Abstract
Liver regeneration may take place after liver injury through replication of hepatocytes or hepatic progenitor cells called oval cells. Interferons (IFN) are natural cytokines with pleiotrophic effects including antiviral and antiproliferative actions. No data are yet available on the physiology and cellular source of natural IFNs during liver regeneration. To address this issue, we have analyzed the levels and biologic activities of IFN-α/IFN-γ in two models of partial hepatectomy. After 2/3rd partial hepatectomy (PH), hepatic levels of IFN-α and IFN-γ declined transiently in contrast to a transient increase of the IFN-γ serum level. After administration of 2-acetylaminofluorene and partial hepatectomy (AAF/PH model), however, both IFN-α and IFN-γ expression were up-regulated in regenerating livers. Again, the IFN-γ serum level was transiently increased. Whereas hepatic IFN-γ was up-regulated early (day 1–5), but not significantly, in the AAF/PH model, IFN-α was significantly up-regulated at later time points in parallel to the peak of oval cell proliferation (days 7–9). Biological activity of IFN-α was shown by activation of IFN-α-specific signal transduction and induction of IFN-α specific-gene expression. We found a significant infiltration of the liver with inflammatory monocyte-like mononuclear phagocytes (MNP) concomitant to the frequency of oval cells. We localized IFN-α production only in MNPs, but not in oval cells. These events were not observed in normal liver regeneration after standard PH. We conclude that IFN-γ functions as an acute-phase cytokine in both models of liver regeneration and may constitute a systemic component of liver regeneration. IFN-α was increased only in the AAF/PH model, and was associated with proliferation of oval cells. However, oval cells seem not to be the source of IFN-α. Instead, inflammatory MNP infiltrating AAF/PH-treated livers produce IFN-α. These inflammatory MNPs may be involved in the regulation of the oval cell compartment through local expression of cytokines, including IFN-α.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Liver regeneration is a unique biological process in which partial organ loss can be compensated by a coordinated proliferation of remaining viable liver cells. Normally, hepatocytes are a quiescent cell population, but they may proliferate under various pathophysiological conditions, including viral or toxic hepatic injury or partial hepatectomy (PH). However, if the capacity of hepatocytes to replicate is severely impaired, liver tissue may still be replaced by a facultative progenitor cell compartment (“oval cells”). In rodents, proliferation of oval cells can be induced by exposure to hepatocarcinogens (e.g., 2-acetyl-aminofluorene, AAF) and by subsequently submitting the animals to proliferative stimuli, e.g., PH. Although this is an alternative regeneration pathway when mature hepatocytes fail to replicate, there is now increasing evidence that activation of these progenitor cells may also give rise to hepatocellular carcinoma (HCC) (Dumble et al. 2002; Braun et al. 1989).
The cell-signaling pathways that characterize alternative regeneration pathways are mainly unknown (Batusic et al. 2005). In particular, very few factors have yet been identified that trigger oval cell-driven liver regeneration. Interferon-gamma (IFN-γ) was suggested as one of these factors as its responsive gene program is activated in oval cell-driven liver regeneration (Bisgaard et al. 1999). IFN-γ was shown to inhibit the proliferation of hepatocytes but to promote the proliferation of oval cells (Brooling et al. 2005). The oval cell response is attenuated in mice deficient in IFN-γ gene expression (Akhurst et al. 2005). As it has been recently shown, its promotion of oval cells seems to be rather indirect upon liver injury by stimulating inflammation and aggravating liver damage (Knight et al. 2007).
Administration of IFN-α results in impaired rat liver regeneration after PH by inhibition of hepatocyte replication and total protein synthesis (Theocharis et al. 1995; Theocharis et al. 1997; Nishiguchi et al. 1986; Favre et al. 2001). When IFN-α is administered in rats after AAF/PH treatment, the frequency of oval cells and the incidence of HCCs are reduced (Nakaji et al. 2004; Alvarez et al. 2001).
Apparently, the absence of IFN-γ or the addition of IFN-α/-γ have significant effects on liver regeneration. However, no data are yet available on the physiology and cellular source of natural IFNs during liver regeneration. To address this issue, we have analyzed the levels and biological activities of IFN-α/IFN-γ in alternative regeneration pathways of the same species. In particular, we aimed to delineate the effects of an acute-phase reaction from that occurring as a result of either proliferation of hepatocytes or oval cells.
Materials and methods
Animals
Male Wistar rats, purchased from Harlan–Winkelmann (Germany), were allowed to acclimatize for 1 week prior to the start of experiments, at which time they weighed between 180 and 220 g. Food and water were available ad libitum.
Treatment of animals
Rats were fed with 1.5 mg 2-acetylaminofuorene (AAF) daily by intragastric administration, and partial hepatectomy (PH) or sham operation (SO) was performed on the 6th day. AAF was not given on the day of surgery. PH was performed under ether anesthesia by midventral laparotomy, ligation of the median anterior and left lateral hepatic lobes separately with a silk suture, and complete excision of ligated lobes. SO consisted of a midventral laparotomy of similar extent, gentle manipulation of the liver, followed by surgical closure of the abdominal wall similar to PH-treated rats. AAF treatment was restarted on the first postoperative day and continued for another 5 days to a total dose of 15 mg per rat. Rats were killed on day 1, 3, 5, 7, 11 and 13 after PH or SO (three rats per time point). A control group received only AAF without any operative procedure (AAF controls, n = 3). Livers were snap frozen in liquid nitrogen and stored at –80°C.
Standard PH or SO was performed without AAF treatment as described above. Since liver regeneration proceeds faster without AAF treatment, earlier time points of up to 7 days were chosen (three rats per time point).
This study was approved by the Animal Ethics and Research Committee of the University Medical Center Göttingen, and the experiments were conducted in accordance with the laws and regulations of the state of Lower Saxony, Germany.
Enzyme-linked immunosorbent assay
Liver samples were homogenized in ice-cold phosphate-buffered saline (PBS), pH 7.8, containing 20 mmol/L Hepes, 10 mmol/L KCl, 0.1 mmol/L ethylene-diamine-tetraacetic acid (EDTA), 1 mmol/L dithiothreitol (DTT) and 1 mmol/L phenylmethylsulfonyl fluoride (PMSF). We used six volumes of lysis buffer for determination of IFN-α in liver samples and eight volumes of buffer for determination of IFN-γ. The homogenate was centrifuged for 30 min at 20,000×g at 4°C, and the supernatant was used in the enzyme-linked immunosorbent assay (ELISA) according to the manufacturer’s instructions (Mouse IFN-α ELISA Kit, Pestka Biomedical Laboratories, New Brunswik, NJ, USA; Rat IFN-γ ELISA Kit, BioSource Europe, Nivelles, Belgium). Serum samples were analyzed undiluted according to the manufacturer’s protocol. The values of the assays were determined in pg/mL serum or pg/g frozen liver tissue, respectively.
