Abstract
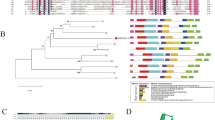
Plants respond to stress conditions through early stress-response factors (ESRF), which serve the function of stress sensing and/or signal transduction. These mainly comprise qualitative and/or quantitative flux in the redox molecules, calcium ions (Ca2+), phosphatidic acid, hexose sugars and phytohormones. The role of resident proteins such as phytohormone receptors and G-proteins as first messengers under stress is well established. Yet, within the modern omics context, most of the stress response at the protein level is injudiciously attributed to substantial up- or down-regulation of expression measured at the RNA or protein level. Proteins such as kinases and transcription factors (TFs) that exhibit cascade effects are primary candidates for studies in plant stress tolerance. However, resident-protein post-translational modification (PTM), specifically in response to particular conditions such as stress, is a candidate for immediate and potent ‘quick reaction force’ (QRF) kind of effects. Stress-mediated SUMOylation of TFs and other proteins have been observed. SUMOylation can change the rate of activity, function or location of the modified protein. Early SUMOylation of resident proteins can act in the stress signal transduction or in adaptive response. Here, we consider brief background information on ESRFs to establish the crosstalk between these factors that impinge on PTMs. We then illustrate connections of protein SUMOylation to phytohormones and TFs. Finally, we present results of an in silico analysis of rice Receptor-Like Kinases, heat-shock and calcium-binding proteins to identify members of these gene families, whose basal expression under drought but potential SUMOylation presents them as QRF candidates for roles in stress signaling/response.
Similar content being viewed by others
References
Agarwal M, Sahi C, Katiyar-Agarwal S, Agarwal S, Young T, Gallie DR, Sharma VM, Ganesan K, Grover A (2003) Molecular characterization of rice hsp101: complementation of yeast hsp104 mutation by disaggregation of protein granules and differential expression in indica and japonica rice types. Plant Mol Biol 51:543–553
Alonso-Ramírez A, Rodríguez D, Reyes D, Jiménez JA, Nicolás G, López-Climent M, Gómez-Cadenas A, Nicolás C (2009) Cross-talk between gibberellins and salicylic acid in early stress responses in Arabidopsis thaliana seeds. Plant Signal Behav 4:750–751
Alvim FC, Carolino SMB, Cascardo JCM, Nunes CC, Martinez CA, Otoni WC, Fontes EBP (2001) Enhanced accumulation of BiP in transgenic plants confers tolerance to water stress. Plant Physiol 126:1042–1054
Apuya NR, Yadegari R, Fischer RL, Harada JJ, Zimmerman JL, Goldberg RB (2001) The Arabidopsis embryo mutant schlepperless has a defect in the chaperonin-60β gene. Plant Physiol 126:717–730
Boonburapong B, Buaboocha T (2007) Genome-wide identification and analyses of the rice calmodulin and related potential calcium sensor proteins. BMC Plant Biol 7:4
Buchner J (1999) Hsp90 & Co.—a holding for folding. Trends Biochem Sci 24:136–141
Bukau B, Horwich AL (1998) The Hsp70 and Hsp60 chaperone machines. Cell 92:351–366
Carvalho MHC (2008) Drought stress and reactive oxygen species: production, scavenging and signaling. Plant Signal Behav 3:156–165
Catala R, Ouyang J, Abreu IA, Hu Y, Seo H, Zhang X, Chua NH (2007) The Arabidopsis E3 SUMO ligase SIZ1 regulates plant growth and drought responses. Plant Cell 19:2952–2966
