Abstract
Key message
Most of the upregulated genes contributed to the accumulation of soluble sugars and ABA in the phloem of ‘Vitis amurensis’ compared to ‘Merlot’ during cold acclimation.
Abstract
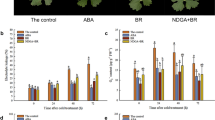
Extreme cold is one of the dominant abiotic factors affecting grape yield and quality. However, the changes in sugars, phytohormones, and gene expression in the branch phloem of different tolerant grape varieties during cold acclimation remain elusive. The data supported that with decreasing temperature, the contents of fructose, sucrose, and ABA in the phloem of Vitis amurensis (cold-tolerant, T) and ‘Merlot’ (cold-sensitive, S) increased during cold acclimation, and these indicators were higher in T than in S. Furthermore, the activities of sucrose synthetase, sucrose phosphate synthetase, and acid invertase peaked in the early phase of cold acclimation (approximately 5 °C) compared to other phases (approximately 28 °C, 0 °C, − 5 °C and − 10 °C). Moreover, the RNA sequencing results helped identify a total of 11,343 differentially expressed genes in the phloem of T and S, among which 4912 were upregulated and 6431 were downregulated. In the abscisic acid pathway, CRTISO, PSPY1-1, CYCP707A4-2, PYL4-1, PYL4-2, P2C08, SAPK2, TARAB1, and DBF3 were more highly expressed in T than in S. In the starch and sucrose metabolism pathway, HXK1, PGMP, GLGL1, SUS6, VCINV, BGL11, SSY1, GPS, BAM1 and BAM3 were also more highly expressed in T than in S. Moreover, the genes related to oxidative phosphorylation, such as NDHF, ND4, ND1, NAD7, NAD2, ATPB, YMF19, ATP9, PMA1 and AHA8, were upregulated in T. These results will be beneficial for understanding the potential differences in tolerance across two different cold-tolerant grapes with respect to sugar metabolism and gene expression.







Similar content being viewed by others
Data availability
The RNA-sequencing data have been deposited with NCBI (https://submit.ncbi.nlm.nih.gov/subs/sra/) under BioProject PRJNA793284.
Abbreviations
- RNA-seq:
-
Transcriptome sequencing
- T:
-
Vitis amurensis
- S:
-
Merlot
- IAA:
-
Auxin
- ABA:
-
Abscisic acid
- SuSy:
-
Sucrose synthase
- SPS:
-
Sucrose phosphate synthase
- AI:
-
Acid invertase
- DEGs:
-
Differentially expressed genes
- ROS:
-
Reactive oxygen species
- d:
-
Day
- UDP:
-
Nucleoside diphosphate
- GO:
-
Gene Ontology resource database
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes database
- TFs:
-
Transcription factors
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
References
Ali M, Baek K (2020) Jasmonic acid signaling pathway in response to abiotic stresses in plants. Int J Mol Sci 21(2):621
Améglio T, Decourteix M, Alves G, Valentin V, Sakr S, Julien JL, Petel G, Guilliot A, Lacointe A (2004) Temperature effects on xylem sap osmolarity in walnut trees: evidence for a vitalistic model of winter embolism repair. Tree Physiol 24:785–793
An D, Yang J, Zhang P (2012) Transcriptome profiling of low temperature treated cassava apical shoots showed dynamic responses of tropical plant to cold stress. BMC Genomics 13(1):1–25
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11(10):R106
Anna L, Teemu P, Tuula J et al (2016) Osmolality and non-structural carbohydrate composition in the secondary phloem of trees across a latitudinal gradient in Europe. Front Plant Sci 7:726
Aslam M, Sugita K, Qin Y, Rahman A (2020) Aux/IAA14 regulates microRNA-mediated cold stress response in Arabidopsis roots. Int J Mol Sci 21:8441
Cai Y, Yan J, Li Q, Deng Z, Liu S, Lu J, Zhang Y (2019) Sucrose transporters of resistant grapevine are involved in stress resistance. Plant Mol Biol 100(1):111–132
Carvalho LC, Coito JL, Gonçalves EF, Chaves MM, Amâncio S (2015) Differential physiological response of the grapevine varieties Touriga Nacional and Trincadeira to combined heat, drought and light stresses. Plant Biol 18(suppl 1):101–111
Charrier G, Cochard H, Améglio T (2013) Evaluation of the impact of frost resistances on potential altitudinal limit of trees. Tree Physiol 33:891–902
Choi H, Hong J, Ha J, Kang J, Kim S (2000) ABFs, a family of ABA-responsive element binding factors. J Biol Chem 275:1723–1730
