Abstract
When decomposition of a recovered body is fairly advanced, identification based on common morphologic features is often impossible. In these cases, short tandem repeat (STR) marker genotyping has established itself as a convenient and reliable alternative. However, at very progressed stages of decomposition, postmortem tissue putrefaction processes can decrease DNA yields considerably. Hence, not all types of tissue are equally suitable for successful STR marker-based postmortem identification.
Bone or dental material is often analysed in corpses with advanced decompositional changes. However, processing of these materials is very elaborate and time and resource consuming. We have therefore focused on the suitableness of various types of soft tissue swabs, where DNA extraction is easier and faster. By sampling 28 bodies at various stages of decomposition, we evaluated the suitability of different tissues for genotyping at varying degrees of physical decay. This was achieved by a systematic classification of the sampled bodies by morphological scoring and subsequent analysis of multiple tissue swabs of the aortic wall, urinary bladder wall, brain, liver, oral mucosa and skeletal muscle.
In summary, we found variable degrees of suitability of different types of soft tissue swabs for DNA-based identification. Swabs of the aortic wall, the urinary bladder wall and brain tissue yielded the best results — in descending order — even at advanced levels of decay.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A reliable and rapid forensic identification after the recovery of a human body is a crucial and morally obligatory task in many respects. However, when decomposition processes are already fairly advanced and common morphological identifying features are no longer reliable, alternative methods are required. Here, in most cases even with highly decomposed bodies, the identity of a human corpse can still be established by DNA profiling — more specifically by short tandem repeat (STR) marker profiling [1,2,3,4]. Still, DNA yields decrease with progressing decay due to microorganism activity and autolytic processes such as DNAse activity, hydrolysis and oxidation [5]. Some kinds of tissue are known to be less susceptible to these processes than others. Therefore, the choice of tissue is relevant to gain suitable material for achieving the task of forensic DNA profiling.
When bodies are highly decomposed or even skeletonised or mummified, DNA extracted from dental or bone material is a valuable approach for a successful forensic identification [6,7,8,9,10,11]. However, the process of DNA extraction from this tissue type is very elaborate and time and resource consuming. DNA extraction from soft tissues, on the other hand, is faster and less extensive. Another valuable source for DNA typing of highly decomposed bodies is human fingernails or toenails [12,13,14]. Here, DNA extraction is similarly unsophisticated. But with this type of tissue, contamination can be an issue, especially in criminal cases with rape scenarios before death. Moreover, at advanced stages of decomposition, fingernails and toenails can get detached from the body and can possibly not be ascribed with undisputable certainty to a recovered body.
Several studies centre on the suitability of different human bone types or specific dental tissue for a successful forensic identification [15,16,17,18,19,20,21]. Although the success of STR typing, dependent on the progress of decay, has generally been investigated in soft tissue [2, 22,23,24,25,26,27,28], literature directly comparing different soft tissue types at various stages of decomposition for forensic identification purposes is sparse [29, 30]. Therefore, we focused on a selection of swabs from soft tissues, such as the aortic wall, the urinary bladder wall, brain tissue, liver tissue and skeletal muscle tissue to gain more information about the reliability of those tissues even at more advanced stages of decomposition. Tissue selection was based on experiences gained in our routine casework. Aorta and urinary bladder are well-protected hollow organs, easily swabbed. The brain, liver and muscle were selected as they originate from different body regions and underly very different internal decomposition influences. Oral swabs were collected as a reference. Swabs were chosen as sampling method, as this is a straightforward, common, valid standard method in most DNA laboratories. Other than that, this sampling method is fast and easily applicable during autopsies. Furthermore, DNA extraction from swabs is less elaborate than DNA extraction from original tissue samples. There are several environmental factors that can influence decomposition of the whole human body as one entity but also degradation of single organs and tissue types can vary based on changed external influences. Those environmental factors include clothing, temperature, humidity, insect- or microorganism-activity, scavenging and body composition before death [31,32,33,34,35,36]. To achieve a better comparability between cases and sampled tissue types for this study, we exclusively selected cases of highly decomposed bodies that were discovered indoors. Another factor to achieve a better comparability and to ensure improved evaluation of these tissue types is a categorization of the state of decomposition of the bodies analysed in this study. To additionally implement this factor into our investigation, two separate scoring methods based on morphological features of ongoing decay were used: The Total Decomposition Score (TDS) and the Total Body Score (TBS) [37, 38]. The first scoring method by Gelderman et. al provides a guide to calculate a “Total Decomposition Score” (TDS). This score represents the sum of three separately evaluated regional scores: “facial decomposition score” (FDS), “body decomposition score” (BDS) and the “limbs decomposition score” (LDS). These have specific point-values depending on morphological features in relation to the decomposition process, ranging from 3 (fresh) to 18 (complete skeletonization) possible points [37]. The second scoring method by Megyesi et al. has similar features. In this case, the values representing the separately evaluated regional scores add up to the Total Body Score (TBS) ranging from 3 (fresh) to 35 (dry bone) points, also depending on the degree of decomposition [38]. The aim of this study was to establish whether a swab of a certain type of soft tissue is still suitable for a valid forensic identification even at advanced degrees of putrefaction.