Chaikam V, Karlson DT (2010) Response and transcriptional regulation of rice SUMOylation system during development and stress conditions. Biochem Mol Biol Rep 43:103–109
Chauhan H, Khurana N, Agarwal P, Khurana P (2011) Heat shock factors in rice (Oryza sativa L.): genome-wide expression analysis during reproductive development and abiotic stress. Mol Genet Genomics 286:171–187
Chinnusamy V, Zhu J, Zhu JK (2007) Cold stress regulation of gene expression in plants. Trends Plant Sci 12:444–451
Chosed R, Mukherjee S, Lois LM, Orth K (2006) Evolution of a signalling system that incorporates both redundancy and diversity: Arabidopsis SUMOylation. Biochem J 15:521–529
Cohen-Peer R, Schuster S, Meiri D, Breiman A, Avni A (2010) Sumoylation of Arabidopsis heat shock factor A2 (HsfA2) modifies its activity during acquired thermo tolerance. Plant Mol Biol 74:33–45
Colby T, Matthai A, Boechelmann A, Stuible HP (2006) SUMO-Conjugating and SUMO-Deconjugating Enzymes from Arabidopsis. Plant Physiol 142:318–332
Conti L, Kioumourtzoglou D, O’Donnell E, Dominy P, Sadanandom A (2009) OTS1 and OTS2 SUMO proteases link plant development and survival under salt stress. Plant Signal Behav 4:225–227
Cui X, Luan S (2012) A new wave of hormone research: crosstalk mechanisms. Mol Plant 5:959–960
Day I, Reddy V, Ali GS, Reddy ASN (2002) Analysis of EF-hand-containing proteins in Arabidopsis. Genome Biol 3:research0056–research0056.24
DeFalco TA, Bender KW, Snedden WA (2010) Breaking the code: Ca2+ sensors in plant signalling. Biochem J 425:27–40
Elrouby N, Coupland G (2010) Proteome-wide screens for small ubiquitin-like modifier (SUMO) substrates identify Arabidopsis proteins implicated in diverse biological processes. Proc Natl Acad Sci USA 107:17415–17420
Gao LL, Xue HW (2012) Global analysis of expression profiles of rice receptor-like kinase genes. Mol Plant 5:143–153
Harrison M (2012) Cross-talk between phytohormone signaling pathways under both optimal and stressful environmental conditions. In: Khan et al. (eds) Phytohormones and abiotic stress tolerance in plants. Springer, Berlin, pp 49–76
Hart FU (1996) Molecular chaperones in cellular protein folding. Nature 381:571–580
Hay RT (2005) SUMO: a history of modification. Mol Cell 18:1–12
Hermkes R, Fu YF, Nürrenberg K, Budhiraja R, Schmelzer E, Elrouby N, Dohmen RJ, Bachmair A, Coupland G (2011) Distinct roles for Arabidopsis SUMO protease ESD4 and its closest homolog ELS1. Planta 233:63–73
Hruz T, Laule O, Szabo G, Wessendorp F, Bleuler S, Oertle L, Widmayer P, Gruissem W, Zimmermann P (2008) Genevestigator V3: a reference expression database for the meta-analysis of transcriptomes. Adv Bioinforma 2008(420747):1–5. doi:10.1155/2008/420747
Hu H, Dai M, Yao J, Xiao B, Li X, Zhang Q, Xiong L (2006) Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci USA 103:12987–12992
Hu W, Hu G, Han B (2009) Genome-wide survey and expression profiling of heat shock proteins and heat shock factors revealed overlapped and stress specific response under abiotic stresses in rice. Plant Sci 176:583–590
Huang L, Yang S, Zhang S, Liu M, Lai J, Qi Y, Shi S, Wang J, Wang Y, Xie Q, Yang C (2009) The Arabidopsis SUMO E3 ligase AtMMS21, a homologue of NSE2/MMS21, regulates cell proliferation in the root. Plant J 60:666–678
Huang GT, Ma SL, Bai LP, Zhang L, Ma H, Jia P, Liu J, Zhong M, Guo ZF (2012) Signal transduction during cold, salt, and drought stresses in plants. Mol Biol Rep 39:969–987
