM, Dai ZB, Zhou YJ, Huang LQ (2015) Research progress of synthetic biology for tanshinones. China J Chin Mater Med 40(13):2486–2491
Gu L, Han ZX, Zhang LF, Downie B, Zhao TY (2013) Functional analysis of the 5′ regulatory region of the maize GALACTINOL SYNTHASE2 gene. Plant Sci 213(3):38–45
Guo BJ, Wei YF, Xu RB, Lin S, Luan HY, Lv C, Zhang XZ, Song XY, Xu RG (2016) Genome-wide analysis of APETALA2/ethylene-responsive factor (AP2/ERF) gene family in barley (Hordeum vulgare L.). PLoS ONE 11(9):e0161322. https://doi.org/10.1371/journal.pone.0161322
Hua W, Zhang Y, Song J, Zhao L, Wang Z (2011) De novo transcriptome sequencing in Salvia miltiorrhiza to identify genes involved in the biosynthesis of active ingredients. Genomics 98(4):272–279
Kai G, Xu H, Zhou C, Pan L, Xiao J, Luo X, You L, Zhang L (2011) Metabolic engineering tanshinone biosynthetic pathway in Salvia miltiorrhiza hairy root cultures. Metab Eng 13(3):319–327
Li X (2011) A transient expression assay using Arabidopsis mesophyll protoplasts. Bio-protocol/BIO 101:e70. https://doi.org/10.21769/bioprotoc.70
Li X, Xue ZP, Zhu WX (2011) Antioxidant activities and contents of total flavonoids and phenols from different parts of Salvia miltiorrhiza bunge. Food Sci 32(3):108–111
Li B, Wang B, Li H, Liang P, Mei R, Liang Z, Yan X, Zhu Y (2016a) Establishment of Salvia castanea Diels f. tomentosa Stib. hairy root cultures and the promotion of tanshinone accumulation and gene expression with Ag+, methyl jasmonate, and yeast extract elicitation. Protoplasma 253(1):87–100
Li X, Wang J, Lei M, Li L, Fu Y, Wang Z, Ao M, Li Z (2016b) Transcriptome analysis of storage roots and fibrous roots of the traditional medicinal herb Callerya speciosa (Champ.) ScHot. PLoS ONE 11(8):e0160338. https://doi.org/10.1371/journal.pone.0160338
Li B, Cui GH, Shen GA, Zhan ZL, Huang LQ, Chen JC, Qi XQ (2017) Targeted mutagenesis in the medicinal plant Salvia miltiorrhiza. Sci Rep 7:43320. https://doi.org/10.1038/srep43320
Liu Y, Zhang S (2004) Phosphorylation of 1-aminocyclopropane-1-carboxylic acid synthase by MPK6, a stress-responsive mitogen-activated protein kinase, induces ethylene biosynthesis in Arabidopsis. Plant Cell 16(12):3386–3399
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(− Delta Delta C(T)) method. Methods 25(4):402–408
Ludidi NN (2004) Characterization of two Arabidopsis thaliana genes with roles in plant homeostasis. Dissertation, Cape Town University of the Western Cape. http://hdl.handle.net/11394/14602
Nakano T, Suzuki K, Fujimura T, Shinshi H (2006) Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol 140(2):411–432
Ohme-Takagi M, Shinshi H (1995) Ethylene-inducible DNA binding proteins that interact with an ethylene-responsive element. Plant Cell 7(2):173–182
Ramadan AM, Eissa HF, El-Domyati FM, Saleh OM, Ibrahim NE, Salama M, Mahfouz MM, Bahieldin A (2011) Characterization of inhibitor(s) of β-glucuronidase enzyme activity in GUS -transgenic wheat. Plant Cell Tissue Organ Cult 107(3):373–381
Riechmann JL, Meyerowitz EM (1998) The AP2/EREBP family of plant transcription factors. Biol Chem 379(6):633–646
Scarpeci TE, Frea VS, Zanor MI, Valle EM (2017) Overexpression of AtERF019 delays plant growth and senescence and improves drought tolerance in Arabidopsis. J Exp Bot 68(3):673–685
Song CP, Agarwal M, Ohta M, Guo Y, Halfter U, Wang P, Zhu JK (2005) Role of an Arabidopsis AP2/EREBP-type transcriptional repressor in abscisic acid and drought stress responses. Plant Cell 17(8):2384–2396
