Abstract
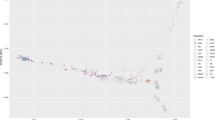
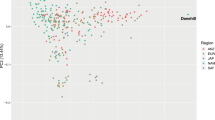
Alaskan sled dogs are a genetically distinct population shaped by generations of selective interbreeding with purebred dogs to create a group of high-performance athletes. As a result of selective breeding strategies, sled dogs present a unique opportunity to employ admixture-mapping techniques to investigate how breed composition and trait selection impact genomic structure. We used admixture mapping to investigate genetic ancestry across the genomes of two classes of sled dogs, sprint and long-distance racers, and combined that with genome-wide association studies (GWAS) to identify regions that correlate with performance-enhancing traits. The sled dog genome is enhanced by differential contributions from four non-admixed breeds (Alaskan Malamute, Siberian Husky, German Shorthaired Pointer, and Borzoi). A principal components analysis (PCA) of 115,000 genome-wide SNPs clearly resolved the sprint and distance populations as distinct genetic groups, with longer blocks of linkage disequilibrium (LD) observed in the distance versus sprint dogs (7.5–10 and 2.5–3.75 kb, respectively). Furthermore, we identified eight regions with the genomic signal from either a selective sweep or an association analysis, corroborated by an excess of ancestry when comparing sprint and distance dogs. A comparison of elite and poor-performing sled dogs identified a single region significantly associated with heat tolerance. Within the region we identified seven SNPs within the myosin heavy chain 9 gene (MYH9) that were significantly associated with heat tolerance in sprint dogs, two of which correspond to conserved promoter and enhancer regions in the human ortholog.







Similar content being viewed by others
References
ADMA (2011) Alaska Dog Musher’s Association, Fairbanks, AK. Available at http://www.sleddog.org
Barbier E, Wang JB (2009) Anti-depressant and anxiolytic like behaviors in PKCI/HINT1 knockout mice associated with elevated plasma corticosterone level. BMC Neurosci 10:132
Barrett JC, Fry B, Maller J, Daly MJ (2005) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21:263–265
Bhangale TR, Stephens M, Nickerson DA (2006) Automating resequencing-based detection of insertion-deletion polymorphisms. Nat Genet 38:1457–1462
Boyko AR, Quignon P, Li L, Schoenebeck JJ, Degenhardt JD, Lohmueller KE, Zhao K, Brisbin A, Parker HG, von Holdt BM, Cargill M, Auton A, Reynolds A, Elkahloun AG, Castelhano M, Mosher DS, Sutter NB, Johnson GS, Novembre J, Hubisz MJ, Siepel A, Wayne RK, Bustamante CD, Ostrander EA (2010) A simple genetic architecture underlies morphological variation in dogs. PLoS Biol 8:e1000451
Buerkle CA, Lexer C (2008) Admixture as the basis for genetic mapping. Cell 23:686–694
Burniston JG (2009) Adaptation of the rat cardiac proteome in response to intensity-controlled endurance exercise. Proteomics 9:106–115
Cheng CY, Reich D, Wong TY, Klein R, Klein BE, Patterson N, Tandon A, Li M, Boerwinkle E, Sharrett AR, Kao WH (2010) Admixture mapping scans identify a locus affecting retinal vascular caliber in hypertensive African Americans: the Atherosclerosis Risk In Communities (ARIC) study. PLoS Genet 6:e1000908
Collins MJC (1991) Dog Driver: A Guide for the Serious Musher, 1st edn. Alpine Publications, Crawford
Ernst J, Kellis M (2010) Discovery and characterization of chromatin states for systematic annotation of the human genome. Nat Biotechnol 28:817–825
Ernst J, Kheradpour P, Mikkelsen TS, Shoresh N, Ward LD, Epstein CB, Zhang X, Wang L, Issner R, Coyne M, Ku M, Durham T, Kellis M, Bernstein BE (2011) Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473:43–49