Materials and methods
Sampling and morphological scoring
In this study, samples of in total 28 bodies with a higher degree of decomposition were taken (Tables 1 and 2). Those bodies were sampled and analysed between April 2019 and February 2020 at the Institute for Forensic Medicine in Salzburg, Austria. We chose a variety of soft tissues for sampling and subsequent DNA profiling. Sampling was mostly of an invasive nature during the autopsy of our cases. Two swabs each (one as backup) of the aortic wall, the urinary bladder wall, brain tissue, liver tissue and skeletal muscle tissue were taken as well as oral swabs. The muscle samples were all taken from the Musculus vastus lateralis. In this manner, a set of 336 samples was produced and further processed. To accomplish a systematic classification of the decomposition state of all analysed bodies, the degree of decomposition was evaluated with the two specific scoring methods based on morphologic features TDS and TBS.
DNA extraction
DNA extraction of all samples was performed with the GEN-IAL ®—First DNA all-tissue DNA Kit according to the manufacturer’s instructions [39].
DNA quantification and amplification
DNA extracts were quantified using the Quantifiler™ Trio DNA Quantification Kit (Thermo Fisher Scientific Corporation) and a 7500 Real-Time PCR System for Human Identification (Thermo Fisher Scientific Corporation). Subsequently, PCR amplification was performed on a Gene Amp PCR Systems 9700 system using the AmpFlSTR™ NGM SElect™ PCR Amplification Kit (Thermo Fisher Scientific Corporation) at 29 cycles with max 1 ng of DNA input based on the DNA yields determined in the quantification step. For all samples, two individual PCR runs were performed as internal quality control and to eliminate possible methodological errors.
Capillary electrophoresis and data analysis
For fragment separation, a mixture of 2–5 µl amplified PCR product, 23.5 µl HiDiTM formamide (Thermo Fisher Scientific Corporation) and 1.5 µl GeneScanTM 600 LIZ ® dye Size Standard v2.0 (Thermo Fisher Scientific Corporation) was injected in an ABI 3500 Genetic Analyzer (Thermo Fisher Scientific Corporation). The obtained DNA STR marker profiles were then analysed, and profile quality was evaluated (GeneMapper ® ID-X software Version 1.4, Thermo Fisher Scientific Corporation). Here, the minimum signal strength analysed was 50 relative fluorescent units (RFU). Profile completeness was calculated as the percentage of observed alleles out of a full STR marker profile. A full profile is represented by 68 alleles in total: 17 STR marker systems with two heterozygous or homozygous single alleles per system in two PCR approaches. Average success rates per tissue type were then accessorily calculated.
DNA re-extraction and re-amplification
Of all samples that showed very low DNA yields or signs of contamination, the backup swab was additionally analysed (n = 22) in the same manner as the first swab. If the second swab was also of poor quality, or samples still showed signs of contamination, they were labelled accordingly and excluded from further evaluation.