Stork M (2008) Genetic analysis reveals that C19-GA 2-oxidation is a major gibberellin inactivation pathway in Arabidopsis. Plant Cell 20(9):2420–2436
Wan L, Zhang J, Zhang H, Zhang Z, Quan R, Zhou S, Huang R (2011) Transcriptional activation of OsDERF1 in OsERF3 and OsAP2-39 negatively modulates ethylene synthesis and drought tolerance in rice. PLoS ONE 6(9):e25216. https://doi.org/10.21371/journal.pone.0025216
Wang DL (1999) CO2 enrichment and allelopathy. Acta Ecol Sin 19(1):122–127
Wang JW, Wu JY (2010) Tanshinone biosynthesis in Salvia miltiorrhiza and production in plant tissue cultures. Appl Microbiol Biotechnol 88(2):437–449
Wang X, Morrisnatschke SL, Lee KH (2007) New developments in the chemistry and biology of the bioactive constituents of tanshen. Med Res Rev 27(1):133–148
Wang P, Du Y, Zhao X, Miao Y, Song CP (2013) The MPK6–ERF6–ROS-responsive cis-acting element 7/GCC box complex modulates oxidative gene transcription and the oxidative response in Arabidopsis. Plant Physiol 161(3):1392–1408
Xu Z, Peters RJ, Weirather J, Luo H, Liao B, Zhang X, Zhu Y, Ji A, Zhang B, Hu S (2015) Full-length transcriptome sequences and splice variants obtained by a combination of sequencing platforms applied to different root tissues of Salvia miltiorrhiza and tanshinone biosynthesis. Plant J 82(6):951–961
Xu HB, Song JY, Luo HM, Zhang YJ, Li QM et al (2016) Analysis of the genome sequence of the medicinal plant Salvia miltiorrhiza. Mol Plant 9(6):949–952
Yan XF, Yang W (2007) Plant secondary metabolism and its response to environment. Acta Ecol Sin 27(6):2554–2562
Yang D, Ma P, Liang X, Wei Z, Liang Z, Liu Y, Liu F (2012) PEG and ABA trigger methyl jasmonate accumulation to induce the MEP pathway and increase tanshinone production in Salvia miltiorrhiza hairy roots. Physiol Plant 146(2):173–183
Zhao X, Zheng X, Fan T, Li Z, Zhang Y, Zheng J (2015) A novel drug discovery strategy inspired by traditional medicine philosophies. Science 347(6219 Suppl):S38–S40
Zhou L, Dong C, Liu J (2011) Increased resistance of Arabidopsis to cold and salt stresses by suppressing the transcription repressors of the A-5 group among the DREB subfamily transcription factors through artificial microRNA. China Biotechnol 31(5):34–41
Acknowledgements
We are grateful to professors Chao Sun, Shilin Chen and Weibo Jin for kindly providing the genome resources and transcriptome data of S. miltiorrhiza used in this study, respectively. We thanks for the supports from the National Natural Science Foundation of China (81773835, 81373908 and 81373536).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have declared no conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2018_2884_MOESM2_ESM.tif
Supplementary material 2 Fig. S2 The minimal inhibitory concentration of Aureobasidin A (AbA) screens for the bait strain of the SmCPS1 and SmKSL1 promoter (TIFF 3681 kb)
425_2018_2884_MOESM3_ESM.tif
Supplementary material 3 Fig. S3 The Western-blotting analysis of MBP and SmERF6 and the precursor microRNA structure of amiR159b-SmERF6 (TIFF 1532 kb)
Rights and permissions
About this article
Cite this article
Bai, Z., Li, W., Jia, Y. et al. The ethylene response factor SmERF6 co-regulates the transcription of SmCPS1 and SmKSL1 and is involved in tanshinone biosynthesis in Salvia miltiorrhiza hairy roots. Planta 248, 243–255 (2018). https://doi.org/10.1007/s00425-018-2884-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-018-2884-z