Abstract
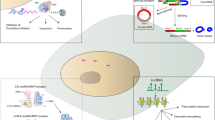
MicroRNAs (miRs) are small non-coding RNAs, which control gene expression either by inducing mRNA degradation or by blocking translation, and play a crucial role in tissue homeostasis. In the cardiovascular system, miRs were shown to control cardiac hypertrophy, fibrosis and apoptosis, angiogenesis, and vessel remodeling. In addition, miRs regulate stem cell maintenance and some miRs induced cell fate decisions. This review summarizes the current insights into the control of stem cells and lineage commitment by miRs focusing specifically to the regulation of endothelial, smooth muscle, and cardiac lineage.

Similar content being viewed by others
References
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297. doi:S0092867404000455
Small EM, Frost RJ, Olson EN (2010) MicroRNAs add a new dimension to cardiovascular disease. Circulation 121:1022–1032. doi:10.1161/CIRCULATIONAHA.109.889048
Bonauer A, Boon RA, Dimmeler S (2010) Vascular microRNAs. Curr Drug Targets 11:943–949. doi:BSP/CDT/E-Pub/00088
Thum T, Catalucci D, Bauersachs J (2008) MicroRNAs: novel regulators in cardiac development and disease. Cardiovasc Res 79:562–570. doi:10.1093/cvr/cvn137
Martinez NJ, Gregory RI (2010) MicroRNA gene regulatory pathways in the establishment and maintenance of ESC identity. Cell Stem Cell 7:31–35. doi:10.1016/j.stem.2010.06.011
Sinkkonen L, Hugenschmidt T, Berninger P, Gaidatzis D, Mohn F, Artus-Revel CG, Zavolan M, Svoboda P, Filipowicz W (2008) MicroRNAs control de novo DNA methylation through regulation of transcriptional repressors in mouse embryonic stem cells. Nat Struct Mol Biol 15:259–267. doi:10.1038/nsmb.1391
Wang Y, Baskerville S, Shenoy A, Babiarz JE, Baehner L, Blelloch R (2008) Embryonic stem cell-specific microRNAs regulate the G1-S transition and promote rapid proliferation. Nat Genet 40:1478–1483. doi:10.1038/ng.250
Tay Y, Zhang J, Thomson AM, Lim B, Rigoutsos I (2008) MicroRNAs to Nanog, Oct4 and Sox2 coding regions modulate embryonic stem cell differentiation. Nature 455:1124–1128. doi:10.1038/nature07299
Wellner U, Schubert J, Burk UC, Schmalhofer O, Zhu F, Sonntag A, Waldvogel B, Vannier C, Darling D, zur Hausen A, Brunton VG, Morton J, Sansom O, Schuler J, Stemmler MP, Herzberger C, Hopt U, Keck T, Brabletz S, Brabletz T (2009) The EMT-activator ZEB1 promotes tumorigenicity by repressing stemness-inhibiting microRNAs. Nat Cell Biol 11:1487–1495. doi:10.1038/ncb1998
Xu N, Papagiannakopoulos T, Pan G, Thomson JA, Kosik KS (2009) MicroRNA-145 regulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell 137:647–658. doi:10.1016/j.cell.2009.02.038
Melton C, Judson RL, Blelloch R (2010) Opposing microRNA families regulate self-renewal in mouse embryonic stem cells. Nature 463:621–626. doi:10.1038/nature08725
Yang WJ, Yang DD, Na S, Sandusky GE, Zhang Q, Zhao G (2005) Dicer is required for embryonic angiogenesis during mouse development. J Biol Chem 280:9330–9335. doi:10.1074/jbc.M413394200
Kuehbacher A, Urbich C, Zeiher AM, Dimmeler S (2007) Role of Dicer and Drosha for endothelial microRNA expression and angiogenesis. Circ Res 101:59–68. doi:10.1161/CIRCRESAHA.107.153916
Suarez Y, Fernandez-Hernando C, Pober JS, Sessa WC (2007) Dicer dependent microRNAs regulate gene expression and functions in human endothelial cells. Circ Res 100:1164–1173. doi:10.1161/01.RES.0000265065.26744.17
Giraldez AJ, Cinalli RM, Glasner ME, Enright AJ, Thomson JM, Baskerville S, Hammond SM, Bartel DP, Schier AF (2005) MicroRNAs regulate brain morphogenesis in zebrafish. Science 308:833–838. doi:10.1126/science.1109020
Suarez Y, Fernandez-Hernando C, Yu J, Gerber SA, Harrison KD, Pober JS, Iruela-Arispe ML, Merkenschlager M, Sessa WC (2008) Dicer-dependent endothelial microRNAs are necessary for postnatal angiogenesis. Proc Natl Acad Sci USA 105:14082–14087. doi:10.1073/pnas.0804597105
