Abstract
The inference of biogeographical ancestry (BGA) can provide useful information for forensic investigators when there are no suspects to be compared with DNA collected at the crime scene or when no DNA database matches exist. Although public databases are increasing in size and population scope, there is a lack of information regarding genetic variation in Eurasian populations, especially in central regions such as the Middle East. Inhabitants of these regions show a high degree of genetic admixture, characterized by an allele frequency cline running from NW Europe to East Asia. Although a proper differentiation has been established between the cline extremes of western Europe and South Asia, populations geographically located in between, i.e, Middle East and Mediterranean populations, require more detailed study in order to characterize their genetic background as well as to further understand their demographic histories. To initiate these studies, three ancestry informative SNP (AI-SNP) multiplex panels: the SNPforID 34-plex, Eurasiaplex and a novel 33-plex assay were used to describe the ancestry patterns of a total of 24 populations ranging across the longitudinal axis from NW Europe to East Asia. Different ancestry inference approaches, including STRUCTURE, PCA, DAPC and Snipper Bayes analysis, were applied to determine relationships among populations. The structure results show differentiation between continental groups and a NW to SE allele frequency cline running across Eurasian populations. This study adds useful population data that could be used as reference genotypes for future ancestry investigations in forensic cases. The 33-plex assay also includes pigmentation predictive SNPs, but this study primarily focused on Eurasian population differentiation using 33-plex and its combination with the other two AI-SNP sets.


Similar content being viewed by others
References
Kayser M, de Knijff P (2011) Improving human forensics through advances in genetics, genomics and molecular biology. Nat Rev Genet 12:179–192
Phillips C (2015) Forensic genetic analysis of bio-geographical ancestry. Forensic Sci. Int. Genet. doi: 10.1016/j.fsigen.2015.05.012 (e-pub ahead of print)
Phillips C (2013) Ancestry informative markers. In: J.A. Siegel, P.J. Saukko (Eds.) Encyclopedia of Forensic Sciences, 2nd ed., vol. 1, Academic Press, 323–331
Phillips C, Salas A, Sánchez JJ, Fondevila M, Gómez-Tato A, Álvarez-Dios J, Calaza M, Casares de Cal M, Ballard D, Lareu MV, Carracedo Á (2007) The SNPforID consortium, inferring ancestral origin using a single multiplex assay of ancestry—informative marker SNPs. Forensic Sci Int Genet 1:273–280
Halder I, Shriver M, Thomas M, Fernandez JR, Frudakis TN (2008) A panel of ancestry informative markers for estimating individual biogeographical ancestry and admixture from four continents: utility and applications. Hum Mutat 29(5):648–658
Kersbergen P, van Duijn K, Kloosterman AD, den Dunnen JT, Kayser M, de Knijff P (2009) Developing a set of ancestry-sensitive DNA markers reflecting continental origins of humans. BMC Genet 10:69
Phillips C, Freire Aradas A, Kriegel AK, Fondevila M, Bulbul O, Santos C, Serrulla Rech F, Perez Carceles MD, Carracedo Á, Schneider PM, Lareu MV (2013) Eurasiaplex: a forensic SNP assay for differentiating European and South Asian ancestries. Forensic Sci Int Genet 7:359–366
Reich D, Thangaraj K, Patterson N, Price AL, Singh L (2009) Reconstructing Indian population history. Nature 461:489–494
Zerjal T, Spencer Wells R, Yuldasheva N, Ruzibakiev R, Tyler-Smith C (2002) A genetic landscape reshaped by recent events: Y-chromosomal insights into Central Asia. Am J Hum Genet 71:466–482
Spencer Wells R, Yuldasheva N, Ruzibakiev R, Underhill PA, Evseeva I, Blue-Smith J, Jin L, Su B, Pitchappan R, Shanmugalakshmi S et al (2001) The Eurasian heartland: a continental perspective on Y-chromosome diversity. Proc Natl Acad Sci U S A 98:10244–10249