Secondary amplification and RFU analysis
In a next step, all samples that provided a higher DNA concentration yield than 0.1 ng/µl and a full 17 locus DNA profile (34 alleles out of 17 STR Marker systems in a PCR double approach adding up to 68 alleles) were further processed in the following manner: Samples were diluted with DNA-free water to achieve a standardised concentration of 0.1 ng/µl. After that, another amplification step was performed in the same setup as mentioned above with 5 µl DNA (= 500 pg) input. For a subsequent fragment separation step, 2 µl of the amplified products was used. To obtain an additional parameter for evaluation of the obtained DNA profiles of all tissue kinds, DNA degradation was evaluated by RFU analysis. The RFU per allele was calculated as the average of the heterozygous peak heights or the homozygous peak height divided by two. The max:min ratio was then calculated by dividing the value of the highest RFU per allele of all loci within an electropherogram of a sample with the lowest RFU per allele of all loci in the same electropherogram [17]. This was performed with both PCR approaches per sample and then an average was calculated. Statistical calculations were performed with Microsoft Excel 2019 (Version 16.33).
Results
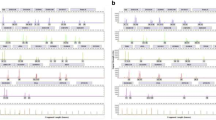
In general, success rates were rather high regarding STR marker profile completeness (see Table 2 and Fig. 1). The swab of the aortic wall showed the best results with a 99.5% average success rate in STR marker profile completeness. Sampling the urinary bladder led to 25 out of 28 samples with full profiles (average success rate of 99.3%). The brain and liver swabs showed quite similar results with 95.5% and 96.8%, followed by the muscle samples with 89.8%. The oral swabs did provide the least successful profile completeness rate with only 74.2%. As sampling the aorta provided a full profile in 27 out of 28 cases, this is not surprising. To gain information about the degradation of the analysed samples, a max:min ratio of the RFU values was calculated. The higher the max:min ratio, the higher the difference between peak heights within an electropherogram of one sample, indicating a “ski-slope-effect” and therefore higher degradation. This effect is caused by the observation that in degraded DNA, signal intensities decrease with increasing fragment length [40]. To ensure a better comparability between samples, only samples with a concentration above 0.1 ng/µl were further processed with standardised conditions. Here, as shown in Fig. 2 and Table 3, the liver samples showed the highest degradation compared to other samples and a quite high standard deviation. The degradation values calculated for the aorta samples are the lowest compared to the other soft tissue samples. Our samples were evaluated with morphological scores ranging from 6 to 13 (TDS) and 6 to 25 (TBS) (see Table 1, Supplementary Table S2). Up to a TDS of 9 and a TBS of 14, all sampled tissues do still provide a full quality STR marker profile. All sampled cases except one with a TDS below 10 and a TBS value below 16 do provide a profile completeness of more than 95%. If the TDS score increases above 11 and the TBS above 18 at least one of the sampled soft tissue types does not provide a 100% quality profile. The oral swab does not provide a 100% profile completeness in all but one sample with a TDS above 10 and a TBS higher than 16. In 27 out of 28 cases, the aorta samples did provide a full profile even at the highest TDS and TBS scores analysed. The same picture applies for the urinary bladder wall swabs with a profile completeness of 95% in all samples but one.
Discussion
To ensure a valid and rapid forensic identification, the selection of tissue type for a DNA analysis seems to be an important factor. With progressing decomposition in a human body, choices are increasingly limited due to tissue putrefaction and advanced DNA degradation processes. Bone and dental material are recommendable and widely used as the tissue of choice and at some point, during these processes when skeletonization or mummification is already fairly advanced, those hard tissue types are the only tissues available for sampling. However, if still possible, the sampling of soft tissues for forensic identification through DNA typing is by far less elaborate. In the present study, six types of soft tissue swabs sampled from highly decomposed bodies were examined: aortic wall, urinary bladder wall, brain, liver, skeletal muscle and oral mucosa. To achieve this, profile quality and DNA degradation from all tissue swabs were analysed and a possible correlation with two different degradation scores based on morphologic features was evaluated.
For amplification and subsequent generation of STR marker profiles, solely the AmpFlSTR™ NGM SElect™ PCR Amplification Kit was used as we have positive experiences with using this kit for the identification of heavily putrefied bodies. For the application in routine forensic laboratory case work, it is recommendable though to use a combination of kits (e.g. in combination with the Power Plex® ESI 17 Pro System).
