Abstract
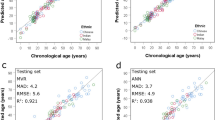
Age-related CpG sites (AR-CpGs) are currently the most promising biomarkers for forensic age estimation. In our previous studies, we first validated the age correlation of seven reported AR-CpGs in blood samples of Chinese Han population. Subsequently, we screened some good age predictors from blood samples of Chinese Han population, and built pyrosequencing-based age prediction models. However, it is still important to select a set of high-performance AR-CpGs in a specific racial group and establish a simple and efficient method for accurate age estimation for forensic purpose. In this study, eight AR-CpGs, namely chr6: 11,044,628 (ELOVL2), cg06639320 (FHL2), chr1: 207,823,723 (C1orf132), cg19283806 (CCDC102B), cg14361627 (KLF14), cg17740900 (SYNE2), cg07553761 (TRIM59), and cg26947034, were selected based on our previous studies, and a multiplex methylation SNaPshot assay was developed to investigate DNA methylation levels at these AR-CpGs in 529 blood samples (aged 2–82 years) from Han Chinese population. All selected CpG sites showed strong age correlation with the correlation coefficient (r) from 0.8363 to 0.9251. Multiple linear regression (MLR) and support vector regression (SVR) age prediction models were simultaneously established to fit change characteristics of DNA methylation levels of eight AR-CpGs with the age in 374 donors’ blood samples. The MLR model enabled age prediction with R2 = 0.923, mean absolute error (MAE) = 3.52, while the SVR model enabled age prediction with R2 = 0.935, MAE = 2.88. One hundred fifty-five independent samples were used as a validation set to test the two models’ performance, and the prediction MAE for the validation set was 3.71 and 3.34 for the MLR and SVR models, respectively. For the MLR and SVR models, the correct prediction rate at ± 5 years reached a high level of 79.35% and 83.23%, respectively. In general, these statistical parameters indicated that the SVR model outperformed the MLR model in age prediction of the Han Chinese population. In addition, our method provides sufficient sensitivity in forensic applications and allows for 100% efficiency when examining bloodstains kept in room conditions for up to 43 days. These results indicate that our multiplex methylation SNaPshot assay is a reliable, effective, and accurate method for age prediction in blood samples from the Chinese Han population.




Similar content being viewed by others
References
Lynnerup N, Kjeldsen H, Zweihoff R, Heegaard S, Jacobsen C, Heinemeier J (2010) Ascertaining year of birth/age at death in forensic cases: a review of conventional methods and methods allowing for absolute chronology. Forensic Sci Int 201(1–3):74–78. https://doi.org/10.1016/j.forsciint.2010.03.026
Hayakawa M, Hattori K, Sugiyama S, Ozawa T (1992) Age-associated oxygen damage and mutations in mitochondrial DNA in human hearts. Biochem Biophys Res Commun 189(2):979–985. https://doi.org/10.1016/0006-291X(92)92300-M
Cortopassi GA, Shibata D, Soong NW, Arnheim N (1992) A pattern of accumulation of a somatic deletion of mitochondrial DNA in aging human tissues. Proc Natl Acad Sci U S A 89(16):7370–7374. https://doi.org/10.1073/pnas.89.16.7370
Baynes JW (2001) The role of AGEs in aging: causation or correlation. Exp Gerontol 36(9):1527–1537. https://doi.org/10.1016/S0531-5565(01)00138-3
Blasco MA (2005) Telomeres and human disease: ageing, cancer and beyond. Nat Rev Genet 6(8):611–622. https://doi.org/10.1038/nrg1656
Meissner C, Ritz-Timme S (2010) Molecular pathology and age estimation. Forensic Sci Int 203(1–3):34–43. https://doi.org/10.1016/j.forsciint.2010.07.010
Zubakov D, Liu F, van Zelm MC, Vermeulen J, Oostra BA, van Duijn CM, Driessen GJ, van Dongen JJ, Kayser M, Langerak AW (2010) Estimating human age from T-cell DNA rearrangements. Curr Biol 20(22):R970–R971. https://doi.org/10.1016/j.cub.2010.10.022
Ou X, Zhao H, Sun H, Yang Z, Xie B, Shi Y, Wu X (2011) Detection and quantification of the age-related sjTREC decline in human peripheral blood. Int J Legal Med 125(4):603–608. https://doi.org/10.1007/s00414-010-0528-3
