Abstract.
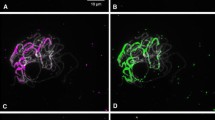
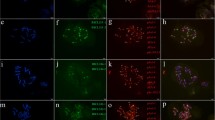
The physical distribution of ten simple-sequence repeated DNA motifs (SSRs) was studied on chromosomes of bread wheat, rye and hexaploid triticale. Oligomers with repeated di-, tri- or tetra-nucleotide motifs were used as probes for fluorescence in situ hybridization to root-tip metaphase and anther pachytene chromosomes. All motifs showed dispersed hybridization signals of varying strengths on all chromosomes. In addition, the motifs (AG)12, (CAT)5, (AAG)5, (GCC)5 and, in particular, (GACA)4 hybridized strongly to pericentromeric and multiple intercalary sites on the B genome chromosomes and on chromosome 4A of wheat, giving diagnostic patterns that resembled N-banding. In rye, all chromosomes showed strong hybridization of (GACA)4 at many intercalary sites that did not correspond to any other known banding pattern, but allowed identification of all R genome chromosome arms. Overall, SSR hybridization signals were found in related chromosome positions independently of the motif used and showed remarkably similar distribution patterns in wheat and rye, indicating the special role of SSRs in chromosome organization as a possible ancient genomic component of the tribe Triticeae (Gramineae).
Similar content being viewed by others

Author information
Authors and Affiliations
Additional information
Received: 13 February 1998; in revised form: 18 August 1998 / Accepted: 18 August 1998
Rights and permissions
About this article
Cite this article
Cuadrado, A., Schwarzacher, T. The chromosomal organization of simple sequence repeats in wheat and rye genomes. Chromosoma 107, 587–594 (1998). https://doi.org/10.1007/s004120050345
Issue Date:
DOI: https://doi.org/10.1007/s004120050345