Déjardin A, Sokolov LN, Kleczkowski LA (1999) Sugar/osmoticum levels modulate differential abscisic acid-independent expression of two stress-responsive sucrose synthase genes in Arabidopsis. Biochem J 344(Pt 2):503–509
Dong S, Beckles DM (2019) Dynamic changes in the starch-sugar interconversion within plant source and sink tissues promote a better abiotic stress response. J Plant Physiol 234–235:80–93
Du H, Wu N, Fu J, Wang S, Li X, Xiao J, Xiong L (2012) A GH3 family member, OsGH3-2, modulates auxin and abscisic acid levels and differentially affects drought and cold tolerance in rice. J Exp Bot 63(18):6467–6480
Fryzova R, Pohanka M, Martinkova P, Cihlarova H, Brtnicky M, Hladky J, Kynicky J (2018) Oxidative stress and heavy metals in plants. Rev Environ Contam Toxicol 245:129–156
Gong X, Liu M, Zhang L, Ruan Y, Ding R, Ji Y, Zhang N, Zhang S, Farmer J, Wang C (2015) Arabidopsis AtSUC2 and AtSUC4, encoding sucrose transporters, are required for abiotic stress tolerance in an ABA-dependent pathway. Physiol Plant 153(1):119–136
Gong Z, Xiong L, Shi H, Yang S, Herrera-Estrella LR, Xu G, Chao DY, Li J, Wang PY, Qin F, Li J, Ding Y, Shi Y, Wang Y, Yang Y, Guo Y, Zhu JK (2020) Plant abiotic stress response and nutrient use efficiency. Sci China Life Sci 63(5):635–674
Halimaa P, Blande D, Aarts MGM, Tuomainen M, Tervahauta A, Kärenlampi S (2014) Comparative transcriptome analysis of the metal hyperaccumulator Noccaea caerulescens. Front Plant Sci 5:213–220
He J, Duan Y, Hua D, Fan G, Wang L, Liu Y, Chen Z, Han L, Qu LJ, Gong Z (2012) DEXH box RNA helicase-mediated mitochondrial reactive oxygen species production in Arabidopsis mediates crosstalk between abscisic acid and auxin signaling. Plant Cell 24(5):1815–1833
Höll J, Vannozzi A, Czemmel S, D’Onofrio C, Walker A, Rausch T, Lucchin M, Boss P, Dry I, Bogs J (2013) The R2R3-MYB transcription factors MYB14 and MYB15 regulate stilbene biosynthesis in Vitis vinifera. Plant Cell 25(10):4135–4149
Hopff D, Wienkoop S, Lüthje S (2013) The plasma membrane proteome of maize roots grown under low and high iron conditions. J Proteomics 91:605–618
Horiuchi R, Arakawa K, Kasuga J, Suzuki T, Jitsuyama Y (2021) Freezing resistance and behavior of winter buds and canes of wine grapes cultivated in northern Japan. Cryobiology 101:44–51
Huang X, Chen M, Yang L, Li Y, Wu J (2015) Effects of exogenous abscisic acid on cell membrane and endogenous hormone contents in leaves of sugarcane seedlings under cold stress. Sugar Tech 17(1):59–64
Huang X, Shi H, Hu Z, Liu A, Amombo E, Chen L, Fu J (2017) ABA is involved in regulation of cold stress response in bermudagrass. Front Plant Sci 8:1613
Jain M, Khurana J (2009) Transcript profiling reveals diverse roles of auxin-responsive genes during reproductive development and abiotic stress in rice. FEBS J 276(11):3148–3162
Kenchanmane RSK, Barnes AC, Schnable JC, Roston RL (2018) Low-temperature tolerance in land plants: are transcript and membrane responses conserved? Plant Sci 276:73–86
Kim S, Kang JY, Cho DI, Park JH, Kim SY (2004) ABF2, an ABRE-binding bZIP factor, is an essential component of glucose signaling, and its overexpression affects multiple stress to lerance. Plant J 40(1):75–87
Kim Y, Choi K, Khan A, Waqas M, Lee I (2016) Exogenous application of abscisic acid regulates endogenous gibberellins homeostasis and enhances resistance of oriental melon (Cucumismelo var. L.) against low temperature. Sci Hortic 207:41–47
Knight MR, Knight H (2012) Low-temperature perception leading to gene expression and cold tolerance in higher plants. New Phytol 195(4):737–751
Krasensky J, Jonak C (2012) Drought, salt, and temperature stress-induced metabolic rearrangements and regulatory networks. J Exp Bot 63(4):1593–1608
Livak K, Schmittgen T (2001) Analysis of relative gene expression data using real-time quantitative and the 2-ΔΔCT method. Methods 25(4):402–408
Mao J, Li W, Mi B, Ma Z, Dawuda M, Zuo C, Zhang Y, Jiang X, Chen B (2018) Transcriptome analysis revealed glucose application affects plant hormone signal transduction pathway in “Red Globe” grape plantlets. Plant Growth Regul 84:45–56
