Abstract
The AP1/FUL clade of MADS box genes have undergone multiple duplication events among angiosperm species. While initially identified as having floral meristem identity and floral organ identity function in Arabidopsis, the role of AP1 homologs does not appear to be universally conserved even among eudicots. In comparison, the role of FRUITFULL has not been extensively explored in non-model species. We report on the isolation of three AP1/FUL genes from cultivated spinach, Spinacia oleracea L. Two genes, designated SpAPETALA1-1 (SpAP1-1) and SpAPETALA1-2 (SpAP1-2), cluster as paralogous genes within the Caryophyllales AP1 clade. They are highly differentiated in the 3′, carboxyl-end encoding region of the gene following the third amphipathic alpha-helix region, while still retaining some elements of a signature AP1 carboxyl motifs. In situ hybridization studies also demonstrate that the two paralogs have evolved different temporal and spatial expression patterns, and that neither gene is expressed in the developing sepal whorl, suggesting that the AP1 floral organ identity function is not conserved in spinach. The spinach FRUITFULL homolog, SpFRUITFULL (SpFUL), has retained the conserved motif and groups with Caryophyllales FRUITFULL homologs. SpFUL is expressed in leaf as well as in floral tissue, and shows strong expression late in flower development, particularly in the tapetal layer in males, and in the endothecium layer and stigma, in the females. The combined evidence of high rates of non-synonymous substitutions and differential expression patterns supports a scenario in which the AP1 homologs in the spinach AP1/FUL gene family have experienced rapid evolution following duplication.






Similar content being viewed by others
Abbreviations
- AP1 :
-
APETALA1 gene
- Fl :
-
FRUITFULL-LIKE gene
- FUL :
-
FRUITFULL gene
References
Anthony RG, James PE, Jordan BR (1995) The cDNA sequence of a cauliflower apetala-1/squamosa homolog. Plant Physiol 108:441–442
Beales J, Laurie DA, Devos KM (2005) Allelic variation at the linked AP1 and PhyC loci in hexaploid wheat is associated but not perfectly correlated with vernalization response. Theor Appl Genet 110:1099–1107
Berbel A, Navarro C, Ferrándiz C, Cañas LA, Madueño F, Beltrán J-P (2001) Analysis of PEAM4, the pea AP1 functional homologue, supports a model for AP1-like genes controlling both floral meristem and floral organ identity in different plant species. Plant J 25:441–451
Bowman JL, Alvarez J, Weigel D, Meyerowitz EM, Smyth DR (1993) Control of flower development in Arabidopsis thaliana by APETALA1 and interacting genes. Development 119:721–743
Bowman JL, Drews GN, Meyerowitz EM (1991) Expression of the Arabidopsis floral homeotic gene AGAMOUS is restricted to specific cell types late in flower development. Plant Cell 3:749–758
Bowman JL, Smyth DR, Meyerowitz EM (1989) Genes directing flower development in Arabidopsis. Plant Cell 1:37–52
Causier B, Castillo R, Zhou J, Ingram R, Xue Y, Schwarz-Sommer Z, Davies B (2005) Evolution in action: following function in duplicated floral homeotic genes. Curr Biol 15:1508–1512
Davies B, Cartolano M, Schwarz-Sommer Z (2006) Flower development: the Antirrhinum perspective. Adv Bot Res 44:279–321
Davies B, Motte P, Keck E, Saedler H, Sommer H, Schwarz-Sommer Z (1999) PLENA and FARINELLI: redundancy and regulatory interactions between two Antirrhinum MADS-box factors controlling flower development. EMBO J 18:4023–4034
Devon RS, Porteous DJ, Brookes AJ (1995) Splinkerettes-improved vectorettes for greater efficiency in PCR walking. Nucleic Acids Res 23:1644–1645
Drea S, Hileman LC, de Martino G, Irish VF (2007) Functional analyses of genetic pathways controlling petal specification in poppy. Development 134:4157–4166
Elo A, Lemmetyinen J, Turunen ML, Tikka L, Sopanen T (2001) Three MADS-box genes similar to APETALA1 and FRUITFULL from silver birch (Betula pendula). Physiol Plant 112:95–103