RNA extraction, Northern blot hybridization and real-time PCR
Total RNA was extracted from rat liver and from freshly isolated and cultured cells according to Chirgwin et al. (1979), separated on agarose gel by electrophoresis, blotted onto nylon membranes and hybridized with a 32P-labeled cDNA probe for rat Mx-2 (1.1 kb cDNA). Radiolabeled oligonucleotide specific for 28S ribosomal RNA was used as a control.
For real-time PCR, 1 μg of total RNA was converted into cDNA using Superscript II RT (Invitrogen, Carlsbad, CA, USA) and oligo (dT)15 primer. The cDNA was amplified with SYBR Green Master Mix (Applied Biosystems) according to the manufacturer’s instructions in an ABI PRISM 7000 Sequence Detection System (Applied Biosystems), and relative expression was calculated as described elsewhere (Batusic et al. 2005). We used specific primer pairs for rat IFN-α (TGCAACCCTCCTAGACTCATTCT/CCCCTACCTGCTGCATCAGA), IFN-γ (GCCCTCTCTGGCTGTTACTG/CCAAGAGGAGGCTCTTTCCT), α-fetoprotein (AFP; GCCCAGCATACGAAGAAAACA/TCTCTTTGTCTGGAAGCATTCCT), cyclin D1 (GCCATCCAT GCGGAAAATC/AGAGACAAGAACCGGTCCAGGT), Mx-2 (CCCTTCAGCTAACCACTACCC/CCTGGCAGGGTTCTAAAATG), and ubiquitin c (CACCAAGAAGGTCAAACAGGAA/AAGACACCTCCCCATCAAACC) as a housekeeping gene.
In situ hybridization
In situ hybridization experiments were performed according to a protocol described by Braissant and Wahli (1998). Antisense and sense IFN-α cDNAs were synthesized by a standard PCR protocol (Invitrogen Platinum Taq) using a cDNA template from a rat liver sample of acute liver injury (24 h after CCl4 ingestion). The following primers containing the T7 promoter were used: Antisense IFN-αfor 5′-CGGCTCTGTGCTTTCTTGAT-3′; Antisense IFN-αT7rev 5′-TAATACGACTCACTATAGGG-AGTTGGCTGAGGAAGACAGG-3′; Sense IFN-αT7for 5′-TAATACGACTCACTATAGGG-CGGCTCTGTGCTTTCTTGAT-3′; Sense IFN-α rev 5′AGTTGGCTGAGGAAGACAGG-3′. The PCR products were purified by Qiagen Minielute Kit, and antisense or sense riboprobes were synthesized using the DIG labeling kit and T7 enzyme (Roche). Fixation, hybridization and washing steps were performed on liver sections of control rats and of AAF-treated rats at day 7 after PH, as described previously (Cimica et al. 2005).
Laser capture microdissection (LCM)
Frozen liver tissue specimens were cut as a series of 6 μm-thick sections with a clean blade at −20°C and mounted on RNase-free slides. Sections were processed as described earlier (Blaschke et al. 2003). Periportal and pericentral parts of the hepatic lobule were separately collected by laser microbeam microdissection and captured by laser pressure catapulting directly into the cap tubes. Three to five microdissected periportal or pericentral zones were captured on a single cap and RNA was extracted using the RNAqueous-Micro RNA isolation kit (Ambion Europe, Huntingdon, Cambridgeshire, UK) according to the manufacturer’s protocol. We synthesized cDNA using SuperScript II reverse transcriptase (Invitrogen) and performed PCR with specific primer pairs for rat IFN-α (5′-CGGCTCTGTGCTTTCTTGAT-3′ and 5′-TGCATCCCAAGCAGT–AGATG-3′) as described earlier (Batusic et al. 2004).
Immunohistochemistry
Immunohistochemical detection of proliferating cell nuclear antigen (PCNA; Novocastra, UK) and CD68/CD163 antigens (monoclonal anti CD68/clone ED1, monoclonal anti CD163/clone ED2, both from Serotec, UK) was performed by the immunoperoxidase method according to a standard protocol (McCormick et al. 1993). A polyclonal HRP-conjugated anti-mouse immunoglobulin antibody (DAKO, P0161) was used in a second reagent step. Alpha-fetoprotein (anti-AFP; DAKO) was detected by the immunoperoxidase technique using the DAKO Envision system according to the manufacturer’s instructions (DAKO, Glostrup, Denmark). Immunohistochemical detection of natural killer (NK) cells in the rat liver was performed by using the alkaline phosphatase-anti-alkaline phosphatase (APAAP) method as described previously in more detail (Batusic et al. 2005). We used a monoclonal antibody against NKp30, an NK cell receptor that is expressed in the majority of rat NK cells (clone CLH9; dilution of primary antibody 1:100; Santa Cruz Biotechnology, Santa Cruz, CA, USA). Negative controls were incubated with mouse IgM (DAKO, Denmark) instead of primary antibodies.
Western blot analysis
Lysates were prepared by homogenization at 4°C in 20 mM Tris–HCl (pH 8.0), 5 mM EDTA, 3 mM EGTA, 1 mM DTT, 1% SDS, 1 mM PMSF and protease inhibitor cocktail (Sigma-Aldrich Inc., USA). Samples were cleared by centrifugation at 15,000×g for 15 min at 4°C, and the protein concentration was measured by BCA assay (Pierce, Rockford, IL, USA), using bovine serum albumin as standard. Protein lysates were separated on SDS–polyacrylamide gels, electrotransferred to polyvinylidene difluoride membranes (Invitrogen; USA) and probed with primary antibodies overnight. The appropriate peroxidase-conjugated secondary antibodies (DAKO, Glostrup, Denmark) were then added in a dilution of 1:5,000 and incubation continued for 1 h at room temperature. Bound antibodies were visualized using chemiluminescent substrate (ECL; AmershamPharmacia, UK). Equal loading was previously controlled by transient Ponceau S staining. The primary antibodies included: mouse monoclonal anti-Mx (mAB M143, 1:500, generous gift from Dr. O. Haller, Freiburg, Germany), anti-JAK1 (Upstate Biotechnology, USA), anti-Tyk2 (C-8; Santa Cruz, USA) and anti-β-actin (clone AC-15, Sigma-Aldrich, USA).