Imai J, Maruya M, Yashiroda H, Yahara I, Tanaka K (2003) The molecular chaperone Hsp90 plays a role in the assembly and maintenance of the 26S proteasome. EMBO J 22:3557–3567
Ishikawa A, Tanaka H, Nakai M, Asahi T (2003) Deletion of a chaperonin 60β gene leads to cell death in the Arabidopsis lesion initiation 1 mutant. Plant Cell Physiol 44:255–261
Jacob T, Ritchie S, Assmann SM, Gilroy S (1999) Abscisic acid signal transduction in guard cells is mediated by phospholipase D activity. Proc Natl Acad Sci USA 96:12192–12197
Jaglo-Ottosen KR, Gilmour SJ, Zarka DG, Schabenberger O, Thomashow MF (1998) Arabidopsis CBF1 overexpression induces COR genes and enhances freezing tolerance. Science 280:104–106
Jin JB, Jin YH, Lee J, Miura K, Yoo CY, Kim WY, Van Oosten M, Hyun Y, Somers DE, Lee I, Yun DJ, Bressan RA, Hasegawa PM (2008) The SUMO E3 ligase, AtSIZ1, regulates flowering by controlling a salicylic acid-mediated floral promotion pathway and through affects on FLC chromatin structure. Plant J 53:530–540
Kasuga M, Liu Q, Miura S, Yamaguchi-Shinozaki K, Shinozaki K (1999) Improving plant drought, salt, and freezing tolerance by gene transfer of a single stress-inducible transcription factor. Nat Biotechnol 17:287–291
Kilian J, Whitehead D, Horak J, Wanke D, Weinl S, Batistic O, D’Angelo C, Bornberg-Bauer E, Kudla J, Harter K (2007) The AtGenExpress global stress expression data set: protocols, evaluation and model data analysis of UV-B light, drought and cold stress responses. Plant J 50:347–363
Kim BH, Schoffl F (2002) Interaction between Arabidopsis heat shock transcription factor 1 and 70 kDa heat shock proteins. J Exp Bot 53:371–375
Knight H, Knight MR (2001) Abiotic stress signalling pathways: specificity and cross-talk. Trends Plant Sci 6:262–267
Ko JH, Yang SH, Han KH (2006) Upregulation of an Arabidopsis RING-H2 gene, XERICO, confers drought tolerance through increased abscisic acid biosynthesis. Plant J 47:343–355
Kobayashi Y, Murata M, Minami H, Yamamoto S, Kagaya Y, Hobo T, Yamamoto A, Hattori T (2005) Abscisic acid-activated SNRK2 protein kinases function in the gene-regulation pathway of ABA signal transduction by phosphorylating ABA response element-binding factors. Plant J 44:939–949
Kohli A, Sreenivasulu N, Kumar P (2013) The phytohormone crosstalk paradigm takes center stage in understanding how plants respond to abiotic stresses. Plant Cell Rep (in press)
Kolukisaoglu U, Weinl S, Blazevic D, Batistic O, Kudla J (2004) Calcium sensors and their interacting protein kinases: genomics of the Arabidopsis and rice CBL–CIPK signaling networks. Plant Physiol 134:43–58
Kudla J, Batisic O, Hashimoto K (2010) Calcium signals: the lead currency of plant information processing. Plant Cell 22:541–563
Kurepa J, Walker JM, Smalle J, Gosink MM, Davis SJ, Durham TL, Sung DY, Vierstra RD (2003) The small ubiquitin-like modifier (SUMO) protein modification system in Arabidopsis. J Biol Chem 278:6862–6872
Lee JH, Schöffl F (1996) An Hsp70 antisense gene affects the expression of HSP70/HSC70, the regulation of HSF, and the acquisition of thermo tolerance in transgenic Arabidopsis thaliana. Mol Gen Genet 252:11–19
Lee J, Nam J, Park HC, Na G et al (2006) Salicylic acid-mediated innate immunity in Arabidopsis is regulated by SIZ1 SUMO E3 ligase. Plant J 49:79–90
Li WX, Oono Y, Zhu J, He XJ, Wu JM, Iida K, Lu XY, Cui X, Jin H, Zhu JK (2008) The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and posttranscriptionally to promote drought resistance. Plant Cell 20:2238–2251