Ferrandiz C, Gu Q, Martienssen R, Yanofsky MF (2000a) Redundant regulation of meristem identity and plant architecture by FRUITFULL, APETALA1 and CAULIFLOWER. Development 127:725–734
Ferrandiz C, Liljegren SJ, Yanofsky MF (2000b) Negative regulation of the SHATTERPROOF genes by FRUITFULL during Arabidopsis fruit development. Science 289:436–438
Ferrario S, Busscher J, Franken J, Gerats T, Vandenbussche M, Angenent GC, Immink RGH (2004) Ectopic expression of the Petunia MADS box gene UNSHAVEN accelerates flowering and confers leaf-like characteristics to floral organs in a dominant-negative manner. Plant Cell 16:1490–1505
Gu Q, Ferrandiz C, Yanofsky MF, Martienssen R (1998) The FRUITFULL MADS-box gene mediates cell differentiation during Arabidopsis fruit development. Development 125:1509–1517
Guindon S, Gascuel O (2003) A simple, fast and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Gustafson-Brown C, Savidge B, Yanofsky MF (1994) Regulation of the Arabidopsis homeotic gene APETALA1. Cell 76:131–143
Hardenack S, Ye D, Saedler H, Grant S (1994) Comparison of MADS box gene expression in developing male and female flowers of the dioecious plant White Campion. Plant Cell 6:1775–1787
Hongyan S, Kunmei S, Wenliang L, Hongzhi K, Zhiduan C, Zheng M (2006) Conservation and divergence of candidate class B genes in Akebia trifoliata (Lardizabalaceae). Dev Gene Evol V216:785–795
Honma T, Goto K (2001) Complexes of MADS-box proteins are sufficient to convert leaves into floral organs. Nature 409:525–529
Huijser P, Klein J, Lönnig W, Meijer H, Saedler H, Sommer H (1992) Bracteomania, an inflorescence anomaly, is caused by the loss of function of the MADS-box gene squamosa in Antirrhinum majus. EMBO J 11:1239–1249
Irish VF (2003) The evolution of floral homeotic gene function. Bioessays 25:637–646
Jack T (2004) Molecular and genetic mechanisms of floral control. Plant Cell 16:S1–S17
Katoh K, Kuma K-I, Toh H, Miyata T (2005) MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res 33:511–518
Kramer EM, Jaramillo MA, Di Stilio VS (2004) Patterns of gene duplication and functional evolution during the diversification of the AGAMOUS subfamily of MADS box genes in Angiosperms. Genetics 166:1011–1023
Krizek BA, Meyerowitz EM (1996) Mapping the protein regions responsible for the functional specificities of the Arabidopsis MADS domain organ-identity proteins. Proc Natl Acad Sci USA 93:4063–4070
Kumar S, Tamura K, Nei M (2004) MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Briefings Bioinf 5:150–163
Kyozuka J, Konishi S, Nemoto K, Izawa T, Shimamoto K (1998) Down-regulation of RFL, the FLO/LFY homolog of rice, accompanied with panicle branch initiation. Proc Natl Acad Sci USA 95:1979–1982
Lee H, Suh SS, Park E, Cho E, Ahn JH, Kim SG, Lee JS, Kwon YM, Lee I (2000) The AGAMOUS-LIKE 20 MADS domain protein integrates floral inductive pathways in Arabidopsis. Genes Dev 14:2366–2376
Lee J-Y, Baum SF, Alvarez J, Patel A, Chitwood DH, Bowman JL (2005) Activation of CRABS CLAW in the nectaries and carpels of Arabidopsis. Plant Cell 17:25–36
Liljegren SJ, Roeder AHK, Kempin SA, Gremski K, Ostergaard L, Guimil S, Reyes DK, Yanofsky MF (2004) Control of fruit patterning in Arabidopsis by indehiscent. Cell 116:843–853
Litt A (2007) An evaluation of A-function: evidence from the APETALA1 and APETALA2 gene lineages. Int J Plant Sci 168:73–91
Litt A, Irish VF (2003) Duplication and diversification in the APETALA1/FRUITFULL floral homeotic gene lineage: implications for the evolution of floral development. Genetics 165:821–833
Liu Z, Meyerowitz EM (1995) LEUNIG regulates AGAMOUS expression in Arabidopsis flowers. Development 121:975–991
Mandel MA, Yanofsky MF (1995) The Arabidopsis AGL8 MADS box gene is expressed in inflorescence meristems and is negatively regulated by APETALA1. Plant Cell 7:1763–1771
