Abstract
The abundance and chromosomal distribution of six class-II transposable elements (TEs) of Drosophila buzzatii have been analyzed by Southern blotting and in situ hybridization. These six transposons had been previously found at the breakpoints of inversions 2j and 2q 7 of D. buzzatii. These two polymorphic inversions were generated by an ectopic recombination event between two copies of Galileo, a Foldback element. The four breakpoints became hotspots for TE insertions after the generation of the inversion and the transposons analyzed in this work are considered to be secondary invaders of these regions. Insertions of the six transposons are present in the euchromatin but show an increased density in the pericentromeric euchromatin–heterochromatin transition region and the dot chromosome. They are also more abundant in the inverted segments of chromosome 2 rearrangements. We further observed that the accumulation of TE insertions varies between elements and is correlated between dot, proximal regions, and inverted segments. These observations fully agree with previous data in Drosophila melanogaster and support recombination rate as the chief force explaining the chromosomal distribution of TEs.


Similar content being viewed by others
References
Andolfatto P, Depaulis F, Navarro A (2001) Inversion polymorphisms and nucleotide variability in Drosophila. Genet Res 77:1–8
Ashburner M (1989) Drosophila. A laboratory handbook. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Bachtrog D (2003) Accumulation of Spock and Worf, two novel non-LTR retrotransposons, on the neo-Y chromosome of Drosophila miranda. Mol Biol Evol 20:173–181
Bartolomé C, Maside X (2004) The lack of recombination drives the fixation of transposable elements on the fourth chromosome of Drosophila melanogaster. Genet Res 83:91–100
Bartolomé C, Maside X, Charlesworth B (2002) On the abundance and distribution of transposable elements in the genome of Drosophila melanogaster. Mol Biol Evol 19:926–937
Biemont C, Cizeron G (1999) Distribution of transposable elements in Drosophila species. Genetica 105:43–62
Cáceres M, Barbadilla A, Ruiz A (1999a) Recombination rate predicts inversion size in Diptera. Genetics 153:251–259
Cáceres M, Ranz JM, Barbadilla A, Long M, Ruiz A (1999b) Generation of a widespread Drosophila inversion by a transposable element. Science 285:415–418
Cáceres M, Puig M, Ruiz A (2001) Molecular characterization of two natural hotspots in the Drosophila buzzatii genome induced by transposon insertions. Genome Res 11:1353–1364
Carmena M, González C (1995) Transposable elements map in a conserved pattern of distribution extending from beta-heterochromatin to centromeres in Drosophila melanogaster. Chromosoma 103:676–684
Casals F, Cáceres M, Ruiz A (2003) The foldback-like transposon Galileo is involved in the generation of two different natural chromosomal inversions of Drosophila buzzatii. Mol Biol Evol 20:674–685
Casals F, Cáceres M, Manfrin MH, González J, Ruiz A (2005) Molecular characterization and chromosomal distribution of Galileo, Kepler and Newton, three foldback transposable elements of the Drosophila buzzatii species complex. Genetics 169:2047–2059
Charlesworth B (1996) Background selection and patterns of genetic diversity in Drosophila melanogaster. Genet Res 68:131–149
Charlesworth B, Langley CH (1989) The population genetics of Drosophila transposable elements. Annu Rev Genet 23:251–287
Charlesworth B, Lapid A, Canada D (1992) The distribution of transposable elements within and between chromosomes in a population of Drosophila melanogaster. II. Inferences on the nature of selection against elements. Genet Res 60:115–130
Charlesworth B, Sniegowski P, Stephan W (1994) The evolutionary dynamics of repetitive DNA in eukaryotes. Nature 371:215–220
Craig NL (1997) Target site selection in transposition. Annu Rev Biochem 66:437–474
