Abstract
Background
Spirometric measurements of pulmonary function are important in diagnosing and determining the severity of chronic obstructive pulmonary disease (COPD). We performed this study to determine whether candidate genes identified in genome-wide association studies of spirometric measurements were associated with COPD and if they interacted with smoking intensity.
Methods
The current analysis included 1,000 COPD subjects and 1,000 controls recruited from 24 hospital-based pulmonary clinics. Thirteen SNPs, chosen based on genome-wide association studies of spirometric measurements in the Korean population cohorts, were genotyped. Genetic association tests were performed, adjusting for age, sex, and smoking intensity, using models including a SNP-by-smoking interaction term.
Results
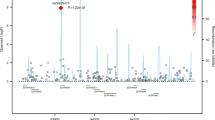
PID1 and FAM13A were significantly associated with COPD susceptibility. There were also significant interactions between SNPs in ACN9 and FAM13A and smoking pack-years, and an association of ACN9 with COPD in the lowest smoking tertile. The risk allele of FAM13A was associated with increased expression of FAM13A in the lung.
Conclusions
We have validated associations of FAM13A and PID1 with COPD. ACN9 showed significant interaction with smoking and is a potential candidate gene for COPD. Significant associations of genetic variants of FAM13A with gene expression levels suggest that the associated loci may act as genetic regulatory elements for FAM13A gene expression.

Similar content being viewed by others
References
Mathers CD, Loncar D (2006) Projections of global mortality and burden of disease from 2002 to 2030. PLoS Med 3:e442
Vestbo J, Hurd SS, Agustí AG, Jones PW, Vogelmeier C, Anzueto A et al (2013) Global strategy for the diagnosis, management, and prevention of chronic obstructive pulmonary disease: GOLD executive summary. Am J Respir Crit Care Med 187:347–365
Todd JL, Goldstein DB, Ge D, Christie J, Palmer SM (2011) The state of genome-wide association studies in pulmonary disease. Am J Respir Crit Care Med 184:873–880
Soler Artigas M, Wain LV, Repapi E, Me Obeidat, Sayers I, Burton PR et al (2012) Effect of five genetic variants associated with lung function on the risk of chronic obstructive lung disease, and their joint effects on lung function. Am J Respir Crit Care Med 184:786–795
Wilk JB, Shrine NRG, Loehr LR, Zhao JH, Manichaikul A, Lopez LM et al (2012) Genome-wide association studies identify CHRNA5/3 and HTR4 in the development of airflow obstruction. Am J Respir Crit Care Med 186:622–632
Castaldi PJ, Cho MH, Litonjua AA, Bakke P, Gulsvik A, Lomas DA et al (2011) The association of genome-wide significant spirometric loci with COPD susceptibility. Am J Respir Cell Mol Biol 45:1147–1153
Hardin M, Zielinski J, Wan ES, Hersh CP, Castaldi PJ, Schwinder E et al (2012) CHRNA3/5, IREB2, and ADCY2 are associated with severe chronic obstructive pulmonary disease in Poland. Am J Respir Cell Mol Biol 47:203–208
Hancock DB, Artigas MS, Gharib SA, Henry A, Manichaikul A, Ramasamy A et al (2012) Genome-wide joint meta-analysis of SNP and SNP-by-smoking interaction identifies novel loci for pulmonary function. PLoS Genet 8:e1003098
Miller MR, Hankinson J, Brusasco V, Burgos F, Casaburi R, Coates A et al (2005) Standardisation of spirometry. Eur Respir J 26:319–338
Kim WJ, Lee MK, Shin C, Cho NH, Lee SD, Oh YM, Sung J (2014) Genome-wide association studies identify locus on 6p21 influencing lung function in the Korean population. Respirology 19(3):360–368
Kim WJ, Oh YM, Lim JH, Jeong JG, Kim JH, Lee JS et al (2012) Gene expression profiling of lung tissue from COPD subjects [abstract]. Respirology 17(S2):35
Hancock DB, Eijgelsheim M, Wilk JB, Gharib SA, Loehr LR, Marciante KD et al (2010) Meta-analyses of genome-wide association studies identify multiple loci associated with pulmonary function. Nat Genet 42:45–52