Lois LM, Lima CD, Chua NH (2003) Small ubiquitin-like modifier modulates abscisic acid signaling in Arabidopsis. Plant Cell 15:1347–1359
Lott N, Ross T (2006) Tracking and evaluating US billion dollar weather disasters, 1980–2005. AMS forum: environmental risk and impacts on society: successes and challenges. http://www1.ncdc.noaa.gov/pub/data/papers/200686ams1.2nlfree.pdf. Accessed 30 Jan 2006
Lyzenga WJ, Stone SL (2011) Protein ubiquitination: an emerging theme in plant abiotic stress tolerance. Am J Plant Sci Biotechnol (Special Issue 2) 5:1–11
Lyzenga WJ, Stone SL (2012) Abiotic stress tolerance mediated by protein ubiquitination. J Exp Bot 63:599–616
Marshall A, Aalen RB, Audenaert D, Beeckman T, Broadley MR et al (2012) Tackling drought stress: RECEPTOR-LIKE KINASES present new approaches. Plant Cell 24:2262–2278
Matarasso N, Schuster S, Avni A (2005) A novel plant cysteine protease has a dual function as a regulator of 1-aminocyclopropane-1-carboxylic acid synthase gene expression. Plant Cell 17:1205–1216
Mazzucotelli E, Mastrangelo AM, Crosatti C, Guerra D, Stanca AM, Cattivelli L (2008) Abiotic stress response in plants: when post-transcriptional and post-translational regulations control transcription. Plant Sci 174:420–431
Melchior F, Schergaut M, Pichler A (2003) SUMO: ligases, isopeptidases and nuclear pores. Trends Biochem Sci 28:612–618
Mittler R, Vanderauwera S, Gollery M, Van Breusegem F (2004) Reactive oxygen gene network of plants. Trends Plant Sci 9:490–498
Miura K, Hasegawa PM (2010) Sumoylation and other ubiquitin-like post-translational modifications in plants. Trends Cell Biol 20:223–232
Miura K, Rus A, Sharkhuu A, Yokoi S, Karthikeyan AS, Raghothama KG, Baek D, Koo YD, Jin JB, Bressan RA, Yun DJ, Hasegawa PM (2005) The Arabidopsis SUMO E3 ligase SIZ1 controls phosphate deficiency responses. Proc Natl Acad Sci USA 102:7760–7765
Miura K, Jin JB, Lee J, Yoo CY, Stirm V, Miura T, Ashworth EN, Bressan RA, Yun DJ, Hasegawa PM (2007) SIZ1-mediated sumoylation of ICE1 controls CBF3/DREB1A expression and freezing tolerance in Arabidopsis. Plant Cell 19:1403–1414
Miura K, Leec J, Jina JB, Yooa CY, Miuraa T, Hasegawaa PM (2009) Sumoylation of ABI5 by the Arabidopsis SUMO E3 ligase SIZ1 negatively regulates abscisic acid signalling. Proc Natl Acad Sci USA 106:5418–5423
Miura K, Lee J, Miura T, Hasegawa PM (2010) SIZ1 controls cell growth and plant development in arabidopsis through salicylic acid. Plant Cell Physiol 51:103–113
Miura K, Lee J, Gong Q, Ma S, Jin JB, Yoo CY, Miura T, Sato A, Bohnert HJ, Hasegawa PM (2011) SIZ1 regulation of phosphate starvation-induced root architecture remodeling involves the control of auxin accumulation. Plant Physiol 155:1000–1012
Mori IC, Schroeder JI (2004) Reactive oxygen species activation of plant Ca2+ channels. A signaling mechanism in polar growth, hormone transduction, stress signaling, and hypothetically mechanotransduction. Plant Physiol 135:702–708
Morimoto RI (1998) Regulation of the heat shook transcriptional response: cross talk between a family of heat shock factors, molecular chaperones, and negative regulators. Genes Dev 12:3788–3796
Mukhopadhyay D, Dasso M (2007) Modification in reverse: the SUMO proteases. Trends Biochem Sci 32:286–295
Murakami T, Matsuba S, Funatsuki H, Kawaguchi K, Saruyama H, Tanida M, Sato Y (2004) Over-expression of a small heat shock protein, sHSP17.7, confers both heat tolerance and UV-B resistance to rice plants. Mol Breed 13:165–175