Daveran-Mingot ML, Campo N, Ritzenthaler P, Le Bourgeois P (1998) A natural large chromosomal inversion in Lactococcus lactis is mediated by homologous recombination between two insertion sequences. J Bacteriol 180:4834–4842
Dimitri P (1997) Constitutive heterochromatin and transposable elements in Drosophila melanogaster. Genetica 100:85–93
Dimitri P, Junakovic N, Arca B (2003) Colonization of heterochromatic genes by transposable elements in Drosophila. Mol Biol Evol 20:503–512
Eanes WF, Wesley C, Charlesworth B (1992) Accumulation of P elements in minority inversions in natural populations of Drosophila melanogaster. Genet Res 59:1–9
Evgen’ev MB, Zelentsova H, Poluectova H, Lyozin GT, Veleikodvorskaja V, Pyatkov KI, Zhivotovsky LA, Kidwell MG (2000) Mobile elements and chromosomal evolution in the virilis group of Drosophila. Proc Natl Acad Sci U S A 97:11337–11342
Francino O, Cabré O, Fontdevila A (1993) Distribution of the copia transposable element in the repleta group of Drosophila. Genet Sel Evol 25:501–516
Goldman AS, Lichten M (1996) The efficiency of meiotic recombination between dispersed sequences in Saccharomyces cerevisiae depends upon their chromosomal location. Genetics 144:43–55
Gordo I, Charlesworth B (2001) Genetic linkage and molecular evolution. Curr Biol 11:R684–686
Hill WG, Robertson A (1966) The effect of linkage on limits to artificial selection. Genet Res 8:269–294
Hoskins RA, Smith CD, Carlson JW, Carvalho AB, Halpern A, Kaminker JS, Kennedy C, Mungall CJ, Sullivan BA, Sutton GG, Yasuhara JC, Wakimoto BT, Myers EW, Celniker SE, Rubin GM, Karpen GH (2002) Heterochromatic sequences in a Drosophila whole-genome shotgun assembly. Genome Biol 3:research 0085.1–85.16
Junakovic N, Terrinoni A, Di Franco C, Vieira C, Loevenbruck C (1998) Accumulation of transposable elements in the heterochromatin and on the Y chromosome of Drosophila simulans and Drosophila melanogaster. J Mol Evol 46:661–668
Kaminker JS, Bergman CM, Kronmiller B, Carlson J, Svirskas R, Patel S, Frise E, Wheeler DA, Lewis SE, Rubin GM, Ashburner M, Celniker SE (2002) The transposable elements of the Drosophila melanogaster euchromatin: a genomics perspective. Genome Biol 3:research0084.1–84.2
Kim JM, Vanguri S, Boeke JD, Gabriel A, Voytas DF (1998) Transposable elements and genome organization: a comprehensive survey of retrotransposons revealed by the complete Saccharomyces cerevisiae genome sequence. Genome Res 8:464–478
Labrador M, Fontdevila A (1994) High transposition rates of Osvaldo, a new Drosophila buzzatii retrotransposon. Mol Gen Genet 245:661–674
Langley CH, Montgomery E, Hudson R, Kaplan N, Charlesworth B (1988) On the role of unequal exchange in the containment of transposable element copy number. Genet Res 52:223–235
Lathe WC 3rd, Burke WD, Eickbush DG, Eickbush TH (1995) Evolutionary stability of the R1 retrotransposable element in the genus Drosophila. Mol Biol Evol 12:1094–1105
Locke J, Podemski L, Roy K, Pilgrim D, Hodgetts RR (1999) Analysis of two cosmid clones from chromosome 4 of Drosophila melanogaster reveals two new genes amid an unusual arrangement of repeated sequences. Genome Res. 9:137–149
Maside X, Bartolome C, Assimacopoulos S, Charlesworth B (2001) Rates of movement and distribution of transposable elements in Drosophila melanogaster: in situ hybridization vs Southern blotting data. Genet Res 78:121–136
Montgomery E, Charlesworth B, Langley CH (1987) A test for the role of natural selection in the stabilization of transposable element copy number in a population of Drosophila melanogaster. Genet Res 49:31–41
Montgomery EA, Huang SM, Langley CH, Judd BH (1991) Chromosome rearrangement by ectopic recombination in Drosophila melanogaster: genome structure and evolution. Genetics 129:1085–1098
Navarro A, Betran E, Barbadilla A, Ruiz A (1997) Recombination and gene flux caused by gene conversion and crossing over in inversion heterokaryotypes. Genetics 146:695–709