To answer the research question of this study, it is quite important to “quantify” and categorise the state of decomposition of the recovered bodies. Both scoring methods, the TDS and the TBS, are based on specific morphologic phenomena which are linked to certain stages of decomposition and both showed similar results for each body analysed in this study. Notably, organ or tissue decomposition can differ from overall morphological decomposition scoring. However, in regard of the lack of decomposition scoring methods for these units, a global decomposition score comprising three body regions was considered best practice.
The results of the current study show that the aortic swabs provided the best profile completeness at all analysed levels of decay with a 99.5% success rate of achieving a full STR marker profile with a sample size of 28 analysed bodies. Sato et al. found that the aorta is one of the most useful samples for forensic identification after analysing 47 bodies with a variety of causes of death, including drowning and burning [26, 41]. These findings are supported by the results of a study of Shintani-Ishida analysing blood vessels in general as a promising source for DNA typing [42]. Thus, the aortic wall seems to be a quite resilient and dependable type of tissue for sampling even under severe conditions. Another reliable tissue under challenging forensic conditions is the urinary bladder. It is the tissue of choice for the identification of burnt bodies due to its anatomically protected situation within the body [23]. As antemortem bacterial colonization in this area is highly suppressed by innate immune system activity, also postmortem decay is delayed until very advanced body degradation. This could explain why the urinary bladder wall swabs showed the second-best results regarding profile completeness and low degradation values compared to other tissues analysed in this study.
Brain tissue provided a 95.5% average success rate in this study and good-quality DNA even at higher TDS/TBS rates. This could be explained by its enclosed location in the skull, preserving it from environmental factors such as bacterial or insect colonization for a longer time period than abdominal or thoracic organs. Other than that, the brain mass is not only protected by the ossa cranii but also by the dura mater. Motani et al. showed in their study that the dura mater itself is still a good source for DNA profiling even at higher degrees of decomposition [25]. Likewise, Ludes et al. and Pooniya et al. denominated brain tissue as a good source for DNA fingerprinting [30, 43].
Enhanced postmortal enzymatic and bacterial activity can be seen in the area of another organ analysed in this study: the liver. Due to these DNA degrading processes, this type of tissue is not highly dependable for DNA typing. Our results showed only in 22 out of 28 cases a full STR marker profile. In a study by Hoff-Olsen et al., incomplete profiles could be seen in more than half of their investigated cases [2]. Insect activity and especially maggot infestation are another factor that can severely influence soft tissue quality for DNA profiling. As postmortem, maggots enter the body primarily through natural body openings like the mouth, nose and the eyes, and putrefaction processes can be accelerated in these areas. Antemortem, a buccal swab is standard procedure for a fast and valid identification of a person. But during decomposition, the amount of liquid decompositional products is quite high which can make sampling difficult. Together with the aforementioned insect activity, this leads to poor-quality DNA extraction of oral swabs of highly decomposed bodies.
The DNA profiling results for skeletal muscle tissue are oscillating rather much compared to other soft tissue types. This phenomenon could also be observed in other studies [24, 27].
When TDS/TBS values are correlated to profile completeness, our data suggest that all investigated tissues are viable sources for DNA sampling up to a TDS/TBS of 9/14 (e.g. grey to green discoloration of the skin, early bloating stages) and respectively 95% profile completeness up to 10/16 (e.g. purging of decompositional fluids, brown to black discoloration of limbs). All cases with higher scores failed to yield complete profiles from oral swabs, and this source should thus be avoided. Aorta and urinary bladder performed extremely well, with only a single case yielding under 95% profile completeness respectively. Notably, these were not the cases with the highest overall decomposition scores (12/16 and 12/18, e.g. brown discoloration of the head and neck, sagging of flesh, caving in of the abdominal cavity), suggesting additional influences. Muscle, the brain and liver also performed surprisingly well, with average success rates of around 90–95%. Although there was a trend observed that cases with higher TDS/TBS values produced less conclusive outcome, even some with very high values (e.g. 13/21, 12/25, e.g. bone exposure, (partial) mummification) yielded complete profiles in these tissues.