Ewing B, Hillier L, Wendl MC, Green P (1998) Base-calling of automated sequencer traces using phred. I. Accuracy assessment. Genome Res 8:175–185
Gaut BS, Long AD (2003) The lowdown on linkage disequilibrium. Plant Cell 15:1502–1506
Gordon D, Abajian C, Green P (1998) Consed: a graphical tool for sequence finishing. Genome Res 8:195–202
Gray SR, De Vito G, Nimmo MA, Farina D, Ferguson RA (2006) Skeletal muscle ATP turnover and muscle fiber conduction velocity are elevated at higher muscle temperatures during maximal power output development in humans. Am J Physiol Regul Integr Comp Physiol 290:R376–R382
Gray MM, Granka JM, Bustamante CD, Sutter NB, Boyko AR, Zhu L, Ostrander EA, Wayne RK (2009) Linkage disequilibrium and demographic history of wild and domestic canids. Genet 181:1493–1505
Hakonarson H, Grant SF (2011) Planning a genome-wide association study: Points to consider. Ann Med 43:451–460
Huson HJ, Parker HG, Runstadler J, Ostrander EA (2010) A genetic dissection of breed composition and performance enhancement in the Alaskan sled dog. BMC Genet 11:71
Iditarod (2011) The Official Site of the Iditarod, Iditarod Trail Committee, Inc., Anchorage, AK. Available at http://www.iditarod.com/
Ingalls CP, Warren GL, Armstrong RB (1998) Dissociation of force production from MHC and actin contents in muscles injured by eccentric contractions. J Muscle Res Cell Motil 19:215–224
Kang HM (2010) Efficient Mixed_model Association eXpediated (EMMAX). Los Angeles, CA: University of California, Los Angeles
KC A (1998) The Complete Dog Book, Official Publication of The American Kennel Club, 19th edn. Howell Book House, New York
Kwak GH, Kim JR, Kim HY (2009) Expression, subcellular localization, and antioxidant role of mammalian methionine sulfoxide reductases in Saccharomyces cerevisiae. BMB Rep 42:113–118
Luciano M, Hansell NK, Lahti J, Davies G, Medland SE, Raikkonen K, Tenesa A, Widen E, McGhee KA, Palotie A, Liewald D, Porteous DJ, Starr JM, Montgomery GW, Martin NG, Eriksson JG, Wright MJ, Deary IJ (2011) Whole genome association scan for genetic polymorphisms influencing information processing speed. Biol Psychol 86:193–202
Nickerson DA, Tobe VO, Taylor SL (1997) PolyPhred: automating the detection and genotyping of single nucleotide substitutions using fluorescence-based resequencing. Nucleic Acids Res 15:2745–2751
Ostrander EA, Huson HJ, Ostrander GK (2009) Genetics of atheletic performance. Annu Rev Genomics Hum Genet 10:407–429
Parker HG, Shearin AL, Ostrander EA (2010) Man’s best friend becomes biology’s best in show: genome analyses in the domestic dog. Annu Rev Genet 44:309–336
Patterson N, Hattangadi N, Lane B, Lohmueller KE, Hafler DA et al (2004) Methods for high-density admixture mapping of disease genes. Am J Hum Genet 74:979–1000
Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D (2006) Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38:904–909
Pritchard JK, Przeworski M (2001) Linkage disequilibrium in humans: models and data. Am J Hum Genet 69:1–14
Purcell S (2009) PLINK_v1.07, Whole genome association analysis toolset. Available at http://pngu.mgh.harvard.edu/~purcell/plink/download.shtml
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ, Sham PC (2007) PLINK: A tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
Rennick P (ed) (1987) Dogs of the North. Alaska Geographic Society, Anchorage
Rosenberg NA, Huang L, Jewett EM, Szpiech ZA, Jankovic I, Boehnke M (2010) Genome-wide association studies in diverse populations. Nat Rev Genet 11:356–366