Immunoprecipitation and SDS-PAGE analysis
Liver samples used for immunoprecipitation were lysed in NP-40 lysis buffer containing 150 mM NaCl, 1% NP-40, 50 mM Tris–HCl (pH 8.0), 1 mM PMSF, 1 mM sodium orthovanadate and an aliquot of protease inhibitor cocktail (Sigma-Aldrich Inc., USA). After insoluble material was removed by centrifugation, the lysates were incubated with 5 μg of antibody for 1 h at 4°C. The following antibodies were used (all from Upstate Biotechnology, USA): anti-Stat1, anti-Stat2, anti-Stat3, and corresponding phospho-specific antibodies. The formed antibody–antigen complexes were precipitated using protein G Sepharose beads (AmerhamPharmacia, UK) and washed several times before being redissolved in 20 μL SDS-PAGE sample buffer. Electrophoresis, transfer and immunoblotting were carried out according to our protocols for Western blotting. HepG2 cells treated with IFN-α (500 U/mL) were used as positive controls after precipitation with Stat-1 or Stat-2.
Statistical analysis
The results are expressed as mean ± SEM. Significance in differences was tested by Student’s t test, and p < 0.05 was considered significant.
Results
Two models of liver regeneration
To study hepatocyte-driven liver regeneration, we performed PH or SO in rats without AAF treatment and monitored weight and PCNA staining. Liver weight was almost restored at day 7 after PH (90% ± 5%). PCNA staining index already started to increase 8 h after PH (2.5% positive nuclei/high power field vs. 0.4% in normal liver), and reached a maximum of 24 h after PH (54% positive nuclei/high power field). A second peak was observed 72 h after PH (32% positive nuclei/high power field) (Fig. 1a–d).
Normal liver regeneration (a–d) and regeneration via oval cells (e–f). (a–d) PCNA staining of liver sections from normal rats at (a), 8 h (b), 24 h (c) and 72 h (d) after standard PH in rats. (e–f) Serial sections (5 μm) of AAF-treated rats at 7 days after PH demonstrating co-localization of PCNA (e) and AFP (f)-positive cells. Scale bar = 100 μm (applies to a–f)
In AAF-treated rats, however, PCNA-positive nuclei were not seen until day 3 after PH and were found exclusively around portal tracts. A maximum of PCNA-positive cells was observed at day 7 after PH (24 ± 7% positive nuclei/HPF), all of which were identical or adjacent to AFP-immunopositive cells (Fig. 1e–f).
Protein levels of IFN-α/-γ in hepatocyte-driven liver regeneration
IFN-α and IFN-γ levels were measured in liver tissue samples and in serum samples by ELISA. We found 81 ± 22 pg/g IFN-α in homogenates of normal liver tissue. The IFN-α tissue level did not differ significantly from the control value after PH or SO (Fig. 2a). Serum levels of IFN-α were measured at the lowest detectable range of 5–10 pg/mL and did not change significantly above the lower detection range after standard PH or SO (figure not shown).
Quantification of IFN-γ in normal liver samples demonstrated a concentration that is more than 100-fold above the IFN-α concentration (10,143 pg/g liver). Immediately after PH or SO, the IFN-γ tissue level decreased to a minimum at 4 h (6,836 pg/g after PH, 6,993 pg/g after SO), followed by a constant increase back to control level (Fig. 2b). In serum samples, however, mean IFN-γ concentration was up-regulated from a normal level of 65.1 ± 15 pg/mL maximally to 343 ± 33 pg/mL in PH rats or to 226 ± 23 pg/mL in SO rats at 8 h (*p = 0.036 between PH and SO rats) (Fig. 2c).
Expression of IFNs and IFN- inducible genes in hepatocyte-driven liver regeneration
The expression IFN-inducible genes Mx-2, PKR, 2–5 OA was detectable by both real-time PCR and Northern blotting, but did not change significantly after standard PH or SO (<2-fold variations; data not shown).
Protein levels of IFN-α/-γ in oval cell-driven liver regeneration
In AAF-treated rats, hepatic IFN-α concentration increased significantly at day 7 after PH compared to sham-operated rats (187.6 ± 16.5 pg/g in PH-group vs. 55.7 ± 12.2 pg/g in SO-group; p = 0.0152). Maximal IFN-α concentration was measured at day 9 after PH (213.2 ± 32 pg/g vs. 50 ± 7 pg/g in SO-rats; p = 0.002). At days 11 and 13, the IFN-α concentration declined to levels slightly above the control level (Fig. 3a). There was no difference in the IFN-α level between normal control rats and AAF-fed controls (81.6 and 83 pg/g). Serum levels of IFN-α were below the minimum threshold as previously observed in non-AAF-treated rats.
Tissue and serum concentration of IFN-α and IFN-γ in AAF-treated rats after PH or SO (sham operation): ELISAs of liver tissue homogenates (a, b) and serum samples (c). Average protein levels were determined from three rats for each time point (±SEM). IFN-α concentration was not measurable in serum samples
IFN-γ tissue concentration increased from a normal level of 10,185 ± 155 pg/g to a maximum of 16,019 ± 1,102 pg/g at day 3 in AAF/PH-treated rats (vs. 14,670 ± 880 pg/g in AAF/SO-treated rats; p = 0.66). At day 7, the IFN-γ level fell to control level in PH- and SO-treated rats and remained at control level until day 13 (10,398 ± 194 pg/g liver tissue in PH rats) (Fig. 3b). Notably, the mean IFN-γ tissue level was higher in AAF-fed control rats than in normal controls (12,891 ± 1,565 vs. 10,185 ± 155 pg/g; p = 0.22). In serum samples, IFN-γ concentration gradually increased after PH from normal level (67 ± 9 pg/mL) to a maximum at day 7 (310 ± 49 pg/ml AAF/PH vs. 189 ± 27 pg/mL AAF/SO, p = 0.045). At day 9, serum IFN-γ concentration fell to control level. The mean IFN-γ serum level was insignificantly higher in AAF controls than in normal controls (82.4 ± 16 vs. 64 ± 9 pg/mL; p = 0.38) (Fig. 3c).