Nakashima K, Shinwari ZK, Sakuma Y et al (2000) Organization and expression of two Arabidopsis DREB2 genes encoding DRE-binding proteins involved in dehydration- and highsalinity-responsive gene expression. Plant Mol Biol 42:657–665
Nigam N, Singh A, Sabi C, Chandramouli A, Grover A (2008) SUMO-conjugating enzyme (Sce) and FK506-binding protein (FKBP) encoding rice (Oryza sativa L.) genes: genome-wide analysis, expression studies and evidence for their involvement in abiotic stress response. Mol Genet Genomics 279:371–383
Novatchkova M, Tomanov K, Hofmann K, Stuible HP, Bachmair A (2012) Update on sumoylation: defining core components of the plant SUMO conjugation system by phylogenetic comparison. New Phytol 195:23–31
Osakabe Y, Yamaguchi-Shinozaki K, Shinozaki K, Tran LP (2013) Sensing the environment: key roles of membrane-localized kinases in plant perception and response to abiotic stress. J Exp Bot 64:445–458
Park HC, Kim H, Koo SC, Park HJ, Cheong MS, Hong H, Baek D, Chung WS, Kim DH, Bressan RA, Lee SY, Bohnert HJ, Yun DJ (2010) Functional characterization of the SIZ/PIAS-type SUMO E3 ligases, OsSIZ1 and OsSIZ2 in rice. Plant Cell Environ 33:1923–1934
Park HJ, Kim WY, Park HC, Lee SY, Bohnert HJ, Yun DJ (2011) SUMO and SUMOylation in plants. Mol Cells 32:305–316
Peleg Z, Blumwald E (2011) Hormone balance and abiotic stress tolerance in crop plants. Curr Opin Plant Biol 14:290–295
Pratt WB, Toft DO (2003) Regulation of signaling protein function and trafficking by the hsp90/hsp70-based chaperone machinery. Exp Biol Med 228:111–133
Ray S, Agarwal P, Arora R, Kapoor S, Tyagi AK (2007) Expression analysis of calcium-dependent protein kinase gene family during reproductive development and abiotic stress conditions in rice (Oryza sativa L. ssp. indica). Mol Genet Genomics 278:493–505
Sakuma Y, Liu Q, Dubouzet JG, Abe H, Shinozaki K, Yamaguchi-Shinozaki K (2002) DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem Biophys Res Commun 290:998–1009
Sakuma Y, Maruyama K, Osakabe Y et al (2006) Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 18:1292–1309
Santner A, Calderon-Villalobos LIA, Estelle M (2009) Plant hormones are versatile chemical regulators of plant growth. Nat Chem Biol 5:301–308
Saracco SA, Miller MJ, Kurepa J, Vierstra RD (2007) Genetic analysis of SUMOylation in Arabidopsis: conjugation of SUMO1 and SUMO2 to nuclear proteins is essential. Plant Physiol 145:119–134
Sarkar NK, Kundnani P, Grover A (2012) Functional analysis of Hsp70 superfamily proteins of rice (Oryza sativa). Cell Stress Chaperon. doi:10.1007/s12192-012-0395-6
Sato Y, Yokoya S (2008) Enhanced tolerance to drought stress in transgenic rice plants overexpressing a small heat-shock protein, sHSP17.7. Plant Cell Rep 27:329–334
Shindo H, Suzuki R, Tsuchiya W, Taichi M, Nishiuchi Y, Yamazaki T (2012) PHD finger of the SUMO ligase Siz/PIAS family in rice reveals specific binding for methylated histone H3 at lysine 4 and arginine 2. Fed Eur Biochem Soc Lett 586:1783–1789
Shinozaki K, Yamaguchi-Shinozaki K (2000) Molecular responses to dehydration and low temperature: differences and cross-talk between two stress signaling pathways. Curr Opin Plant Biol 3:217–223
Shinozaki K, Yamaguchi-Shinozaki K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 6:410–417
Shiu SH, Karlowski WM, Pan R, Tzeng YH, Mayer KF, Li WH (2004) Comparative analysis of the receptor-like kinase family in Arabidopsis and rice. Plant Cell 16:1220–1234
Shiu MD, Zou C, Hanada K, Shiu SH (2009) