Abstract
Formyl peptide receptor 3 (FPR3) is a potential player in innate immunity and appears with FPR2 as a FPR cluster during primate evolution. Comparative genome analyses indicate that a segmental duplication (SD) event upstream of the FPR3 gene after the divergence of New and Old World monkeys led to the emergence of an alternative promoter. In this study we combined computational and experimental approaches to identify a FPR3 gene that is controlled by an alternative promoter derived during a SD event. Its transcriptional activity was detected by quantitative reverse transcription polymerase chain reaction. Human alternative transcripts (FPR3-1 and FPR3-2) showed tissue-specific patterns with strong expressions in lung or uterus, while the FPR3-1 transcript of rhesus macaque is broadly expressed in various tissues. Overall, transcriptional variations of FPR3 occur by an alternative promoter during primate evolution.






Similar content being viewed by others
References
Ahn K, Huh JW, Park SJ, Kim DS, Ha HS et al (2008) Selection of internal reference genes for SYBR green qRT-PCR studies of rhesus monkey (Macaca mulatta) tissues. BMC Mol Biol 9:78
Akam M (1998) Hox genes, homeosis and the evolution of segment identity: no need for hopeless monsters. Int J Dev Biol 42:445–451
Alvarez V, Coto E, Setién F, González-Roces S, López-Larrea C (1996) Molecular evolution of the N-formyl peptide and C5a receptors in non-human primates. Immunogenetics 44:446–452
Bailey JA, Eichler EE (2006) Primate segmental duplications: crucibles of evolution, diversity and disease. Nat Rev Genet 7:552–564
Bailey JA, Gu Z, Clark RA, Reinert K, Samonte RV et al (2002) Recent segmental duplications in the human genome. Science 297:1003–1007
Bao L, Gerard NP, Eddy RL Jr, Shows TB, Gerard C (1992) Mapping of genes for the human C5a receptor (C5AR), human FMLP receptor (FPR), and two FMLP receptor homologue orphan receptors (FPRH1, FPRH2) to chromosome 19. Genomics 13:437–440
Blekhman R, Oshlack A, Gilad Y (2009) Segmental duplications contribute to gene expression differences between humans and chimpanzees. Genetics 182:627–630
Carninci P, Sandelin A, Lenhard B, Katayama S, Shimokawa K et al (2006) Genome-wide analysis of mammalian promoter architecture and evolution. Nat Genet 38:626–635
Cohen CJ, Lock WM, Mager DL (2009) Endogenous retroviral LTRs as promoters for human genes: a critical assessment. Gene 448:105–114
Elagoz A, Henderson D, Babu PS, Salter S, Grahames C et al (2004) A truncated form of CKbeta8–1 is a potent agonist for human formyl peptide-receptor-like 1 receptor. Br J Pharmacol 141:37–46
Flagel LE, Wendel JF (2009) Gene duplication and evolutionary novelty in plants. New Phytol 183:557–564
Frazer KA, Chen X, Hinds DA, Pant PV, Patil N et al (2003) Genomic DNA insertions and deletions occur frequently between humans and nonhuman primates. Genome Res 13:341–346
Gao JL, Chen H, Filie JD, Kozak CA, Murphy PM (1998) Differential expansion of the N-formyl peptide receptor gene cluster in human and mouse. Genomics 51:270–276
Harada M, Habata Y, Hosoya M, Nishi K, Fujii R et al (2004) N-formylated humanin activates both formyl peptide receptor-Like 1 and 2. Biochem Biophys Res Commun 324:255–261
Haviland DL, Borel AC, Fleischer DT, Haviland JC, Wetsel RA (1993) Structure, 5′-flanking sequence, and chromosome location of the human N-formyl peptide receptor gene. A single-copy gene comprised of two exons on chromosome 19q.13.3 that yields two distinct transcripts by alternative polyadenylation. Biochemistry 32:4168–4174
Hughes TA (2006) Regulation of gene expression by alternative untranslated regions. Trends Genet 22:119–122
Hughes JF, Coffin JM (2001) Evidence for genomic rearrangements mediated by human endogenous retroviruses during primate evolution. Nat Genet 29:487–489