Expression of IFNs and IFN-inducible genes in oval cell-driven liver regeneration
IFN-α mRNA levels were similar in normal livers and AAF control-fed livers (± 0.04-fold). At day 3 after PH, IFN-α gene expression was significantly up-regulated (1.86-fold vs. 0.72-fold in SO-rats; p = 0.02). A maximal increase of IFN-α transcripts was found at day 7 (2.52-fold in PH rats vs. 0.75-fold in SO-rats; p = 0.037). At day 9 after PH, IFN-α gene expression was still significantly increased above control or SO level (2.28-fold vs. 1.21-fold; p = 0.02). At day 13 after PH, IFN-α mRNA fell to control or sham level, again. (Fig. 4a).
Gene expression of IFN-α/-γ, alpha-fetoprotein (AFP), Cyclin D1, and IFN-α-inducible Mx-2 gene in the AAF/PH model of liver regeneration. Relative gene expression measured by real-time PCR of IFN-α mRNA (a), IFN-γ mRNA (b), AFP (c), Cyclin D1 (d), and Mx-2 (e). Mean ± SEM; normal control livers were set as 1. (f) Representative Northern blot (NB) and Western blot (WB) hybridization for Mx-2 in normal rats (N), AAF-fed control rats (N#), and AAF-treated rats up to 13 days after 2/3 PH. Total RNA was hybridized with a 32P-labeled probe for Mx-2. Loading controls were performed with an oligonucleotide probe for ribosomal 28S or anti-β-actin antibody, respectively
Hepatic IFN-γ gene expression was up-regulated by 1.5-fold with AAF treatment alone (AAF controls compared to normal controls). At days 1 and 3 after PH, we found more IFN-γ transcripts (2.42-fold and 2.83-fold vs. normal liver). However, up-regulation of IFN-γ mRNAs was also found after SO (1.9-fold and 2.2-fold, respectively). Hence, the difference was not significant between PH- and SO-treated rats (p = 0.23 at day 1 and p = 0.25 at day 3). At day 5 after PH or SO, the level of IFN-γ mRNA returned to normal and remained at control level (±0.5-fold) until day 13 (Fig. 4b).
We used AFP mRNA as a marker for the presence of oval cells. Cyclin D1 was used as a marker of proliferation of liver cells including oval cells. Both AFP and cyclin-D1 gene expression were up-regulated starting at day 1 and reached a 210-fold (AFP) or 52-fold (Cyclin-D1) maximum at day 7 after PH. While AFP gene expression was quickly down-regulated, the expression of cyclin-D1 was still significantly above control level until day 11 (24-fold compared to normal liver) (Fig. 4c, d).
Biological activity of IFN-α was studied by expression of the Mx-2 gene, which is exclusively inducible by IFN-α but not IFN-γ. With real-time PCR, Mx-2 mRNA was up-regulated starting at day 3 after PH, and reached a transient 12-fold increase at day 7 after PH (vs. 1.04-fold after SO). The Mx-2 mRNA level fell rapidly at days 9 and 11 (3.6- and 2.2-fold after PH vs. 1.4- and 1.54-fold after SO) (Fig. 4e). Using Northern blot hybridization, Mx-2 transcripts were hardly detectable in normal rats. The Mx-2 hybridization signal increased at day 3 after PH and showed a marked transient overexpression at day 7 after PH before declining rapidly to control levels (Fig. 4f). Next, we examined Mx-2 protein by Western blotting which showed an increase starting at day 3 and culminated with a strong peak at day 9 after PH (Fig. 4f).
Activation of the IFN-α signal transduction pathway in AAF/PH-treated rat livers
We evaluated the activation of the downstream effectors in the Jak/Stat transduction pathway. Western blot analysis of liver tissue extracts showed an up-regulation of the IFN-α receptor-associated kinases Jak-1 and Tyk-2 in AAF/PH-treated rats (Fig. 5a). Day 3 was the first time point after PH that both kinases were up-regulated altogether.
Activation of the Jak/Stat-signal transduction pathway in the AAF/PH-rat model. a Protein expression of Jak-1, Tyk-2 and β-actin (control) were examined by Western blot analysis using rabbit anti-Jak-1 polyclonal Ab, mouse anti-Tyk-2 mAb and mouse anti-β-actin mAb. b Immunoprecipitations of whole liver tissue homogenates were performed with antibodies to Stat-1, Stat-2 and Stat-3. Tyrosine phosphorylation status of these proteins was then evaluated by immunoblotting using phospho-specific antibodies (upper band). Blots were subsequently stripped and reprobed with the primary anti-Stat antibody (lower band) to control the amount of immunoprecipitated protein. c Co-immunoprecipitation of Stat-3 with Stat-1 and Stat-2 proteins in the AAF/PH-rat model. N, normal controls; N#, AAF-fed control animals
In response to ligand-dependent IFN-α receptor activation, latent cytoplasmic Stat-1, Stat-2 and Stat-3 transcription factors are activated via phosphorylation on tyrosine residues. We immunoprecipitated Stat-1, Stat-2 and Stat-3 from tissue homogenates and subsequently immunoblotted the immunoprecipitates with phospho-specific Stat antibodies. In normal resting livers, Stat-1 activation was not detected but Stat-2 and Stat- 3 activation was found at low levels (Fig. 5b). In AAF-fed control livers, Stat-1 was activated at low levels similar to the ones found for phosphorylated Stat-2 and Stat-3. In AAF/PH-treated animals, we found an enhanced activation of Stat-1, Stat-2 and Stat-3 starting at day 1 after PH. Stat-1 was maximally activated at days 3–7, Stat-2 similarly between days 1 and 11, whereas phosphorylated Stat-3 was maximally detected at day 7 (Fig. 5b).
To detect whether Stat-3 interacts with corresponding Stat-proteins, we immunoprecipitated proteins with an antibody directed to Stat-1, and detected Stat-3 at days of Stat-1 activation (days 3, 7 and 11; Fig. 5c). As shown in Fig. 5b, Stat-2 was activated from day 1 up to day 11 after PH. Interestingly, Stat-3 co-immunoprecipitated with Stat-2 at days 1 and 11 only (Fig. 5c).
IFN-α expression in the periportal zone of AAF/PH-treated rat livers
To localize IFN-α synthesis, we performed in situ hybridization and LCM of AAF-treated rat livers. In normal untreated livers and AAF-fed control livers, hybridization signals for IFN-α were not detectable. Hybridization signals for IFN-α were increasingly detectable from day 3 up to day 7 after PH, and decreased thereafter. Hybridization signals for IFN-α were almost exclusively found in the periportal zone, thus labeling a large proportion of the periportal cellular reaction at day 7 (Fig. 6a, b).