Mcadam S, Brodribb T (2018) Mesophyll cells are the main site of abscisic acid biosynthesis in water-stressed leaves. Plant Physiol 177(3):911–917
McAdam S, Brodribb T, Ross J (2016) Shoot-derived abscisic acid promotes root growth. Plant Cell Environ 39(3):652–659
Meisrimler CN, Planchon S, Renaut J, Sergeant K, Lüthje S (2011) Alteration of plasma membrane-bound redox systems of iron deficient pea roots by chitosan. J Proteomics 74(8):1437–1449
Miura K, Lee J, Gong Q, Ma S, Jin J, Yoo C, Miura T, Sato A, Bohnert H, Hasegawa P (2011) SIZ1 regulation of phosphate starvation-induced root architecture remodeling involves the control of auxin accumulation. Plant Physiol 155:1000–1012
Nemhauser JL, Hong F, Chory J (2006) Different plant hormones regulate similar processes through largely nonoverlapping transcriptional responses. Cell 126(3):467–475
Panagopoulou EA, Chiou A, Nikolidaki EK, Christea M, Karathanos VT (2019) Corinthian raisins (Vitis vinifera L, var Apyrena) antioxidant and sugar content as affected by the drying process: a 3-year study. J Sci Food Agric 99(2):915–922
Paulino P, Mauricio R, Guedes C, Rensing S, Birgit K, Bernd M (2010) Plntfdb: updated content and new features of the plant transcription factor database. Nucleic Acids Res 38(Database issue):D822–D827
Peng T, Zhu XF, Duan N, Liu J (2014) PtrBAM1, a β-amylase-coding gene of Poncirus trifoliata, is a CBF regulon member with function in cold tolerance by modulating soluble sugar levels. Plant Cell Environ 37(12):2754–2767
Prerostova S, Černý M, Dobrev PI, Motyka V, Hluskova L, Zupkova B, Gaudinova A, Knirsch V, Janda T, Brzobohatý B, Vankova R (2021) Light regulates the cytokinin-dependent cold stress responses in Arabidopsis. Front Plant Sci 11:608711
Rahman IU, Afzal A, Iqbal Z et al (2020) Response of plant physiological attributes to altitudinal gradient: plant adaptation to temperature variation in the Himalayan region. Sci Total Environ 706:135714
Reinhold H, Soyk S, Simková K, Hostettler C, Marafino J, Mainiero S, Vaughan CK, Monroe JD, Zeeman SC (2011) β-amylase-like proteins function as transcription factors in Arabidopsis, controlling shoot growth and development. Plant Cell 23(4):1391–1403
Ruan YL, Jin Y, Yang YJ, Li GJ, Boyer JS (2010) Sugar input, metabolism, and signaling mediated by invertase: roles in development, yield potential, and response to drought and heat. Mol Plant 3(6):942–955
Sah S, Reddy K, Li J (2016) Abscisic acid and abiotic stress tolerance in crop plants. Front Plant Sci 7:571
Sheteiwy MS, An J, Yin M, Jia X, Guan Y, He F, Hu J (2019) Cold plasma treatment and exogenous salicylic acid priming enhances salinity tolerance of Oryza sativa seedlings. Protoplasma 256:79–99
Shi Y, Ding Y, Yang S (2015) Cold signal transduction and its interplay with phytohormones during cold acclimation. Plant Cell Physiol 56(1):7–15
Shibasaki K, Uemura M, Tsurumi S, Rahman A (2009) Auxin response in Arabidopsis under cold stress: underlying molecular mechanisms. Plant Cell 21:3823–3838
Shin H, Oh Y, Kim D (2015) Differences in cold hardiness, carbohydrates, dehydrins and related gene expressions under an experimental deacclimation and reacclimation in Prunus persica. Physiol Plant 154:485–499
Steppe K, Sterck F, Deslauriers A (2015) Diel growth dynamics in tree stems: linking anatomy and ecophysiology. Trends Plant Sci 20(6):335–343
Sun X, Matus J, Wong D et al (2018) The GARP/MYB-related grape transcription factor AQUILO improves cold tolerance and promotes the accumulation of raffinose family oligosaccharides. J Exp Bot 69(7):1749–1764
Tan H, Huang H, Tie M, Tang Y, Lai Y, Li H (2016) Transcriptome profiling of two Asparagus Bean (Vigna unguiculata subsp. sesquipedalis) cultivars differing in chilling tolerance under cold stress. PLoS ONE 11(3):e0151105
Wang Y, Xin H, Fan P, Zhang J, Liu Y, Dong Y, Zemin W, Yang Y, Zhang Q, Ming R, Zhaong G, Li S, Liang Z (2020) The genome of Shanputao (Vitis amurensis) provides a new insight into cold tolerance of grapevine. Plant J 105(6):1495–1506