Ivey KN, Muth A, Arnold J, King FW, Yeh RF, Fish JE, Hsiao EC, Schwartz RJ, Conklin BR, Bernstein HS, Srivastava D (2008) MicroRNA regulation of cell lineages in mouse and human embryonic stem cells. Cell Stem Cell 2:219–229. doi:10.1016/j.stem.2008.01.016
Fish JE, Santoro MM, Morton SU, Yu S, Yeh RF, Wythe JD, Ivey KN, Bruneau BG, Stainier DY, Srivastava D (2008) MiR-126 regulates angiogenic signaling and vascular integrity. Dev Cell 15:272–284. doi:10.1016/j.devcel.2008.07.008
Wang S, Aurora AB, Johnson BA, Qi X, McAnally J, Hill JA, Richardson JA, Bassel-Duby R, Olson EN (2008) The endothelial-specific microRNA miR-126 governs vascular integrity and angiogenesis. Dev Cell 15:261–271. doi:10.1016/j.devcel.2008.07.002
Taganov KD, Boldin MP, Chang KJ, Baltimore D (2006) NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc Natl Acad Sci USA 103:12481–12486. doi:10.1073/pnas.0605298103
Kane NM, Meloni M, Spencer HL, Craig MA, Strehl R, Milligan G, Houslay MD, Mountford JC, Emanueli C, Baker AH (2010) Derivation of endothelial cells from human embryonic stem cells by directed differentiation: analysis of microRNA and angiogenesis in vitro and in vivo. Arterioscler Thromb Vasc Biol 30:1389–1397. doi:10.1161/ATVBAHA.110.204800
Fasanaro P, Greco S, Lorenzi M, Pescatori M, Brioschi M, Kulshreshtha R, Banfi C, Stubbs A, Calin GA, Ivan M, Capogrossi MC, Martelli F (2009) An integrated approach for experimental target identification of hypoxia-induced miR-210. J Biol Chem 284:35134–35143. doi:10.1074/jbc.M109.052779
Hornstein E, Mansfield JH, Yekta S, Hu JK, Harfe BD, McManus MT, Baskerville S, Bartel DP, Tabin CJ (2005) The microRNA miR-196 acts upstream of Hoxb8 and Shh in limb development. Nature 438:671–674. doi:10.1038/nature04138
Doebele C, Bonauer A, Fischer A, Scholz A, Reiss Y, Urbich C, Hofmann WK, Zeiher AM, Dimmeler S (2010) Members of the microRNA-17–92 cluster exhibit a cell-intrinsic antiangiogenic function in endothelial cells. Blood 115:4944–4950. doi:10.1182/blood-2010-01-264812
Bonauer A, Carmona G, Iwasaki M, Mione M, Koyanagi M, Fischer A, Burchfield J, Fox H, Doebele C, Ohtani K, Chavakis E, Potente M, Tjwa M, Urbich C, Zeiher AM, Dimmeler S (2009) MicroRNA-92a controls angiogenesis and functional recovery of ischemic tissues in mice. Science 324:1710–1713. doi:10.1126/science.1174381
Kazenwadel J, Michael MZ, Harvey NL (2010) Prox1 expression is negatively regulated by miR-181 in endothelial cells. Blood. doi:10.1182/blood-2009-12-256297
Boettger T, Beetz N, Kostin S, Schneider J, Kruger M, Hein L, Braun T (2009) Acquisition of the contractile phenotype by murine arterial smooth muscle cells depends on the MiR143/145 gene cluster. J Clin Invest 119:2634–2647. doi:10.1172/JCI38864
Cordes KR, Sheehy NT, White MP, Berry EC, Morton SU, Muth AN, Lee TH, Miano JM, Ivey KN, Srivastava D (2009) MiR-145 and miR-143 regulate smooth muscle cell fate and plasticity. Nature 460:705–710. doi:10.1038/nature08195
Xin M, Small EM, Sutherland LB, Qi X, McAnally J, Plato CF, Richardson JA, Bassel-Duby R, Olson EN (2009) MicroRNAs miR-143 and miR-145 modulate cytoskeletal dynamics and responsiveness of smooth muscle cells to injury. Genes Dev 23:2166–2178. doi:10.1101/gad.1842409
Elia L, Quintavalle M, Zhang J, Contu R, Cossu L, Latronico MV, Peterson KL, Indolfi C, Catalucci D, Chen J, Courtneidge SA, Condorelli G (2009) The knockout of miR-143 and -145 alters smooth muscle cell maintenance and vascular homeostasis in mice: correlates with human disease. Cell Death Differ 16:1590–1598. doi:10.1038/cdd.2009.153
Xie C, Huang H, Sun X, Guo Y, Hamblin M, Ritchie RP, Garcia-Barrio MT, Zhang J, Chen YE (2010) MicroRNA-1 regulates smooth muscle cell differentiation by repressing KLF4. Stem Cells Dev. doi:10.1089/scd.2010.0283
Chen JF, Mandel EM, Thomson JM, Wu Q, Callis TE, Hammond SM, Conlon FL, Wang DZ (2006) The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nat Genet 38:228–233. doi:10.1038/ng1725
Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, Bartel DP, Linsley PS, Johnson JM (2005) Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433:769–773. doi:10.1038/nature03315