After sampling hard tissues, like bone or dental material, a good comparability of DNA yields can be achieved by using similar amounts of e.g. bone powder for subsequent DNA extraction. With soft tissue samples, it is not as easy to find a good sampling method that satisfies this criterium. The utilization of swabs is a fast and easy standard sampling method for soft tissues in general and for the forensic identification of fresh human bodies. Therefore, we chose this sampling method in our study. However, DNA concentration values after the extraction process cannot be conclusively compared as while swabbing various types of soft tissue in various stages of decay, it is not possible to sample a comparable “amount” of tissue. Therefore, we did not compare DNA yields in this study and “standardised” samples above 0.1 ng/µl for further analysis to ensure a better comparability. Original DNA yields are summarised in Supplementary Table S1.
In general, the application of swabs for sampling soft tissues of highly decomposed bodies proved to be a valid and convenient method. Nevertheless, future research, directly comparing the tissue swabs to direct tissue extraction, could improve understanding of the suitability of tissue swabs.
The decay of bodies in an outdoor environment is influenced by more factors than the decay of bodies degrading indoors, which alters the decomposition rate and consequently the condition of tissue types more intensely [34]. Nevertheless, also indoors, there are still many environmental factors and inter-individual variation that influence degradation processes [44] and this can be seen as a limitation of this study. Increasing the sample size could average out the large number of variables which can affect DNA preservation. Based on our results, we recommend the sampling of the aorta, urinary bladder or brain tissue for the identification of highly decomposed bodies by DNA fingerprinting. However, further research is required in this field and the comparison of these tissue types with other tissues such as cartilage, Achilles tendon, fingernails and toenails and ocular swabs is desirable as those also have been labelled as useful for DNA extraction in cases of putrefied unidentifiable bodies [24, 28, 45].
Conclusion
In this study, we discovered differences in the suitability of a variety of soft tissue type swabs for the forensic identification of highly decomposed bodies by DNA profiling. Swabs of the aortic wall, the urinary bladder wall and brain tissue showed the most promising results respectively. Swabs of muscle and liver tissue did not provide stable results for viable profiling at varying degrees of decomposition. Oral swabs are highly recommendable for identification purposes antemortem but not postmortem when increased levels of decay are reached.
References
Butler JM, Butler JM (2010) Fundamentals of forensic DNA typing. Academic Press/Elsevier Amsterdam, Boston
Hoff-Olsen P, Jacobsen S, Mevag B, Olaisen B (2001) Microsatellite stability in human post-mortem tissues. Forensic Sci Int 119:273–278
Bär W, Hummel K (1991) DNA fingerprinting: its application in forensic case work. EXS 58:349–355. https://doi.org/10.1007/978-3-0348-7312-3_25
Bär W, Kratzer A, Machler M, Schmid W (1988) Postmortem stability of DNA. Forensic Sci Int 39:59–70. https://doi.org/10.1016/0379-0738(88)90118-1
Lindahl T (1993) Instability and decay of the primary structure of DNA. Nature 362:709–715. https://doi.org/10.1038/362709a0
Andelinovic S, Sutlovic D, Erceg Ivkosic I et al (2005) Twelve-year experience in identification of skeletal remains from mass graves. Croat Med J 46:530–539
Boles TC, Snow CC, Stover E (1995) Forensic DNA testing on skeletal remains from mass graves: a pilot project in Guatemala. J Forensic Sci 40:349–355
Hagelberg E, Gray IC, Jeffreys AJ (1991) Identification of the skeletal remains of a murder victim by DNA analysis. Nature 352:427–429. https://doi.org/10.1038/352427a0
Smith BC, Fisher DL, Weedn VW, Warnock GR, Holland MM (1993) A systematic approach to the sampling of dental DNA. J Forensic Sci 38:1194–1209