Naveira H, Fontdevila A (1985) The evolutionary history of Drosophila buzzatii. IX. High frequencies of new chromosome rearrangements induced by introgressive hybridization. Chromosoma 91:87–94
Petrov DA, Aminetzach YT, Davis JC, Bensasson D, Hirsh AE (2003) Size matters: non-LTR retrotransposable elements and ectopic recombination in Drosophila. Mol Biol Evol 20:880–892
Pimpinelli S, Berloco M, Fanti L, Dimitri P, Bonaccorsi S, Marchetti E, Caizzi R, Caggese C, Gatti M (1995) Transposable elements are stable structural components of Drosophila melanogaster heterochromatin. Proc Natl Acad Sci U S A 92:3804–3808
Quesneville H, Bergman CM, Andrieu O, Autard D, Nouaud D, Ashburner M, Anxolabehere D (2005) Combined evidence annotation of transposable elements in genome sequences. PLoS Comput Biol 1:166–175
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Rizzon C, Marais G, Gouy M, Biemont C (2002) Recombination rate and the distribution of transposable elements in the Drosophila melanogaster genome. Genome Res 12:400–407
Ruiz A, Wasserman M (1993) Evolutionary cytogenetics of the Drosophila buzzatii species complex. Heredity 70(Pt 6):582–596
Sambrook J, Fritsch E, Maniatis T (1989) Molecular cloning, a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
SanMiguel P, Tikhonov A, Jin YK, Motchoulskaia N, Zakharov D, Melake-Berhan A, Springer PS, Edwards KJ, Lee M, Avramova Z, Bennetzen JL (1996) Nested retrotransposons in the intergenic regions of the maize genome. Science 274:765–768
SanMiguel P, Gaut BS, Tikhonov A, Nakajima Y, Bennetzen JL (1998) The paleontology of intergene retrotransposons of maize. Nat Genet 20:43–45
Schwartz A, Chan DC, Brown LG, Alagappan R, Pettay D, Disteche C, McGillivray B, de la Chapelle A, Page DC (1998) Reconstructing hominid Y evolution: X-homologous block, created by X-Y transposition, was disrupted by Yp inversion through LINE–LINE recombination. Hum Mol Genet 7:1–11
Steinemann M, Steinemann S (1991) Preferential Y chromosomal location of TRIM, a novel transposable element of Drosophila miranda, obscura group. Chromosoma 101:169–179
Steinemann M, Steinemann S (1997) The enigma of Y chromosome degeneration: TRAM, a novel retrotransposon is preferentially located on the Neo-Y chromosome of Drosophila miranda. Genetics 145:261–266
Wharton L (1942) Analysis of the repleta group of Drosophila. Tex Univ Publ 4228:23–52
Wilder J, Hollocher H (2001) Mobile elements and the genesis of microsatellites in dipterans. Mol Biol Evol 18:384–392
Zelentsova H, Poluectova H, Mnjoian L, Lyozin G, Veleikodvorskaja V, Zhivotovsky L, Kidwell MG, Evgen’ev MB (1999) Distribution and evolution of mobile elements in the virilis species group of Drosophila. Chromosoma 108:443–456
Acknowledgement
This work was supported by grant BMC2002-01708 from the Dirección General de Enseñanza Superior e Investigación Científica (Ministerio de Educación y Cultura, Spain), which was awarded to A.R.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. Lehner
Sequence data from this article have been deposited in the EMBL/GenBank Data Libraries under accession number DQ402469.
Electronic supplementary materials
Below is the link to the electronic supplementary material.
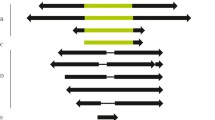
Fig. S1
Southern blot hybridization of (a) BuT1, (b) BuT2, (c) BuT3, (d) BuT4, (e) BuT5 and (f) BuT6 probes to genomic DNA of the following D. buzzatii lines (from left to right): st-1, st-3, st-7, st-10, j-2, j-9, j-19, j-23, j-24, jq7-1, jq7-4, jz3-6, jz3-7, y3-1 and s-1 (JPEG 39 kb)
Rights and permissions
About this article
Cite this article
Casals, F., González, J. & Ruiz, A. Abundance and chromosomal distribution of six Drosophila buzzatii transposons: BuT1, BuT2, BuT3, BuT4, BuT5, and BuT6 . Chromosoma 115, 403–412 (2006). https://doi.org/10.1007/s00412-006-0071-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-006-0071-7