Scheet P, Stephens M (2006) A fast and flexible statistical model for large-scale population genotype data: applications to inferring missing genotypes and haplotypic phase. Am J Hum Genet 78:629–644
Seldin MF, Posaniuc B, Price AL (2011) New approaches to disease mapping in admixed populations. Nat Rev Genet 12:523–528
Sharer JD, Shern JF, Van Valkenburgh H, Wallace DC, Kahn RA (2002) ARL2 and BART enter mitochondria and bind the adenine nucleotide transporter. Mol Biol Cell 13:71–83
Shriner D (2011) Investigating population stratification and admixture using eigenanalysis of dense genotypes. Heredity 107(5):413–420
Stark Z, Bruno DL, Mountford H, Lockhart PJ, Amor DJ (2010) De novo 325 kb microdeletion in chromosome band 10q25.3 including ATRNL1 in a boy with cognitive impairment, autism and dysmorphic features. Eur J Med Genet 53:337–339
Suckale J, Solimena M (2010) The insulin secretory granule as a signaling hub. Trends Endocrinol Metab 21:599–609
Sutter NB, Eberle MA, Parker HG, Pullar BJ, Kirkness EF, Kruglyak L, Ostrander EA (2004) Extensive and breed-specific linkage disequilibrium in Canis familiaris. Genome Res 14:2388–2396
Tang H, Coram M, Wang P, Zhu X, Risch N (2006) Reconstructing genetic ancestry blocks in admixed individuals. Am J Hum Genet 79:1–12
Tang H, Choudhry S, Mei R, Morgan M, Rodriguez-Cintron W, Burchard EG, Risch NJ (2007) Recent genetic selection in the ancestral admixture of Puerto Ricans. Am J Hum Genet 81:626–633
Tian C, Hinds DA, Shigeta R, Kittles R, Ballinger DG, Seldin MF (2006) A genomewide single-nucleotide-polymorphism panel with high ancestry information for African American admixture mapping. Am J Hum Genet 79:640–649
UCSC (2011) University of California, Santa Cruz Genome Browser, human build March 26; canine version 2.0. Available at http://genome.ucsc.edu/
Varadarajulu J, Lebar M, Krishnamoorthy G, Habelt S, Lu J, Bernard Weinstein I, Li H, Holsboer F, Turck CW, Touma C (2011) Increased anxiety-related behaviour in Hint1 knockout mice. Behav Brain Res 220:305–311
Vaudrin B (ed) (1976) Racing Alaskan Sled Dogs, 1st edn. Alaska Northwest Publishing Company, Anchorage
VonHoldt BM, Pollinger JP, Lohmueller KE, Han E, Parker HG, Quignon P, Degenhardt JD, Boyko AR, Earl DA, Auton A, Reynolds A, Bryc K, Brisbin A, Knowles JC, Mosher DS, Spady TC, Elkahloun A, Geffen E, Pilot M, Jedrzejewski W, Greco C, Randi E, Bannasch D, Wilton A, Shearman J, Musiani M, Cargill M, Jones PG, Qian Z, Huang W, Ding ZL, Zhang YP, Bustamante CD, Ostrander EA, Novembre J, Wayne RK (2010) Genome-wide SNP and haplotype analyses reveal a rich history underlying dog domestication. Nature 464:898–902
Wendt R (1999) Alaska Dog Mushing Guide: Facts and Legends, 5th edn. Goldstream Publications, Fairbanks
Winkler CA, Nelson GW, Smith MW (2010) Admixture mapping comes of age. Annu Rev Genomics Hum Genet 11:65–89
Yukon Quest (2011) Yukon Quest Sled Dog Race. Available at http://www.yukonquest.com/
Acknowledgments
We gratefully acknowledge the Intramural Program of the National Human Genome Research Institute. We thank the many owners and participants who provided DNA samples and information regarding their dogs.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Huson, H.J., vonHoldt, B.M., Rimbault, M. et al. Breed-specific ancestry studies and genome-wide association analysis highlight an association between the MYH9 gene and heat tolerance in Alaskan sprint racing sled dogs. Mamm Genome 23, 178–194 (2012). https://doi.org/10.1007/s00335-011-9374-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00335-011-9374-y