Wang Y, Jiang H, Mao Z, Liu W, Jiang S, Xu H, Su M, Zhang J, Wang N, Zhang Z, Chen X (2021) Ethylene increases the cold tolerance of apple via the MdERF1B-MdCIbHLH1 regulatory module. Plant J 106(2):379–393
Wei T, Wang Y, Xie Z, Guo D, Chen C, Fan Q, Deng X, Liu JH (2019) Enhanced ROS scavenging and sugar accumulation contribute to drought tolerance of naturally occurring autotetraploids in Poncirus trifoliata. Plant Biotechnol J 17(7):1394–1407
Xin H, Zhu W, Wang L, Xiang Y, Fang L, Li J, Sun X, Wang N, Londo J, Li S (2013) Genome wide transcriptional profile analysis of Vitis amurensis and Vitis vinifera in response to cold stress. PLoS ONE 8(3):e58740
Xu W, Li R, Zhang N, Ma F, Jiao Y, Wang Z (2014) Transcriptome profiling of Vitis amurensis, an extremely cold-tolerant Chinese wild Vitis species, reveals candidate genes and events that potentially connected to cold stress. Plant Mol Biol 86(4–5):527–541
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Yang H, Wang T, Yu X, Yang Y, Wang X (2020) Enhanced sugar accumulation and regulated plant hormone signalling genes contribute to cold tolerance in hypoploid Saccharum spontaneum. BMC Genomics 21(1):507
Zeng Y, Yu J, Cang J, Liu LJ, Mu Y, Wang J, Zhang D (2011) Detection of sugar accumulation and expression levels of correlative key enzymes in winter wheat (Triticum aestivum) at low temperatures. J Agric Chem Soc Japan 75(4):681–687
Zhang H, Zhao Y, Zhu J (2020) Thriving under stress: how plants balance growth and the stress response. Dev Cell 55(5):529–543
Zhao Y (2010) Auxin biosynthesis and its role in plant development. Annu Rev Plant Biol 61:49–64
Zhu J (2016) Abiotic stress signaling and responses in plants. Cell 167(2):313–324
Zwieniecki M, Tixier A, Sperling O (2015) Temperature-assisted redistribution of carbohydrates in trees. Am J Bot 102(8):1216–1218
Acknowledgements
We thank State Key Laboratory of Aridland Crop Science, Gansu Agricultural University provides service of plant hormones determination.
Funding
This working was supported by the FuXi Foundation of Gansu Agricultural University (No. Ganfx-03J02), Youth Innovation and Entrepreneurship Talent Project of Longyuan (2018LYQN01) and the Science and Technology Major Project of Gansu Province (18ZD2NA006).
Author information
Authors and Affiliations
Contributions
JM and BHC designed the experiments. GPL and JM wrote the manuscript. GPL, ZHM, SXL, WFM and LDF conducted the experiments. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethics approval
Not applicable.
Additional information
Communicated by Emmanuel Guiderdoni.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
299_2022_2862_MOESM1_ESM.tif
Fig. S1 (a) Heatmap clusters representing 154 upregulated DEGs from Fig. 2b (red circles are upregulated genes in the Venn diagram) in the B, C, D, and E phases of T vs S. (b) Likewise, heatmap clusters represent 79 downregulated DEGs from Fig. 2b (blue circles are downregulated genes in the Venn diagram) in the B, C, D, and E phases of T vs S. The expression levels of DEGs are represented by log2 (FPKM) (TIF 3196 KB)
299_2022_2862_MOESM2_ESM.tif
Fig. S2 Co-expression trend analysis of all DEGs. (a), (c), (e), (g), (i), and (k) represent the six different expression trends of DEGs in T and S. (b), (d), (f), (h), (j), and (l) represent the significant pathways of the corresponding gene cluster annotated by KEGG (TIF 3070 KB)
299_2022_2862_MOESM3_ESM.tif
Fig. S3 GO functional enrichment analysis of the DEGs. The X-axis indicates the number of DEGs, and the Y-axis indicates the GO categories (TIF 988 KB)
299_2022_2862_MOESM4_ESM.tif
Fig. S4 Differentially expressed TFs. (a) Types and quantities of TFs. The X-axis indicates the number of DEGs, and the Y-axis indicates the type of TF. (b) The number of similar TF genes in T vs S under different phases during cold acclimation (TIF 2548 KB)
Rights and permissions
About this article
Cite this article
Liang, G., Ma, Z., Lu, S. et al. Temperature-phase transcriptomics reveals that hormones and sugars in the phloem of grape participate in tolerance during cold acclimation. Plant Cell Rep 41, 1357–1373 (2022). https://doi.org/10.1007/s00299-022-02862-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-022-02862-1