Localization of IFN-α synthesis: (a–b) In situ hybridization with an IFN-α-specific probe on sections of AAF-treated livers. IFN-α-specific cDNA was transcribed into DIG-labeled antisense (a) or sense (b) RNA probes and hybridized with sections of livers at day 7 after PH; magnification ×100. (c–e) Laser capture microdissection (LCM) and subsequent RT-PCR analysis of IFN-α mRNA. Representative portal field of a regenerating liver at day 7 after PH/AAF treatment before (c) and after LCM (d). Total RNA was extracted from the LCM sample and subsequently analyzed by RT-PCR for IFN-α transcripts (e). PF, periportal area (zone I); CV, pericentral area (zone III); H2O negative control
Next, we microdissected liver cells by laser capture around portal tracts (zone 1) and central veins (zone 3) at day 7 after PH/AAF treatment and looked for IFN-α transcripts by RT-PCR analysis (Fig. 6c, d). According to the results above, we found IFN-α transcripts in the portal area, but not centrally within the liver lobule (zone 3) (Fig. 6e).
Detection of mononuclear phagocytes (MNP) and natural killer (NK) cells
MNP are known to be a principal source of IFN-α. To check for MNP in AAF/PH-treated livers, we performed immunohistochemical staining with the monoclonal antibodies ED1 and ED2 in serial sections already used for in situ hybridization or LCM. The antigen recognized by ED1 is the rat homolog of human CD68, and the antigen recognized by ED2 is the rat CD163 cell surface glycoprotein. Cells positive for ED1 and ED2 were found along the sinusoids, in accordance with the assumption that mononuclear phagocytes (ED1 positivity) resident in the normal liver are mainly mature tissue macrophages (ED2 positivity) (Fig. 7a, b). In rats treated by PH/AAF administration, ED1+ cells were increasingly found within the periportal cellular reaction containing AFP+ cells. ED1+ cells were found adjacent to AFP+ cells and their numbers increased in parallel. The periportal ED1+ cells were mainly ED2−, hence representing inflammatory cells (Fig. 7c–f).
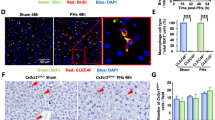
Detection of mononuclear phagocytes (MNP) by POD in AAF/PH-treated rats. Normal livers (a–b): ED1-positive cells (arrowheads) are dispersed throughout the entire parenchyma and the portal tracts of normal livers (a), whereas ED-2-positive cells (short arrows) were mainly found within the parenchyma but not in the portal tracts (b). AAF/PH-treated rats (c–f): ED1-positive cells within periportal clusters of day 3 (c), day 7 (d, e) and day 13 after PH (f). NK cells were detected in serial sections by APAAP staining for the NKp30 cell receptor. g No NKp30-positive cells were detectable in AAF-fed control rats. h Maximum numbers at day 11 after PH (long arrows). Magnification ×100 (a–d), ×200 (e–g), ×400 (h). i Representative Western blot of ED1 protein expression in liver homogenates up to 13 days after AAF/PH treatment (N, normal rat liver; N#, AAF-fed control rat liver). β-actin expression was used as a loading control
NK cells are also known to reside in the liver and are capable of producing significant amounts of IFN-γ and minor amounts of IFN-α (Ahmad and Alvarez 2001). To determine whether NK cells may also be up-regulated in the setting of AAF/PH treatment, we performed APAAP immunostaining for the ubiquitous NK cell receptor NKp30, which plays a prominent role in NK cell activation. In contrast to ED1 immunostaining, we could not detect a significant influx or up-regulation of NKp30-positive cells. NKp30-immunopositive cells were not detectable in normal or AAF-fed control rats (Fig. 7g). We found single NKp30-positive cells at days 7 and 11. Within one enlarged portal field only, a maximum of three immunopositive (red) cells were found at day 11 after PH (Fig. 7h). Real-time PCR of the mRNA encoding the NKp30 cell receptor did not show any significant variation in AAF/PH-treated rats (data not shown). The amount of NKp30 mRNA at day 11 after PH was 1.4-fold compared to normal liver.
We studied total protein content by Western blotting and found increased amounts of ED1 antigens starting at day 1 after PH. Maximal levels were found at day 9 after PH (Fig. 7i). Western blots for Nkp30 showed constant levels without significant changes, thus confirming our previous immunohistochemical and PCR results (not shown).
Discussion
IFNs are a family of related cytokines that mediate a range of diverse functions including immunomodulatory and antiproliferative activities. All subtypes of IFNs elicit their cellular effects through the transcriptional activation of target genes by activated Stat (signal transducer and activator of transcription) proteins. Type I (α and β) and type II (γ) IFN signal through distinct receptors but related Jak-Stat pathways (Heim 2000). Different combinations of these Stats result in different transcription factor complexes and binding to two different classes of response elements (Kalvakolanu 2003). In most cell lines, activation of the IFN-γ receptor leads to phosphorylation of Stat-1 only, followed by nuclear translocation of Stat-1 homodimers and binding to gamma-activated sequence (GAS) response elements of IFN-inducible genes.
On the other hand, the activation of the IFN-α-specific receptor induces the activation of Stat-1, Stat-2 and Stat-3, all of which may activate GAS response elements once they have combined with different combinations of dimers. IFN-α-specific gene expression may ensue when Stat-1 and Stat-2 form heterodimers, which in combination with interferon-regulatory factor 9 (IRF-9) can bind to interferon-stimulated response elements (ISRE) in the promoters of IFN-α target genes (Ozato et al. 2007).
Compared to IFN-γ, Type I IFNs have more potent antiviral, antiproliferative and antitumoral properties. Hence, IFN-α is currently the most frequently used cytokine in the treatment neoplastic diseases or viral hepatitis (Brassard et al. 2002). It is now well understood that its additional capacity to activate ISRE harboring genes accounts best for its unique actions (Heim 2000). Among these ISRE-harboring genes, the Mx gene family is exceptional because it is activated in vivo almost exclusively by IFN-α, but not IFN-γ. Thus, Mx gene products were used as surrogate markers for the biological activity of IFN-α in humans and rodents (Staeheli et al. 1986; von Wussow et al. 1990; Meier et al. 1990).