Mandel MA, Bowman JL, Kempin SA, Ma H, Meyerowitz EM, Yanofsky MF (1992) Manipulation of flower structure in transgenic tobacco. Cell 71:133–143
Matsunaga S, Isono E, Kejnovsky E, Vyskot B, Dolezel J, Kawano S, Charlesworth D (2003) Duplicative transfer of a MADS box gene to a plant Y chromosome. Mol Biol Evol 20:1062–1069
Mellerowicz EJ, Horgan K, Walden A, Coker A, Walter C (1998) PRFLL—a Pinus radiata homologue of FLORICAULA and LEAFY is expressed in buds containing vegetative shoot and undifferentiated male cone primordia. Planta 206:619–629
Müller BM, Saedler H, Zachgo S (2001) The MADS-box gene DEFH28 from Antirrhinum is involved in the regulation of floral meristem identity and fruit development. Plant J 28:169–179
Murai K, Miyamae M, Kato H, Takumi S, Ogihara Y (2003) WAP1, a wheat APETALA1 homolog, plays a central role in the phase transition from vegetative to reproductive growth. Plant Cell Physiol 44:1255–1265
Nawy T, Lee J-Y, Colinas J, Wang JY, Thongrod SC, Malamy JE, Birnbaum K, Benfey PN (2005) Transcriptional profile of the Arabidopsis root quiescent center. Plant Cell 17:1908–1925
Ng M, Yanofsky MF (2001) Activation of the Arabidopsis B class homeotic genes by APETALA1. Plant Cell 13:739–753
Pelaz S, Ditta GS, Baumann E, Wisman E, Yanofsky MF (2000) B and C floral organ identity functions require SEPALLATA MADS-box genes. Nature 405:200–203
Pelaz S, Gustafson-Brown C, Kohalmi SE, Crosby WL, Yanofsky MF (2001) APETALA1 and SEPALLATA3 interact to promote flower development. Plant J 26:385–394
Pfent C, Pobursky KJ, Sather DN, Golenberg EM (2005) Characterization of SpAPETALA3 and SpPISTILLATA, B class floral identity genes in Spinacia oleracea, and their relationship to sexual dimorphism. Dev Gene Evol 215:132–142
Pinyopich A, Ditta GS, Savidge B, Liljegren SJ, Baumann E, Wisman E, Yanofsky MF (2003) Assessing the redundancy of MADS-box genes during carpel and ovule development. Nature 424:85–88
Preston JC, Kellogg EA (2006) Reconstructing the evolutionary history of paralogous APETALA1/FRUITFULL-like genes in grasses (Poaceae). Genetics 174:421–437
Rajani S, Sundaresan V (2001) The Arabidopsis myc/bHLH gene ALCATRAZ enables cell separation in fruit dehiscence. Curr Biol 11:1914–1922
Riechmann JL, Krizek BA, Meyerowitz EM (1996a) Dimerization specificity of Arabidopsis MADS domain homeotic proteins APETALA1, APETALA3, PISTILLATA, and AGAMOUS. Proc Natl Acad Sci USA 93:4793–4798
Riechmann JL, Wang M, Meyerowitz EM (1996b) DNA-binding properties of Arabidopsis MADS domain homeotic proteins APETALA1, APETALA3, PISTILLATA and AGAMOUS. Nucleic Acids Res 24:3134–3141
Rijpkema AS, Gerats T, Vandenbussche M (2007) Evolutionary complexity of MADS complexes. Curr Opin Plant Biol 10:32–38
Robles P, Pelaz S (2005) Flower and fruit development in Arabidopsis thaliana. Int J Dev Biol 49:633–643
Saedler H, Becker A, Winter KU, Kirchner C, Theissen G (2001) MADS-box genes are involved in floral development and evolution. Acta Biochim Pol 48:351–358
Saedler H, Huijser P (1993) Molecular biology of flower development in Antirrhinum majus (snapdragon). Gene 135:239–243
Sather DN, York A, Pobursky KJ, Golenberg EM (2005) Sequence evolution and sex-specific expression patterns of the C class floral identity gene, SpAGAMOUS, in dioecious Spinacia oleracea L. Planta 222:284–292
Shchennikova AV, Shulga OA, Immink R, Skryabin KG, Angenent GC (2004) Identification and characterization of four chrysanthemum MADS-box genes, belonging to the APETALA1/FRUITFULL and SEPALLATA3 subfamilies. Plant Physiol 134:1632–1641
Soltis DE, Ma H, Frohlich MW, Soltis PS, Albert VA, Oppenheimer DG, Altman NS, dePamphilis C, Leebens-Mack J (2007) The floral genome: an evolutionary history of gene duplication and shifting patterns of gene expression. Trends Plant Sci 12:358–367
Song I-J, Nakamura T, Fukuda T, Yokoyama J, Ito T, Ichikawa H, Horikawa Y, Kameya T, Kanno A (2006) Spatiotemporal expression of duplicate AGAMOUS orthologues during floral development in Phalaenopsis. Dev Gene Evol 216:301–313