Fondevila M, Phillips C, Santos C, Freire Aradas A, Vallone PM, Butler JM, Lareu MV, Carracedo Á (2013) Revision of the SNPforID 34-plex forensic ancestry test: assay enhancements, standard reference sample genotypes and extended population studies. Forensic Sci Int Genet 7:63–74
J. Amigo, A. Salas, C. Phillips, Á. Carracedo (2008) SPSmart: adapting population based SNP genotype databases for fast and comprehensive web access, BMC Bioinformatics, 9 428. URL: http://spsmart.cesga.es, accessed September 2014
Liu F, van Duijn K, Vingerling JR, Hofman A, Uitterlinden AG, Janssens ACJW, Kayser M (2009) Eye color and the prediction of complex phenotypes from genotypes. Curr Biol 19:192–193
Walsh S, Liu F, Ballantyne KN, van Oven M, Lao O, Kayser M (2011) Irisplex: a sensitive DNA tool for accurate prediction of blue and brown eye colour in the absence of ancestry information. Forensic Sci Int Genet 5:170–180
Bulbul O, Filoglu G, Altuncul H, Aradas AF, Ruiz Y, Fondevila M, Phillips C, Carracedo Á, Kriegel AK, Schneider PM (2011) A SNP multiplex for the simultaneous prediction of biogeographic ancestry and pigmentation type. Forensic Sci Int Genet Supp Ser 3:500–501
Ruiz Y, Phillips C, Gomez-Tato A, Alvarez-Dios J, Casares de Cal M, Cruz RR, Maroñas O, Söchtig J, Fondevila M, Rodriguez-Cid MJ, Carracedo Á, Lareu MV (2013) Further development of forensic eye color predictive tests. Forensic Sci Int Genet 7:28–40
Untergrasser A, Cutcutache I, Koressaar T, Ye J, Faircloth BC, Remm M, Rozen SG (2012) Primer3—new capabilities and interfaces. Nucleic Acids Res 40:e115
Vallone PM, Butler JM (2004) AutoDimer: a screening tool for primer–dimer and hairpin structures. Biotechniques 37:226–231
Excoffier LG, Schneider S (2005) Arlequin v. 3.0: an integrated software package for population genetics data analysis. Evol Bioinformatics Online 1:47–50
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population STRUCTURE using multilocus genotype data. Genetics 155:945–959
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Core Team R (2014) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Gonzalez JR, Armengol L, Sole X, Guino E, Mercader JM, Estivill X, Moreno V (2007) SNPassoc: an R package to perform whole genome association studies. Bioinformatics 23:644–645
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:94
The Snipper app suite tool: http:// mathgene.usc.es/Snipper/ Cross validation: http://mathgene.usc.es/Snipper/analysispopfile2_new.html, custom ancestry classifications: Accessed September 2014
Jobling M, Hollox E, Hurles M, Kivisild T, Tyler-Smith C (2013) Human evolutionary genetics (2nd ed.). New York: Garland Science. Chapter 14: What happens when populations meet? 443-477 pp
Khodjet-el-Khil H, Fadhlaoui-Zid K, Cherni L, Phillips C, Fondevila M, Carracedo Á, Ben Ammar-Elgaaied A (2011) Genetic analysis of the SNPforID 34-plex ancestry informative SNP panel in Tunisian and Libyan populations. Forensic Sci Int Genet 3:e45–e47
Zalloua PA, Platt DE, El Siba M, Khalife J, Makhoul N, Haber M, Xue MY, Izaabel H, Bosch E et al (2008) Identifying genetic traces of historical expansions: Phoenician footprints in the Mediterranean. Am J Hum Genet 83:633–642
Lazaridis I, Patterson N, Mittnik A, Renaud G, Mallick S, Kirsanow K, Sudmant PH, Schraiber JG et al (2014) Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513:409–413
Kidd KK, Speed WC, Pakstis AJ, Furtado MR, Fang R, Madbouly A, Maiers M, Middha M, Friedlaender FR, Kidd JR (2014) Progress toward an efficient panel of SNPs for ancestry inference. Forensic Sci Int Genet 10:23–32
Kidd JR, Friedlaender FR, Speed WC, Pakstis AJ, De La Vega FM, Kidd KK (2011) Analyses of a set of 128 ancestry informative single-nucleotide polymorphisms in a global set of 119 population samples. Investig Genet 2:1
