Abstract
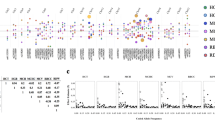
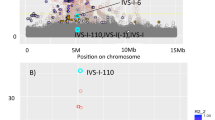
Genetic variations in blood cell parameters can impact clinical traits. We report here the mapping of blood cell traits in a panel of 100 inbred strains of mice of the Hybrid Mouse Diversity Panel (HMDP) using genome-wide association (GWA). We replicated a locus previously identified in using linkage analysis in several genetic crosses for mean corpuscular volume (MCV) and a number of other red blood cell traits on distal chromosome 7. Our peak for SNP association to MCV occurred in a linkage disequilibrium (LD) block spanning from 109.38 to 111.75 Mb that includes Hbb-b1, the likely causal gene. Altogether, we identified five loci controlling red blood cell traits (on chromosomes 1, 7, 11, 12, and 16), and four of these correspond to loci for red blood cell traits reported in a recent human GWA study. For white blood cells, including granulocytes, monocytes, and lymphocytes, a total of six significant loci were identified on chromosomes 1, 6, 8, 11, 12, and 15. An average of ten candidate genes were found at each locus and those were prioritized by examining functional variants in the HMDP such as missense and expression variants. These results provide intermediate phenotypes and candidate loci for genetic studies of atherosclerosis and cancer as well as inflammatory and immune disorders in mice.







Similar content being viewed by others
References
Bennett BJ, Farber CR, Orozco L et al (2010) A high-resolution association mapping panel for the dissection of complex traits in mice. Genome Res 20:281–290
Broman KW, Wu H, Sen S, Churchill GA (2003) R/qtl: QTL mapping in experimental crosses. Bioinformatics 19:889–890
Davis RC, van Nas A, Castellani LW et al (2012) Systems genetics of susceptibility to obesity-induced diabetes in mice. Physiol Genomics 44:1–13
Evans DM, Frazer IH, Martin NG (1999) Genetic and environmental causes of variation in basal levels of blood cells. Twin Res 2:250–257
Farber CR, Bennett BJ, Orozco L et al (2011) Mouse genome-wide association and systems genetics identify Asxl2 as a regulator of bone mineral density and osteoclastogenesis. PLoS Genet 7:e1002038
Flint J, Eskin E (2012) Genome-wide association studies in mice. Nat Rev Genet 13:807–817
Flint J, Valdar W, Shifman S, Mott R (2005) Strategies for mapping and cloning quantitative trait genes in rodents. Nat Rev Genet 6:271–286
Ganesh SK, Zakai NA, van Rooij FJ et al (2009) Multiple loci influence erythrocyte phenotypes in the CHARGE Consortium. Nat Genet 41:1191–1198
Garner C, Tatu T, Reittie JE et al (2000) Genetic influences on F cells and other hematologic variables: a twin heritability study. Blood 95:342–346
Gillum RF, Mussolino ME, Madans JH (2005) Counts of neutrophils, lymphocytes, and monocytes, cause-specific mortality and coronary heart disease: the NHANES-I epidemiologic follow-up study. Ann Epidemiol 15:266–271
Hedrick CC, Castellani LW, Warden CH, Puppione DL, Lusis AJ (1993) Influence of mouse apolipoprotein A-II on plasma lipoproteins in transgenic mice. J Biol Chem 268:20676–20682
Horvat S, Bunger L (1999) Polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) assay for the mouse leptin receptor (Lepr(db)) mutation. Lab Anim 33:380–384
Kang HM, Zaitlen NA, Wade CM et al (2008) Efficient control of population structure in model organism association mapping. Genetics 178:1709–1723
Kelada SN, Aylor DL, Peck BC et al (2012) Genetic analysis of hematological parameters in incipient lines of the collaborative cross. G3 (Bethesda) 2:157–165
Laurie CC, Nickerson DA, Anderson AD et al (2007) Linkage disequilibrium in wild mice. PLoS Genet 3:e144
Lloyd-Jones DM, Camargo CA, Allen LA, Giugliano RP, O’Donnell CJ (2003) Predictors of long-term mortality after hospitalization for primary unstable angina pectoris and non-ST-elevation myocardial infarction. Am J Cardiol 92:1155–1159
Nalls MA, Couper DJ, Tanaka T et al (2011) Multiple loci are associated with white blood cell phenotypes. PLoS Genet 7:e1002113
Orozco LD, Bennett BJ, Farber CR et al (2012) Unraveling inflammatory responses using systems genetics and gene-environment interactions in macrophages. Cell 151:658–670
Payseur BA, Place M, Weber JL (2008) Linkage disequilibrium between STRPs and SNPs across the human genome. Am J Hum Genet 82:1039–1050
Peters LL, Shavit JA, Lambert AJ et al (2010) Sequence variation at multiple loci influences red cell hemoglobin concentration. Blood 116:e139–e149
Puppione DL, Charugundla S (1994) A microprecipitation technique suitable for measuring alpha-lipoprotein cholesterol. Lipids 29:595–597
Reiner AP, Lettre G, Nalls MA et al (2011) Genome-wide association study of white blood cell count in 16,388 African Americans: the continental origins and genetic epidemiology network (COGENT). PLoS Genet 7:e1002108
Shankar A, Wang JJ, Rochtchina E, Yu MC, Kefford R, Mitchell P (2006) Association between circulating white blood cell count and cancer mortality: a population-based cohort study. Arch Intern Med 166:188–194
Soranzo N, Spector TD, Mangino M et al (2009) A genome-wide meta-analysis identifies 22 loci associated with eight hematological parameters in the HaemGen consortium. Nat Genet 41:1182–1190
van der Harst P (2013) Seventy-five genetic loci influencing the human red blood cell. Nature 492:369–375
van Nas A, Ingram-Drake L, Sinsheimer JS et al (2010) Expression quantitative trait loci: replication, tissue- and sex-specificity in mice. Genetics 185:1059–1068
Acknowledgments
This work was supported by NIH grants HL030568, HL028841, DK094311.
Disclosures
The authors have no conflicts of interest to disclose.
Author information
Authors and Affiliations
Corresponding author
Additional information
van Nas and R. C. Davis contributed equally to this study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Davis, R.C., van Nas, A., Bennett, B. et al. Genome-wide association mapping of blood cell traits in mice. Mamm Genome 24, 105–118 (2013). https://doi.org/10.1007/s00335-013-9448-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00335-013-9448-0