Zhao Y, Samal E, Srivastava D (2005) Serum response factor regulates a muscle-specific microRNA that targets Hand2 during cardiogenesis. Nature 436:214–220. doi:10.1038/nature03817
Zhao Y, Ransom JF, Li A, Vedantham V, von Drehle M, Muth AN, Tsuchihashi T, McManus MT, Schwartz RJ, Srivastava D (2007) Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2. Cell 129:303–317. doi:10.1016/j.cell.2007.03.030
Kwon C, Han Z, Olson EN, Srivastava D (2005) MicroRNA1 influences cardiac differentiation in Drosophila and regulates Notch signaling. Proc Natl Acad Sci USA 102:18986–18991. doi:10.1073/pnas.0509535102
Sluijter JP, van Mil A, van Vliet P, Metz CH, Liu J, Doevendans PA, Goumans MJ (2010) MicroRNA-1 and -499 regulate differentiation and proliferation in human-derived cardiomyocyte progenitor cells. Arterioscler Thromb Vasc Biol 30:859–868. doi:10.1161/ATVBAHA.109.197434
Wilson KD, Hu S, Venkatasubrahmanyam S, Fu JD, Abilez OJ, Baugh JJ, Jia F, Ghosh Z, Li RA, Butte AJ, Wu JC (2010) Dynamic microrna expression programs during cardiac differentiation of human embryonic stem cells: role for miR-499. Circ Cardiovasc Genet. doi:10.1161/CIRCGENETICS.109.934281
Takaya T, Ono K, Kawamura T, Takanabe R, Kaichi S, Morimoto T, Wada H, Kita T, Shimatsu A, Hasegawa K (2009) MicroRNA-1 and MicroRNA-133 in spontaneous myocardial differentiation of mouse embryonic stem cells. Circ J 73:1492–1497. doi:JST.JSTAGE/circj/CJ-08-1032
Liu N, Bezprozvannaya S, Williams AH, Qi X, Richardson JA, Bassel-Duby R, Olson EN (2008) MicroRNA-133a regulates cardiomyocyte proliferation and suppresses smooth muscle gene expression in the heart. Genes Dev 22:3242–3254. doi:10.1101/gad.1738708
Condorelli G, Latronico MV, Dorn GW 2nd (2010) MicroRNAs in heart disease: putative novel therapeutic targets? Eur Heart J 31:649–658. doi:10.1093/eurheartj/ehp573
van Rooij E, Sutherland LB, Qi X, Richardson JA, Hill J, Olson EN (2007) Control of stress-dependent cardiac growth and gene expression by a microRNA. Science 316:575–579. doi:10.1126/science.1139089
Ventura A, Young AG, Winslow MM, Lintault L, Meissner A, Erkeland SJ, Newman J, Bronson RT, Crowley D, Stone JR, Jaenisch R, Sharp PA, Jacks T (2008) Targeted deletion reveals essential and overlapping functions of the miR-17 through 92 family of miRNA clusters. Cell 132:875–886. doi:10.1016/j.cell.2008.02.019
Bonauer A, Dimmeler S (2009) The microRNA-17 92 cluster: still a miRacle? Cell Cycle 8:3866–3873. doi:9994
Fleissner F, Jazbutyte V, Fiedler J, Gupta SK, Yin X, Xu Q, Galuppo P, Kneitz S, Mayr M, Ertl G, Bauersachs J, Thum T (2010) Short communication: asymmetric dimethylarginine impairs angiogenic progenitor cell function in patients with coronary artery disease through a microRNA-21-dependent mechanism. Circ Res 107:138–143. doi:10.1161/circresaha.110.216770
Dimmeler S, Leri A (2008) Aging and disease as modifiers of efficacy of cell therapy. Circ Res 102:1319–1330. doi:10.1161/CIRCRESAHA.108.175943
Seeger FH, Sedding D, Langheinrich AC, Haendeler J, Zeiher AM, Dimmeler S (2010) Inhibition of the p38 MAP kinase in vivo improves number and functional activity of vasculogenic cells and reduces atherosclerotic disease progression. Basic Res Cardiol 105:389–397. doi:10.1007/s00395-009-0072-9
Van Craenenbroeck EM, Hoymans VY, Beckers PJ, Possemiers NM, Wuyts K, Paelinck BP, Vrints CJ, Conraads VM (2010) Exercise training improves function of circulating angiogenic cells in patients with chronic heart failure. Basic Res Cardiol 105:665–676. doi:10.1007/s00395-010-0105-4
Acknowledgments
The authors are supported by the Deutsche Forschungsgemeinschaft (SFB834, Exc 147/1) and the European Research Council (Advanced grant “Angiomir”).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ohtani, K., Dimmeler, S. Control of cardiovascular differentiation by microRNAs. Basic Res Cardiol 106, 5–11 (2011). https://doi.org/10.1007/s00395-010-0139-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00395-010-0139-7