Acknowledgments
This research was supported by Istanbul University Scientific Research Projects Unit (BAP) under project number 19042. OB was supported by the German Academic Exchange Service (DAAD) under Research Grants for Doctoral Candidates and Young Academics and Scientists Programme and by The Scientific And Technological Research Council Of Turkey (TÜBİTAK) under 2211-Grant Programme. We sincerely thank all the sample contributors. The authors would like to thank Suad Alfadhli from the Department of Medical Laboratory Sciences, Faculty of Allied Health Sciences, Kuwait University, for collecting the Kuwait samples, Department of Criminalistic Investigations DNA Laboratory, Ministry of Internal Affairs of Azerbaijan Republic and Katharina Läer from the Department of Forensic Medicine, Hannover Medical School, Germany for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. S1
Allele frequency comparisons for Africa, Europe and East Asia for the selected 22 AI-SNPs using SPSmart ENGINES SNP browser (http://spsmart.cesga.es/search.php). One missing AI-SNP data (rs1343879) is not available in SPSmart and it added manually from 1000 Genome Phase3 data below the figure. (GIF 376 kb)
Supplementary Fig. S2
33-plex SNP profile for standard forensic control DNA 9947A (GIF 1645 kb)
Supplementary Fig. S3
Arlequin population divergence analyses of pairwise FST (blue squares), pairwise between-population differences (green) and pairwise within-population differences (orange) for 87 SNPs. Populations represented: Africa (LWK and YRI), Europe/Mediterranean (CEU, FIN, GBR, Germany, northwest Spain, Italy, Greece, Turkey, Russia-Adygei and Azerbaijan), Middle East (Algeria, Israel, Egypt, Kuwait, Libya, Yemen and Morocco), Central South Asia (BEB, GIH, ITU, PJL, STU, India, Pakistan and Uygur) and East Asia (Vietnam, Japan and China). Data was collected from 1000 Genomes, HGDP-CEPH and study populations as described in section 2.1. (GIF 2021 kb)
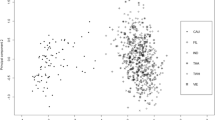
Supplementary Fig. S4
Principal component analysis of the 34-plex alone, 34-plex plus Eurasiaplex and 34-plex, Eurasiaplex plus 33-plex. (PDF 3419 kb)
Supplementary Fig. S5
Bayesian ancestry assignments for European vs. Middle Eastern populations using Snipper. Grey colours represent individuals from 16 populations clustered into three ancestries and ordered by likelihoods ratios. Three ancestry groups represented are: European (CEU; FIN; GBR; Germany and northwest Spain), Mediterranean (Italy: Bergamo, Sardinia and Tuscan; Azerbaijan; Greece, Adygei and Turkey) and Middle Eastern (Algeria; Israel: Druze, Palestinian and Bedouin; Egypt, Kuwait, Libya; Morocco and Yemen). (GIF 1418 kb)
Supplementary Table S1
Details of the 33-plex component SNPs. (XLSX 17 kb)
Supplementary Table S2
Bins and panels for the developed 33-plex using POP4. (XLSX 15 kb)
Supplementary Table S3
a. Cumulative informativeness values of each multiplex between five groups (Africa = AFR, Europe = EUR, Middle East = ME, Central-South Asia = CSA and East Asia = EAS) and values from combined sets. IN OVERALL is the divergence value for the comparison of all five population groups. b. Cumulative informativeness values of each multiplex between three groups and values from combined sets. (XLSX 12 kb)
Supplementary Table S4
Pairwise genetic distance matrix based on the FST values. (XLSX 23 kb)
Supplementary Table S5
Five-group training set for Snipper Bayesian analysis portal. Note this worksheet can be used ‘as is’, because rows 2-5 are not read by Snipper. (XLSX 372 kb)
Rights and permissions
About this article
Cite this article
Bulbul, O., Filoglu, G., Zorlu, T. et al. Inference of biogeographical ancestry across central regions of Eurasia. Int J Legal Med 130, 73–79 (2016). https://doi.org/10.1007/s00414-015-1246-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-015-1246-7