Rubio L, Santos I, Gaitan MJ, Martin de-las Heras S, (2013) Time-dependent changes in DNA stability in decomposing teeth over 18 months. Acta Odontol Scand 71:638–643. https://doi.org/10.3109/00016357.2012.700068
Jakubowska J, Maciejewska A, Pawlowski R (2012) Comparison of three methods of DNA extraction from human bones with different degrees of degradation. Int J Legal Med 126:173–178. https://doi.org/10.1007/s00414-011-0590-5
Inkret J, Podovsovnik E, Zupanc T, Pajnic IZ (2020) Nails as a primary sample type for molecular genetic identification of highly decomposed human remains. Int J Legal Med. https://doi.org/10.1007/s00414-020-02289-x
Allouche M, Hamdoum M, Mangin P, Castella V (2008) Genetic identification of decomposed cadavers using nails as DNA source. Forensic Sci Int Genet 3:46–49. https://doi.org/10.1016/j.fsigen.2008.07.008
Piccinini A, Cucurachi N, Betti F et al (2006) Forensic DNA typing of human nails at various stages of decomposition. Int Congr Ser 1288:586–588. https://doi.org/10.1016/j.ics.2005.08.029
Alonso A, Andelinovic S, Martin P et al (2001) DNA typing from skeletal remains: evaluation of multiplex and megaplex STR systems on DNA isolated from bone and teeth samples. Croat Med J 42:260–266
Malaver PC, Yunis JJ (2003) Different dental tissues as source of DNA for human identification in forensic cases. Croat Med J 44:306–309
Mundorff A, Davoren JM (2014) Examination of DNA yield rates for different skeletal elements at increasing post mortem intervals. Forensic Sci Int Genet 8:55–63. https://doi.org/10.1016/j.fsigen.2013.08.001
Ricaut FX, Keyser-Tracqui C, Crubezy E, Ludes B (2005) STR-genotyping from human medieval tooth and bone samples. Forensic Sci Int 151:31–35. https://doi.org/10.1016/j.forsciint.2004.07.001
Cho S, Shin KJ, Bae SJ, Kwon YL, Lee SD (2020) Improved STR analysis of degraded DNA from human skeletal remains through in-house MPS-STR panel. Electrophoresis. https://doi.org/10.1002/elps.202000070
Ambers A, Turnbough M, Benjamin R et al (2016) Modified DOP-PCR for improved STR typing of degraded DNA from human skeletal remains and bloodstains. Leg Med (Tokyo) 18:7–12. https://doi.org/10.1016/j.legalmed.2015.10.013
Milos A, Selmanovic A, Smajlovic L et al (2007) Success rates of nuclear short tandem repeat typing from different skeletal elements. Croat Med J 48:486–493
Uerlings S, Welter V, Madea B, Grabmuller M (2021) Comparative analysis of DNA extraction processes for DNA-based identification from putrefied bodies in forensic routine work. Forensic Sci Int 320:110707. https://doi.org/10.1016/j.forsciint.2021.110707
Ferreira STG, Paula KA, Nogueira RF, Oliveira ES, Moraes AV (2015) A comparative study between muscle, cartilage and swab from inside the urinary bladder samples for DNA typing of severely burnt bodies in disaster victim identification (DVI). Forens Sci Int-Gen S 5:E617–E618. https://doi.org/10.1016/j.archoralbio.2015.09.243
Trindade-Filho A, Mendes C, Ferreira S et al (2008) DNA obtained from decomposed corpses cartilage: a comparison with skeleton muscle source. Forens Sci Int-Gen S 1:459–461. https://doi.org/10.1016/j.fsigss.2007.10.210
Motani H, Sakurada K, Akutsu T et al (2006) Usefulness of dura mater in providing DNA samples for identifying cadavers. J Forensic Sci 51:888–892. https://doi.org/10.1111/j.1556-4029.2006.00157.x
Sato Y, Motani H, Inoue H et al (2009) Multiplex STR typing of aortic tissues from unidentified cadavers. Leg Med 11:S455–S457. https://doi.org/10.1016/j.legalmed.2009.02.064
Rerkamnuaychoke B, Chantratita W, Jomsawat U, Thanakitgosate J, Pattanasak N, Rojanasunan P (2000) Comparison of DNA extraction from blood stain and decomposed muscle in STR polymorphism analysis. J Med Assoc Thai 83(Suppl 1):S82–S88
Roeper A, Reichert W, Mattern R (2007) The Achilles tendon as a DNA source for STR typing of highly decayed corpses. Forensic Sci Int 173:103–106. https://doi.org/10.1016/j.forsciint.2007.02.004
Schwark T, Heinrich A, von Wurmb-Schwark N (2011) Genetic identification of highly putrefied bodies using DNA from soft tissues. Int J Legal Med 125:891–894. https://doi.org/10.1007/s00414-010-0537-2