de Magalhães JP, Curado J, Church GM (2009) Meta-analysis of age-related gene expression profiles identifies common signatures of aging. Bioinformatics 25(7):875–881. https://doi.org/10.1093/bioinformatics/btp073
Horvath S (2013) DNA methylation age of human tissues and cell types. Genome Biol 14(10):R115. https://doi.org/10.1186/gb-2013-14-10-r115
Zbieć-Piekarska R, Spólnicka M, Kupiec T, Parys-Proszek A, Makowska Ż, Pałeczka A, Kucharczyk K, Płoski R, Branicki W (2015) Development of a forensically useful age prediction method based on DNA methylation analysis. Forensic Sci Int Genet 17:173–179. https://doi.org/10.1016/j.fsigen.2015.05.001
Goel N, Karir P, Garg VK (2017) Role of DNA methylation in human age prediction. Mech Ageing Dev 166:33–41. https://doi.org/10.1016/j.mad.2017.08.012
Zubakov D, Liu F, Kokmeijer I, Choi Y, van Meurs JBJ, van IJcken WFJ, Uitterlinden AG, Hofman A, Broer L, van Duijn CM, Lewin J, Kayser M (2016) Human age estimation from blood using mRNA, DNA methylation, DNA rearrangement, and telomere length. Forensic Sci Int Genet 24:33–43. https://doi.org/10.1016/j.fsigen.2016.05.014
Weidner CI, Lin Q, Koch CM, Eisele L, Beier F, Ziegler P, Bauerschlag DO, Jöckel KH, Erbel R, Mühleisen TW, Zenke M, Brümmendorf TH, Wagner W (2014) Aging of blood can be tracked by DNA methylation changes at just three CpG sites. Genome Biol 15(2):R24. https://doi.org/10.1186/gb-2014-15-2-r24
Bekaert B, Kamalandua A, Zapico SC, Van de Voorde W, Decorte R (2015) A selective set of DNA-methylation markers for age determination of blood, teeth and buccal samples. Forensic Sci Int Genet Suppl Ser 5:e144–e145. https://doi.org/10.1016/j.fsigss.2015.09.058
Freire-Aradas A, Phillips C, Mosquera-Miguel A, Girón-Santamaría L, Gómez-Tato A, Casares de Cal M, Álvarez-Dios J, Ansede-Bermejo J, Torres-Español M, Schneider PM, Pośpiech E, Branicki W, Carracedo Á, Lareu MV (2016) Development of a methylation marker set for forensic age estimation using analysis of public methylation data and the Agena Bioscience EpiTYPER system. Forensic Sci Int Genet 24:65–74. https://doi.org/10.1016/j.fsigen.2016.06.005
Feng L, Peng F, Li S, Jiang L, Sun H, Ji A, Zeng C, Li C, Liu F (2018) Systematic feature selection improves accuracy of methylation-based forensic age estimation in Han Chinese males. Forensic Sci Int Genet 35:38–45. https://doi.org/10.1016/j.fsigen.2018.03.009
Aliferi A, Ballard D, Gallidabino MD, Thurtle H, Barron L, Syndercombe Court D (2018) DNA methylation-based age prediction using massively parallel sequencing data and multiple machine learning models. Forensic Sci Int Genet 37:215–226. https://doi.org/10.1016/j.fsigen.2018.09.003
Jung SE, Lim SM, Hong SR, Lee EH, Shin KJ, Lee HY (2019) DNA methylation of the ELOVL2, FHL2, KLF14, C1orf132/MIR29B2C, and TRIM59 genes for age prediction from blood, saliva, and buccal swab samples. Forensic Sci Int Genet 38:1–8. https://doi.org/10.1016/j.fsigen.2018.09.010
Dias HC, Cordeiro C, Pereira J, Pinto C, Real FC, Cunha E, Manco L (2020) DNA methylation age estimation in blood samples of living and deceased individuals using a multiplex SNaPshot assay. Forensic Sci Int 311:110267. https://doi.org/10.1016/j.forsciint.2020.110267
Pan C, Yi S, Xiao C, Huang Y, Chen X, Huang D (2020) The evaluation of seven age-related CpGs for forensic purpose in blood from Chinese Han population. Forensic Sci Int Genet 46:102251. https://doi.org/10.1016/j.fsigen.2020.102251
Xiao C, Yi S, Huang D (2021) Genome-wide identification of age-related CpG sites for age estimation from blood DNA of Han Chinese individuals. Electrophoresis 42(14–15):1488–1496. https://doi.org/10.1002/elps.202000367
Pisarek A, Pośpiech E, Heidegger A, Xavier C, Papież A, Piniewska-Róg D, Kalamara V, Potabattula R, Bochenek M, Sikora-Polaczek M, Macur A, Woźniak A, Janeczko J, Phillips C, Haaf T, Polańska J, Parson W, Kayser M, Branicki W (2021) Epigenetic age prediction in semen - marker selection and model development. Aging Albany NY 13(15):19145–19164