Li X, Howard TD, Moore WC, Ampleford EJ, Li H, Busse WW et al (2011) Importance of hedgehog interacting protein and other lung function genes in asthma. J Allergy Clin Immunol 127:1457–1465
Xu H, Xu G, Wang D, Zheng C, Wan L (2013) Molecular cloning and tissue distribution of the phosphotyrosine interaction domain containing 1 (PID1) gene in Tianfu goat. Gene 515:71–77
Zhao Y, Zhang C, Chen X, Gao C, Ji C, Chen F et al (2012) Overexpression of NYGGF4 (PID1) induces mitochondrial impairment in 3T3-L1 adipocytes. Mol Cell Biochem 340:41–48
Wu WL, Gan WH, Tong ML, Li XL, Dai JZ, Zhang CM et al (2011) Over-expression of NYGGF4 (PID1) inhibits glucose transport in skeletal myotubes by blocking the IRS1/PI3K/AKT insulin pathway. Mol Genet Metab 102:374–377
Kirkham PA, Barnes PJ (2013) Oxidative stress in COPD. Chest 144:266–273
Cho MH, Boutaoui N, Klanderman BJ, Sylvia JS, Ziniti JP, Hersh CP et al (2010) Variants in FAM13A are associated with chronic obstructive pulmonary disease. Nat Genet 42:200–202
Fingerlin TE, Murphy E, Zhang W, Peljto AL, Brown KK, Steele MP et al (2013) Genome-wide association study identifies multiple susceptibility loci for pulmonary fibrosis. Nat Genet 45:613–620
Cohen M, Reichenstein M, Everts-van der Wind A, Heon-Lee J, Shani M, Lewin HA et al (2004) Cloning and characterization of FAM13A1—a gene near a milk protein QTL on BTA6: evidence for population-wide linkage disequilibrium in Israeli Holsteins. Genomics 84:374–383
Dennis RA, McCammon MT (1999) Acn9 is a novel protein of gluconeogenesis that is located in the mitochondrial intermembrane space. Eur J Biochem 261:236–243
de Graaf RM, Ricard G, van Alen TA, Duarte I, Dutilh BE, Burgtorf C et al (2011) The Organellar genome and metabolic potential of the hydrogen-producing mitochondrion of Nyctotherus ovalis. Mol Biol Evol 28:2379–2391
Dick DM, Aliev F, Wang JC, Saccone S, Hinrichs A, Bertelsen S et al (2008) A systematic single nucleotide polymorphism screen to fine-map alcohol dependence genes on chromosome 7 identifies association with a novel susceptibility gene ACN9. Biol Psychiatry 63:1047–1053
Acknowledgments
This study was supported by the National Project for Personalized Genomic Medicine (A111218-11-GM02) and a grant from the Korean Health 21 R&D Project, Ministry of Health and Welfare, Republic of Korea (A102065). DNA samples were generously provided by the Kangwon National University Hospital Biobank, which is part of the National Biobank of Korea, supported by the Ministry of Health and Welfare, Republic of Korea. For the recruitment of COPD participants and also for the collection of data and samples, we thank all members of the Korean Obstructive Lung Disease (KOLD) Study Group anrsd the ASAN Network: Ji-Hyun Lee, Eun Kyung Kim, Tae-Hyung Kim, Tae Rim Shin, Jin Hwa Lee, Seong Yong Lim, Sang Yeub Lee, Ho Il Yoon, Kwang Ha Yoo, Seung Soo Sheen, Joo Hun Park, Yong Bum Park, Changhwan Kim, Yong Il Hwang, Young Sam Kim, Ji Ye Jung, Yoonki Hong, Seung Won Ra, Joon Beom Seo, Sang Min Lee, In A Jeong, Chang Hoon Lee, Sei Won Lee, Jae Seung Lee, Jin Won Huh, Ji Yong Moon, HyeKyeong Park, Hye Yun Park, Jin Woo Kim, Chin Kook Rhee, Hyoung Kyu Yoon, Woo Jin Kim, Jong Deog Lee, Kang Hyeon Choi, Bock Hyun Jung, Joo Ock Na, Doh Hyung Kim, Hye Sook Choi, Kwang Ha Lee, Myung Jae Park, Sung Soon Lee, Yeon-Mok Oh, and Sang Do Lee.
Conflict of interest
SDL received payment for lecturing from Nycomed, Takeda, and Norvatis. YMO received payment for lecturing from MSD Korea, AstraZeneca Korea, Boehringer Ingelheim Korea, Norvatis, DongWha, Takeda, and GSK Korea. EKS received a consultant fee from GSK, AstraZeneca, and Merck and payment for lecturing from GSK, AstraZeneca, and Merck. The other authors have no conflicts of interest to disclose.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kim, W.J., Lim, M.N., Hong, Y. et al. Association of Lung Function Genes with Chronic Obstructive Pulmonary Disease. Lung 192, 473–480 (2014). https://doi.org/10.1007/s00408-014-9579-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00408-014-9579-4