Sridhar VV, Surendrarao A, Gonzalez D, Conlan RS, Liu Z (2004) Transcriptional repression of target genes by LEUNIG and SEUSS, two interacting regulatory proteins for Arabidopsis flower development. Proc Natl Acad Sci USA 101:11494–11499
Sridhar VV, Surendrarao A, Liu Z (2006) APETALA1 and SEPALLATA3 interact with SEUSS to mediate transcription repression during flower development. Develpment 133:3159–3166
Strimmer K, Haeseler AV (1996) Quartet puzzling: a quartet maximum likelihood method for reconstructing tree topologies. Mol Biol Evol 13:964–969
Sung S-K, Yu G-H, Nam J, Jeong D-H, An G (2000) Developmentally regulated expression of two MADS-box genes, MdMADS3 and MdMADS4, in the morphogenesis of flower buds and fruits in apple. Planta 210:519–528
Theissen G, Melzer R (2007) Molecular mechanisms underlying origin and diversification of the Angiosperm flower. Ann Bot 100:603–619
Tilly JJ, Allen DW, Jack T (1998) The CArG boxes in the promoter of the Arabidopsis floral organ identity gene APETALA3 mediate diverse regulatory effects. Development 125:1647–1657
Vandenbussche M, Theissen G, Van de Peer Y, Gerats T (2003) Structural diversification and neo-functionalization during floral MADS-box gene evolution by C-terminal frameshift mutations. Nucleic Acids Res 31:4401–4409
Vandenbussche M, Zethof J, Royaert S, Weterings K, Gerats T (2004) The duplicated B-class heterodimer model: Whorl-specific effects and complex genetic interactions in Petunia hybrida flower development. Plant Cell 16:741–754
Yang Z (2007) PAML 4: a program package for phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Yang Z, Nielsen R (2000) Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol Biol Evol 17:32–43
Yu D, Kotilainen M, Pollanen E, Mehto M, Elomaa P, Helariutta Y, Albert VA, Teeri TH (1999) Organ identity genes and modified patterns of flower development in Gerbera hybrida (Asteraceae). Plant J 17:51–62
Zahn LM, Leebens-Mack JH, Arrington JM, Hu Y, Landherr LL, de Pamphilis CW, Becker A, Theissen G, Ma H (2006) Conservation and divergence in the AGAMOUS subfamily of MADS-box genes: evidence of independent sub- and neofunctionalization events. Evol Dev 8:30–45
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2008_851_MOESM1_ESM.pdf
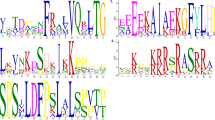
Clustal –like alignment of translated amino acid sequences from FUL gene clade. The asterisks (*) indicate identical residues at a position, whereas the colon (:) indicates chemically similar residues at a position. The dashes (-) in the sequences indicate gaps inserted for alignment purposes (PDF 32 kb)
425_2008_851_MOESM2_ESM.pdf
Clustal –like alignment of translated amino acid sequences from AP1 gene clade. The asterisks (*) indicate identical residues at a position, whereas the colon (:) indicates chemically similar residues at a position. The dashes (-) in the sequences indicate gaps inserted for alignment purposes (PDF 33 kb)
425_2008_851_MOESM3_ESM.jpg
Unrooted phylogenetic gene trees of FUL and AP1 subtrees based on nucleotide and amino acid sequences of the variable C region alone. Data alignment was hypothesize within each subclades. a Neighbor-joining tree of the FUL subclade using the C region nucleotide sequences. b Mimimum-linkage tree of the FUL subclade using the translated C region nucleotide sequences. c Neighbor-joining tree of the AP1 subclade using the C region nucleotide sequences. d Neighbor-joining tree of the AP1 subclade using the translated C region nucleotide sequences
Rights and permissions
About this article
Cite this article
Sather, D.N., Golenberg, E.M. Duplication of AP1 within the Spinacia oleracea L. AP1/FUL clade is followed by rapid amino acid and regulatory evolution. Planta 229, 507–521 (2009). https://doi.org/10.1007/s00425-008-0851-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-008-0851-9