We found increased IFN-γ serum levels after PH in both models of regeneration. Serum concentration of IFN-α remained so low that it was not measurable. Comparing liver tissue levels of IFN-α/-γ after standard PH or SO, we did not find any significant changes or any activation of corresponding signaling pathways. However, hepatic levels of IFN-α and IFN-γ were up-regulated in AAF/PH-treated livers. Whereas IFN-γ was up-regulated at early time points, up-regulation of IFN-α and activation of IFN-α signaling occurred at later time points simultaneously with the peak of oval cell proliferation. In parallel, we found increasing numbers of IFN-expressing, monocyte-like mononuclear phagocytes (MNP) in regenerating livers of AAF/PH-treated rats.
The tyrosine kinase 2 (Tyk-2) is associated with the IFN-α receptor, but not with the IFN-γ receptor. On protein level, we found a constitutive up-regulation of Tyk-2 starting already at day 1 after PH. Stat-1, Stat-2 and Stat-3 were activated starting at day 1 in AAF/PH-treated rats. Stat-1 and Stat-2 phosphorylation was similarly up-regulated at days 1 and 3 after PH, whereas Stat-3 activation occurred later at day 7 after PH.
In the liver, Stat-1 is mainly activated by IFN-α and IFN-γ and its activation is correlated positively with liver injury, but negatively with hepatocyte proliferation or liver regeneration (Gao 2005; Sun et al. 2006). Activation of Stat-1 or Stat-3 plays an essential role in the antiviral and antitumor activity of IFN-α (Meraz et al. 1996; Zhu et al. 2004).
Stat-2 is known to be activated only by Type I IFN (IFN-α/IFN-β), but not by IFN-γ or any other cytokines (Heim 2000; Gao 2005). STAT2-deficient mice have an increased susceptibility to viral infection and the loss of a Type I IFN autocrine/paracrine loop (Park et al. 2000). Its activation is a key signal involved in the antiviral therapy of IFN-α in viral hepatitis patients (Radaeva et al. 2002; Nguyen and Gao 2002).
Stat-3 was shown to be activated in hepatocytes after standard PH (Cressman et al. 1995) and in mature oval cells differentiating along the hepatocyte lineage (Sanchez et al. 2004). Stat-3 can be activated by a variety of cytokines, including interleukin-6, OncostatinM, leukemia inhibiting factor, epidermal growth factor, platelet-derived growth factor, G-CSF, leptin and IFN-α, but not by IFN-γ (Akira 2000; Waelput et al. 2000; Heim 2000; Yang et al. 1998). In models of liver regeneration, IL-6 seems to be by far the most relevant pathway for Stat-3 activation (Cressman et al. 1996). Interaction of IL-6 with its receptor induces the activation of cytoplasmic Stat-3, but not Stat-1 or Stat-2. Activated Stat-3 then translocates to the nucleus to induce the transcription of many target genes, including acute-phase genes, but not antiviral genes such as Mx (Akira 1997). Several studies have demonstrated an important role of the IL-6/Stat-3 pathway in hepatoprotection, liver development and normal liver regeneration after standard PH (Wuestefeld et al. 2003; Streetz et al. 2003). Stat-3 signaling may be also necessary for oval cell-mediated liver regeneration since levels of IL-6 are elevated in corresponding mice models, and IL-6-mediated hyperactive Stat-3 signaling results in enhanced oval cell numbers (Yeoh et al. 2007; Matthews et al. 2004).
We found a constitutive activation of Stat-3 in normal rats and maximal Stat-3 phosphorylation at day 7 after PH when maximal AFP/PCNA immunopositivity occurred. Co-immunoprecipitation studies showed that maximal Stat-3 activation is not associated with Stat-2 activation, indicating that other cytokines besides IFN-α may cause Stat-3 activation during oval cell-driven liver regeneration.
Notably, Stat-1 and Stat-2 were activated earlier than Stat-3. Similar to the increase of the IFN-γ level in AAF/PH-treated rats, Stat-1 was already activated at days 1 and 3 after PH, before maximal oval cell numbers occurred. Tyk-2 and Stat-2 are two members of the IFN-α signaling pathway: the expression and activation of both were simultaneously up-regulated from day 1 up to 11 of the AAF/PH-protocol.
NK cells are important effector cells of innate immunity known to produce large amounts of IFN-γ and eventually IFN-α, once activated (Ahmad and Alvarez 2001). Sun and Gao have shown that hepatic NK cells are increased in numbers after PH and highly activated by demonstrating IFN-γ production in hepatic NK cells (Sun and Gao 2004). We studied the presence of activated NK cells in oval cell-mediated liver regeneration by identification of the ubiquitous NKp30 receptor (Pende et al. 1999). We could not detect NKp30-positive cells in normal rats or before 7 days after PH. Thereafter, single NKp30-positive cells were detectable interspersed in the periportal cellular reaction. Since we could not measure an increased IFN-γ or NKp30 level in liver tissue at days 7 or later, we suggest that the low numbers of NKp30-positive NK cells do not significantly affect IFN-γ or IFN-α tissue levels.
MNP are known to be recruited and activated by IFN-γ in models of chemical-induced hepatocarcinogenesis (Matsuda et al. 2005). The MNP system is characterized by the expression of ED1 or ED2 antigens (Dijkstra et al. 1985). No other cell types except MNP are positive for any ED-monoclonal antibody. ED1 recognizes a cytoplasmic antigen (CD68) present in monocytes and in most macrophages, whereas ED-2 recognizes a membrane antigen (CD163) of tissue macrophages only (e.g., Kupffer cells of the liver) (Armbrust and Ramadori 1995). Several studies have revealed not only morphological but also functional differences between monocytes and resident tissue macrophages (Armbrust et al. 1997, Armbrust et al. 1993). We could show in a previous work that small ED1+/ED2− monocyte-like MNPs constitutively express IFN-α and that Mx-2 is constitutively expressed in those monocyte-like MNPs, but not in resident ED1+/ED2+ macrophages (Kupffer cells) (Batusic et al. 2004). In this paper, we demonstrate an up-regulation of Mx-2 at RNA and protein levels in the AAF/PH model. In parallel, the number of monocyte-like ED1+/ED2− MNPs was increased. In particular, ED1+ MNPs were found adjacent to AFP+/PCNA+ cells of the periportal zone. Using in situ hybridization and LCM, we found increased IFN-α expression in periportal zones of AAF/PH-treated rats, but not in the pericentral zone consisting mainly of resting hepatocytes. We postulate that the up-regulation of IFN-α and Mx expression might be a direct effect of the influx of monocyte-like MNP during the proliferation of the oval cells.