Pooniya S, Lalwani S, Raina A, Millo T, Dogra TD (2014) Quality and quantity of extracted deoxyribonucleic acid (DNA) from preserved soft tissues of putrefied unidentifiable human corpse. J Lab Physicians 6:31–35. https://doi.org/10.4103/0974-2727.129088
Campobasso CP, Di Vella G, Introna F (2001) Factors affecting decomposition and Diptera colonization. Forensic Sci Int 120:18–27. https://doi.org/10.1016/s0379-0738(01)00411-x
Johnson AP, Mikac KM, Wallman JF (2013) Thermogenesis in decomposing carcasses. Forensic Sci Int 231:271–277. https://doi.org/10.1016/j.forsciint.2013.05.031
Matuszewski S, Konwerski S, Fratczak K, Szafalowicz M (2014) Effect of body mass and clothing on decomposition of pig carcasses. Int J Legal Med 128:1039–1048. https://doi.org/10.1007/s00414-014-0965-5
Suckling JK, Spradley MK, Godde K (2016) A longitudinal study on human outdoor decomposition in Central Texas. J Forensic Sci 61:19–25. https://doi.org/10.1111/1556-4029.12892
Sutherland A, Myburgh J, Steyn M, Becker PJ (2013) The effect of body size on the rate of decomposition in a temperate region of South Africa. Forensic Sci Int 231:257–262. https://doi.org/10.1016/j.forsciint.2013.05.035
Zhou C, Byard RW (2011) Factors and processes causing accelerated decomposition in human cadavers - an overview. J Forensic Leg Med 18:6–9. https://doi.org/10.1016/j.jflm.2010.10.003
Gelderman HT, Boer L, Naujocks T, ACM IJ, Duijst W, (2018) The development of a post-mortem interval estimation for human remains found on land in the Netherlands. Int J Legal Med 132:863–873. https://doi.org/10.1007/s00414-017-1700-9
Megyesi MS, Nawrocki SP, Haskell NH (2005) Using accumulated degree-days to estimate the postmortem interval from decomposed human remains. J Forensic Sci 50:618–626
(2007) GEN-IAL —All-tissue DNA Kit. Handbuch für DNA-Extraktion. pp. 11–5.
Tvedebrink T, Eriksen PS, Mogensen HS, Morling N (2012) Statistical model for degraded DNA samples and adjusted probabilities for allelic drop-out. Forensic Sci Int Genet 6:97–101. https://doi.org/10.1016/j.fsigen.2011.03.001
Sato Y, Hayakawa M, Nakajima T, Motani H, Kiuchi M (2003) HLA typing of aortic tissues from unidentified bodies using hot start polymerase chain reaction-sequence specific primers. Leg Med (Tokyo) 5(Suppl 1):S191–S193. https://doi.org/10.1016/s1344-6223(02)00108-6
Shintani-Ishida K, Harada K, Nakajima M, Yoshida K (2010) Usefulness of blood vessels as a DNA source for PCR-based genotyping based on two cases of corpse dismemberment. Leg Med (Tokyo) 12:8–12. https://doi.org/10.1016/j.legalmed.2009.09.004
Ludes B, Pfitzinger H, Mangin P (1993) DNA fingerprinting from tissues after variable postmortem periods. J Forensic Sci 38:686–690
Anderson GS (2011) Comparison of decomposition rates and faunal colonization of carrion in indoor and outdoor environments. J Forensic Sci 56:136–142. https://doi.org/10.1111/j.1556-4029.2010.01539.x
Agostini V, Bailo P, Chiti E et al (2020) Ocular swabs on exhumed bodies: an alternative to the collection of “classical” tissue samples in forensic genetics. Forensic Sci Int Genet 44:102206. https://doi.org/10.1016/j.fsigen.2019.102206
Acknowledgements
We highly appreciate the help and support of the medical pathology assistants (namely Daniela Besic and Michael Peherstorfer) in our institute. Without them, this study would not have been possible. Additionally, we want to thank the whole team of our DNA laboratory.
Funding
Open access funding provided by Paris Lodron University of Salzburg.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All applicable international, national and/or institutional guidelines were followed.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Helm, K., Matzenauer, C., Neuhuber, F. et al. Suitability of specific soft tissue swabs for the forensic identification of highly decomposed bodies. Int J Legal Med 135, 1319–1327 (2021). https://doi.org/10.1007/s00414-021-02601-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-021-02601-3