Hughes JF, Coffin JM (2005) Human endogenous retroviral elements as indicators of ectopic recombination events in the primate genome. Genetics 171:1183–1194
Hwang SB (1990) Specific receptors of platelet-activation factor, receptor heterogeneity, and signal transduction mechanisms. J Lipid Med 2:123–158
International Human Genome Sequencing Consortium (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Jurka J, Milosavljevic A (1991) Reconstruction and analysis of human Alu genes. J Mol Evol 32:105–121
Kimura K, Wakamatsu A, Suzuki Y, Ota T, Nishikawa T et al (2006) Diversification of transcriptional modulation: large-scale identification and characterization of putative alternative promoters of human genes. Genome Res 16:55–65
Koszul R, Caburet S, Dujon B, Fischer G (2004) Eucaryotic genome evolution through the spontaneous duplication of large chromosomal segments. EMBO J 23:234–243
Landry JR, Mager DL, Wilhelm BT (2003) Complex controls: the role of alternative promoters in mammalian genomes. Trends Genet 19:640–648
Le Y, Oppenheim JJ, Wang JM (2001) Pleiotropic roles of formyl peptide receptors. Cytokine Growth Factor Rev 12:91–105
Liberles SD, Horowitz LF, Kuang D, Contos JJ, Wilson KL et al (2009) Formyl peptide receptors are candidate chemosensory receptors in the vomeronasal organ. Proc Natl Acad Sci USA 106:9842–9847
Maldonado-Pérez D, Golightly E, Denison FC, Jabbour HN, Norman JE (2011) A role for lipoxin A4 as anti-inflammatory and proresolution mediator in human parturition. FASEB J 25:569–575
Migeotte I, Communi D, Parmentier M (2006) Formyl peptide receptors: a promiscuous subfamily of G protein-coupled receptors controlling immune responses. Cytokine Growth Factor Rev 17:501–519
Murnane JP (2006) Telomeres and chromosome instability. DNA Repair (Amst) 5:1082–1092
Newman TL, Tuzun E, Morrison VA, Hayden KE, Ventura M et al (2005) A genome-wide survey of structural variation between human and chimpanzee. Genome Res 15:1344–1356
Pâques F, Haber JE (1999) Multiple pathways of recombination induced by double-strand breaks in Saccharomyces cerevisiae. Microbiol Mol Biol Rev 63:349–404
Rhesus Macaque Genome Sequencing Analysis Consortium (2007) Evolutionary and biomedical insights from the rhesus macaque genome. Science 316:222–234
Riboldi E, Musso T, Moroni E, Urbinati C, Bernasconi S et al (2005) Cutting edge: proangiogenic properties of alternatively activated dendritic cells. J Immunol 175:2788–2792
Sequencing Chimpanzee, Consortium Analysis (2005) Initial sequence of the chimpanzee genome and comparison with the human genome. Nature 437:69–87
Sharp AJ, Cheng Z, Eichler EE (2006) Structural variation of the human genome. Annu Rev Genomics Hum Genet 7:407–442
She X, Liu G, Ventura M, Zhao S, Misceo D et al (2006) A preliminary comparative analysis of primate segmental duplications shows elevated substitution rates and a great-ape expansion of intrachromosomal duplications. Genome Res 16:576–583
Smith GP (1976) Evolution of repeated DNA sequences by unequal crossover. Science 191:528–535
Snyderman R, Uhing RJ (1992) Chemoattractant stimulus-response coupling. In: Gallin JI, Goldstein IM, Snyderman R (eds) Inflammation: Basic Principles and Clinical Correlates. Lippincott Williams & Wilkins, Philadelphia, pp 421–439
Stankiewicz P, Lupski JR (2002) Genome architecture, rearrangements and genomic disorders. Trends Genet 18:74–82
Stern DL (1998) A role of Ultrabithorax in morphological differences between Drosophila species. Nature 396:463–466
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tsuritani K, Irie T, Yamashita R, Sakakibara Y, Wakaguri H et al (2007) Distinct class of putative “non-conserved” promoters in humans: comparative studies of alternative promoters of human and mouse genes. Genome Res 17:1005–1014
Van Compernolle SE, Clark KL, Rummel KA, Todd SC (2003) Expression and function of formyl peptide receptors on human fibroblast cells. J Immunol 171:2050–2056