Fraser HB, Lam LL, Neumann SM, Kobor MS (2012) Population-specificity of human DNA methylation. Genome Biol 13(2):R8. https://doi.org/10.1186/gb-2012-13-2-r8
Day K, Waite LL, Thalacker-Mercer A, West A, Bamman MM, Brooks JD, Myers RM, Absher D (2013) Differential DNA methylation with age displays both common and dynamic features across human tissues that are influenced by CpG landscape. Genome Biol 14(9):R102. https://doi.org/10.1186/gb-2013-14-9-r102
Poulin M, Zhou JY, Yan L, Shioda T (2018) Pyrosequencing methylation analysis. Methods Mol Biol 1856:283–296. https://doi.org/10.1007/978-1-4939-8751-1_17
Kaminsky Z, Petronis A (2009) Methylation SNaPshot: a method for the quantification of site-specific DNA methylation levels. Methods Mol Biol 507:241–255. https://doi.org/10.1007/978-1-59745-522-0_18
Lin YC, Tsai LC, Lee JC, Liu KL, Tzen JT, Linacre A, Hsieh HM (2016) Novel identification of biofluids using a multiplex methylation-specific PCR combined with single-base extension system. Forensic Sci Med Pathol 12(2):128–138. https://doi.org/10.1007/s12024-016-9763-3
Li LC, Dahiya R (2002) MethPrimer: designing primers for methylation PCRs. Bioinformatics 18:1427–1431. https://doi.org/10.1093/bioinformatics/18.11.1427
Untergasser A, Cutcutache I, Koressaar T, Ye J, Faircloth BC, Remm M, Rozen SG (2012) Primer3 – new capabilities and interfaces. Nucleic Acids Res 40(15):e115. https://doi.org/10.1093/nar/gks596
Vallone PM, Butler JM (2004) AutoDimer: a screening tool for primer-dimer and hairpin structures. Biotechniques 37(2):226–231. https://doi.org/10.2144/04372ST03
Chang CC, Lin CJ (2011) LIBSVM: A library for support vector machines. Acm T Intel Syst Tec 2:27. https://doi.org/10.1145/1961189.1961199
Kaminsky ZA, Assadzadeh A, Flanagan J, Petronis A (2005) Single nucleotide extension technology for quantitative site-specific evaluation of metC/C in GC-rich regions. Nucleic Acids Res 33(10):e95. https://doi.org/10.1093/nar/gni094
Zbieć-Piekarska R, Spólnicka M, Kupiec T, Makowska Ż, Spas A, Parys-Proszek A, Kucharczyk K, Płoski R, Branicki W (2015) Examination of DNA methylation status of the ELOVL2 marker may be useful for human age prediction in forensic science. Forensic Sci Int Genet 14:161–167. https://doi.org/10.1016/j.fsigen.2014.10.002
Bekaert B, Kamalandua A, Zapico SC, Van de Voorde W, Decorte R (2015) Improved age determination of blood and teeth samples using a selected set of DNA methylation markers. Epigenetics 10(10):922–930. https://doi.org/10.1080/15592294.2015.1080413
Park JL, Kim JH, Seo E, Bae DH, Kim SY, Lee HC, Woo KM, Kim YS (2016) Identification and evaluation of age-correlated DNA methylation markers for forensic use. Forensic Sci Int Genet 23:64–70. https://doi.org/10.1016/j.fsigen.2016.03.005
Huang Y, Yan J, Hou J, Fu X, Li L, Hou Y (2015) Developing a DNA methylation assay for human age prediction in blood and bloodstain. Forensic Sci Int Genet 17:129–136. https://doi.org/10.1016/j.fsigen.2015.05.007
Funding
This research was supported by the National Natural Science Foundation of China (No. 82072121).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
This study was approved by the Medical Ethics Committee of Tongji Medical College, Huazhong University of Science and Technology (2018-IEC-S099).
Consent to participate
Written informed consent was obtained from all individual participants included in the study and/ or from legal representatives.
Conflict of interest
The authors declare no competing interests.
Research involving human participants and/or animals.
All procedures performed in studies involving human blood samples were in concordance with the ethical standards of the institutional research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. This article does not contain any studies with animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Han, X., Xiao, C., Yi, S. et al. Accurate age estimation from blood samples of Han Chinese individuals using eight high-performance age-related CpG sites. Int J Legal Med 136, 1655–1665 (2022). https://doi.org/10.1007/s00414-022-02865-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-022-02865-3