IFN-γ levels were not significantly different between PH and SO rats in both regeneration models except for a transient increase in serum. Treatment with AAF alone tended to increase IFN-γ in serum and tissues, without any operation. Therefore, we conclude that IFN-γ behaves as an acute-phase cytokine in both models of liver regeneration, probably promoting the recruitment/activation of MNP into the liver. The IFN-γ level of PH- or sham-operated rats differed significantly only in serum, indicating additional extrahepatic IFN-γ production. The increased activity of IFN-α in oval cell-driven liver regeneration might be an effect of an influx of monocyte-like MNP, but not NK cells. Those inflammatory MNPs may regulate the proliferation of oval cells through local expression of cytokines including IFN-α.
References
Ahmad A, Alvarez F (2001) Role of NK and NKT cells in the immunopathogenesis of HCV-induced hepatitis. J Leukoc Biol 76:743–759
Akhurst B, Matthews V, Husk K, Smyth MJ, Abraham LJ, Yeoh GC (2005) Differential lymphotoxin-beta and interferon gamma signaling during mouse liver regeneration induced by chronic and acute injury. Hepatology 41(2):327–335
Akira S (1997) IL-6-regulated transcription factors. Int J Biochem Cell Biol 29(12):1401–1418
Akira S (2000) Roles of STAT3 defined by tissue-specific gene targeting. Oncogene 19:2607–2611
Alvarez ML, Cerliani JP, Monti JA, Carnovale CE, Ronco T, Pellegrino JM, Pisani G, Lugano C, Carrillo MC (2001) In vivo apoptotic effect of alpha-2b interferon (IFN) on rat preneoplastic liver. Medicina (B Aires) 61(5 Pt 2):666–669
Armbrust T, Ramadori G (1995) Mononuclear phagocytes of acutely injured rat liver abundantly synthesize and secrete fibronectin in contrast to Kupffer cells of normal liver. Biochem Biophys Res Commun 213(3):752–758
Armbrust T, Schwogler S, Zohrens G, Ramadori G (1993) C1 esterase inhibitor gene expression in rat Kupffer cells, peritoneal macrophages and blood monocytes: modulation by interferon gamma. J Exp Med 178(2):373–380
Armbrust T, Nordmann B, Kreissig M, Ramadori G (1997) C1Q synthesis by tissue mononuclear phagocytes from normal and from damaged rat liver: up-regulation by dexamethasone, down-regulation by interferon gamma, and lipopolysaccharide. Hepatology 26(1):98–106
Batusic DS, Armbrust T, Saile B, Ramadori G (2004) Induction of Mx-2 in rat liver by toxic injury. J Hepatol 40(3):446–453
Batusic DS, Cimica V, Chen Y, Tron K, Hollemann T, Pieler T, Ramadori G (2005) Identification of genes specific to “oval cells” in the rat 2-acetylaminofluorene/partial hepatectomy model. Histochem Cell Biol 124(3–4):245–260
Bisgaard HC, Muller S, Nagy P, Rasmussen LJ, Thorgeirsson SS (1999) Modulation of the gene network connected to interferon-gamma in liver regeneration from oval cells. Am J Pathol 155(4):1075–1085
Blaschke S, Koziolek M, Schwarz A, Benohr P, Middel P, Schwarz G, Hummel KM, Muller GA (2003) Proinflammatory role of fractalkine (CX3CL1) in rheumatoid arthritis. J Rheumatol 30(9):1918–1927
Braissant O, Wahli W (1998) Simplified in situ hybridization protocol using non-radioactively labelled probes to detect abundant and rare mRNAs on tissue section. Biochemica 1:10–16
Brassard DL, Grace MJ, Bordens RW (2002) Interferon-alpha as an immunotherapeutic protein. J Leukoc Biol 71(4):565–581
Braun L, Mikumo R, Fausto N (1989) Production of hepatocellular carcinoma by oval cells: cell cycle expression of c-myc and p53 at different stages of oval cell transformation. Cancer Res 49(6):1554–1561
Brooling JT, Campbell JS, Mitchell C, Yeoh GC, Fausto N (2005) Differential regulation of rodent hepatocyte and oval cell proliferation by interferon gamma. Hepatology 41(4):906–915
Chirgwin JM, Przybyla AE, MacDonald RJ, Rutter WJ (1979) Isolation of biologically active ribonucleic acid from sources enriched in ribonuclease. Biochemistry 18(24):5294–5299
Cimica V, Batusic D, Chen Y, Hollemann T, Pieler T, Ramadori G (2005) Transcriptome analysis of rat liver regeneration in a model of oval hepatic stem cells. Genomics 86(3):352–364
Cressman DE, Diamond RH, Taub R (1995) Rapid activation of the Stat3 transcription complex in liver regeneration. Hepatology 21:1443–1449
Cressman DE, Greenbaum LE, DeAngelis RA, Ciliberto G, Furth EE, Poli V, Taub R (1996) Liver failure and defective hepatocyte regeneration in interleukin-6-deficient mice. Science 274:1379–1383
Dijkstra CD, Dopp EA, Joling P, Kraal G (1985) The heterogeneity of mononuclear phagocytes in lymphoid organs: distinct macrophage subpopulations in the rat recognized by monoclonal antibodies ED1, ED2 and ED3. Immunology 54(3):589–599
Dumble ML, Croager EJ, Yeoh GC, Quail EA (2002) Generation and characterization of p53 null transformed hepatic progenitor cells: oval cells give rise to hepatocellular carcinoma. Carcinogenesis 23(3):435–445
Favre C, Carnovale CE, Monti JA, Carrillo MC (2001) Inhibition by interferon alpha-2b of rat liver regeneration: effect on ornithine decarboxylase and total protein synthesis. Biochem Pharmacol 61(12):1587–1593
Gao B (2005) Cytokines, STATs and liver disease. Cell Mol Immunol 2(2):92–100
Heim MH (2000) Intracellular signalling and antiviral effects of interferons. Dig Liver Dis 32(3):257–263
Kalvakolanu DV (2003) Alternate interferon signaling pathways. Pharmacol Ther 100(1):1–29
Knight B, Lim R, Yeoh GC, Olynyk JK (2007) Interferon-gamma exacerbates liver damage, the hepatic progenitor cell response and fibrosis in a mouse model of chronic liver injury. J Hepatol 47(6):826–833