Wang ZG, Ye RD (2002) Characterization of two new members of the formyl peptide receptor gene family from 129S6 mice. Gene 299:57–63
Wilson MD, Cheung J, Martindale DW, Scherer SW, Koop BF (2006) Comparative analysis of the paired immunoglobulin-like receptor (PILR) locus in six mammalian genomes: duplication, conversion, and the birth of new genes. Physiol Genomics 27:201–218
Wu C, Orozco C, Boyer J, Leglise M, Goodale J et al (2009) BioGPS: an extensible and customizable portal for querying and organizing gene annotation resources. Genome Biol 10:R130
Yang D, Chen Q, Gertz B, He R, Phulsuksombati M, Ye RD, Oppenheim JJ (2002) Human dendritic cells express functional formyl peptide receptor-like-2 (FPRL2) throughout maturation. J Leukoc Biol 72:598–607
Yang EM, Kim SH, Kim NH, Park HS (2010) The genetic association of the FPRL1 promoter polymorphism with chronic urticaria in a Korean population. Ann Allergy Asthma Immunol 105:96–97
Acknowledgments
We thank Dr. Jungnam Lee and Nicholas Meyerson for comments on the manuscript. This study was financially supported by Pusan National University in program Post-Doc 2010.
Accession numbers
AB500093: Callithrix jacchus DNA, MER21C element including AluSx, complete sequence.
AB500092: Ateles geoffroyi DNA, LTR 54 element, complete sequence.
AB500091: Aotus trivirgatus DNA, LTR 54 element, complete sequence.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
335_2011_9341_MOESM1_ESM.tif
Supplementary material 1: Phylogenetic tree of LTR54 related FPR3 gene in various primates. The DR-LTR54 and OR-LTR54 families were completely separated into DR and OR groups, and single LTR54 clustered with the OR group. The black boxes indicate DR-LTR54 in various primates, and the white boxes indicate OR-LTR54 in various primates. The white circles show single LTR54 in New World monkeys. As an out group, paralogous LTR54 randomly selected in human genome was used (TIFF 111 kb)
335_2011_9341_MOESM2_ESM.tif
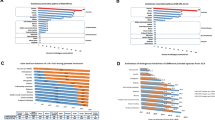
Supplementary material 2: Comparison of expression patterns among FPR family. Expression data was previously reported by Gao et al. 1998; Wang et al. 2002; Elagoz et al. 2004; Harada et al. 2004; Riboldi et al. 2005 (direct real-time PCR data, transformed RT-PCR and Northern data through ImageJ program) (TIFF 142 kb)
335_2011_9341_MOESM3_ESM.tif
Supplementary material 3: Quantitative analyses of human FPR2-1 transcripts. Graphs indicate expression of human FPR2-1 in various tissue panels. Each column represents the mean value with a standard error in triplicate determinations (TIFF 99 kb)
335_2011_9341_MOESM4_ESM.tif
Supplementary material 4: Quantitative analyses of FPR3-1 and FPR3-2 transcripts in A549 and HeLa cells. Graphs show expression pattern of (a) human FPR3-1, (b) human FPR3-2, in A549 and HeLa cells. Notably, the human FPR3-2 was not expressed in A549 cells. Each column represents the mean value with a standard error in triplicate determinations (TIFF 75 kb)
335_2011_9341_MOESM5_ESM.tif
Supplementary material 5: Conservation of FPR3 gene. Analysis of conservation by Phylop (a) and Phastcons (b) of UCSC genome browser and VISTA browser (c) (TIFF 426 kb)
335_2011_9341_MOESM6_ESM.tif
Supplementary material 6: Analysis of histone modification status. (a) Histone modification status of SR within human FPR2 gene (b) Histone modification status of DR and OR within human FPR3 gene. Boxes indicate SR, DR, and OR (TIFF 394 kb)
Rights and permissions
About this article
Cite this article
Ha, HS., Huh, JW., Gim, JA. et al. Transcriptional variations mediated by an alternative promoter of the FPR3 gene. Mamm Genome 22, 621–633 (2011). https://doi.org/10.1007/s00335-011-9341-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00335-011-9341-7