Matsuda M, Nakamoto Y, Suzuki S, Kurata T, Kaneko S (2005) Interferon-gamma-mediated hepatocarcinogenesis in mice treated with diethylnitrosamine. Lab Invest 85:655–663
Matthews VB, Klinken E, Yeoh GC (2004) Direct effects of interleukin-6 on liver progenitor oval cells in culture. Wound Repair Regen 12(6):650–656
McCormick D, Yu C, Hobbs C, Hall PA (1993) The relevance of antibody concentration to the immunohistological quantification of cell proliferation-associated antigens. Histopathology 22(6):543–547
Meier E, Kunz G, Haller O, Arnheiter H (1990) Activity of rat Mx proteins against a rhabdovirus. J Virol 64(12):6263–6269
Meraz MA, White JM, Sheehan KC, Bach EA, Rodig SJ, Dighe AS, Kaplan DH, Riley JK, Greenlund AC, Campbell D, Carver-Moore K, DuBois RN, Clark R, Aguet M, Schreiber RD (1996) Targeted disruption of the Stat1 gene in mice reveals unexpected physiologic specificity in the JAK-STAT signaling pathway. Cell 84(3):431–442
Nakaji M, Yano Y, Ninomiya T, Seo Y, Hamano K, Yoon S, Kasuga M, Teramoto T, Hayashi Y, Yokozaki H (2004) IFN-alpha prevents the growth of pre-neoplastic lesions and inhibits the development of hepatocellular carcinoma in the rat. Carcinogenesis 25(3):389–397
Nguyen VA, Gao B (2002) Expression of interferon alfa signaling components in human alcoholic liver disease. Hepatology 35(2):425–432
Nishiguchi S, Otani S, Matsui-Yuasa I, Morisawa S, Monna T, Kuroki T, Kobayashi K, Yamamoto S (1986) Inhibition by interferon (alpha + beta) of mouse liver regeneration and its reversal by putrescine. FEBS Lett 205(1):61–65
Ozato K, Tailor P, Kubota T (2007) The interferon regulatory family in host defense: mechanism of action. J Biol Chem 282(28):20065–20069
Park C, Li S, Cha E, Schindler C (2000) Immune response in Stat2 knockout mice. Immunity 13(6):795–804
Pende D, Parolini S, Pessino A, Sivori S, Augugliaro R, Morelli L, Marcenaro E, Accame L, Malaspina A, Biassoni R, Bottino C, Moretta L, Moretta A (1999) Identification and molecular characterization of NKp30, a novel triggering receptor involved in natural cytotoxicity mediated by human natural killer cells. J Exp Med 190(10):1505–1516
Radaeva S, Jaruga B, Hong F, Kim WH, Fan S, Cai H, Strom S, Liu Y, El-Assal O, Gao B (2002) Interferon-alpha activates multiple STAT signals and down-regulates c-Met in primary human hepatocytes. Gastroenterology 122(4):1020–1034
Sanchez A, Factor VM, Schroeder IS, Nagy P, Thorgeirsson SS (2004) Activation of NF-kappaB and STAT3 in rat oval cells during 2-acetylaminofluorene/partial hepatectomy-induced liver regeneration. Hepatology 39(2):376–385
Staeheli P, Danielson P, Haller O, Sutcliffe JG (1986) Transcriptional activation of the mouse Mx gene by Type I interferon. Mol Cell Biol 6(12):4770–4774
Streetz KL, Tacke F, Leifeld L, Wustefeld T, Graw A, Klein C, Kamino K, Spengler U, Kreipe H, Kubicka S, Muller W, Manns MP, Trautwein C (2003) Interleukin 6/gp130-dependent pathways are protective during chronic liver diseases. Hepatology 38(1):218–229
Sun R, Gao B (2004) Negative regulation of liver regeneration by innate immunity (natural killer cells/interferon-gamma). Gastroenterology 127(5):1525–1539
Sun R, Park O, Horiguchi N, Kulkarni S, Jeong WI, Sun HY, Radaeva S, Gao B (2006) STAT1 contributes to dsRNA inhibition of liver regeneration after partial hepatectomy in mice. Hepatology 44(4):955–966
Theocharis SE, Margeli AP, Tsokos MG (1995) Alpha 2b-interferon inhibits rat liver regeneration after partial hepatectomy without affecting thymidine kinase activity. J Lab Clin Med 125(5):588–596
Theocharis SE, Margeli AP, Skaltsas SD, Skopelitou AS, Mykoniatis MG, Kittas CN (1997) Effect of interferon-alpha2b administration on rat liver regeneration after partial hepatectomy. Dig Dis Sci 42(9):1981–1986
von Wussow P, Jakschies D, Hochkeppel HK, Fibich C, Penner L, Deicher H (1990) The human intracellular Mx-homologous protein is specifically induced by Type I interferons. Eur J Immunol 20(9):2015–2019
Waelput W, Verhee A, Broekaert D, Eyckerman S, Vanderkerckhove J, Beattie JH, Tavernier J (2000) Identification and expression analysis of leptin-regulated immediate early response and late target genes. Biochem J 48:55–61
Wuestefeld T, Klein C, Streetz KL, Betz U, Lauber J, Buer J, Manns MP, Muller W, Trautwein C (2003) Interleukin-6/glycoprotein 130-dependent pathways are protective during liver regeneration. J Biol Chem 278(13):11281–11288
Yang CH, Murti A, Pfeffer LM (1998) STAT3 complements defects in an interferon-resistant cell line: evidence for an essential role for STAT3 in interferon signaling and biological activities. Proc Natl Acad Sci USA 95(10):5568–5572
Yeoh GC, Ernst M, Rose-John S, Akhurst B, Payne C, Long S, Alexander W, Croker B, Grail D, Matthews VB (2007) Opposing roles of gp130-mediated STAT-3 and ERK-1/2 signaling in liver progenitor cell migration and proliferation. Hepatology 45(2):486–494
Zhu H, Shang X, Terada N, Liu C (2004) STAT3 induces anti-hepatitis C viral activity in liver cells. Biochem Biophys Res Commun 324(2):518–528
Acknowledgments
This work was supported by the Deutsche Forschungsgemeinschaft SFB 402, project D3. The authors wish to thank Prof. Dr. O. Haller for his kind provision of the monoclonal Mx antibody (mAB M143) and Mrs. C. Hoffmann for her excellent technical assistance.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Batusic, D.S., von Bargen, A., Blaschke, S. et al. Different physiology of interferon-α/-γ in models of liver regeneration in the rat. Histochem Cell Biol 136, 131–144 (2011). https://doi.org/10.1007/s00418-011-0838-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00418-011-0838-7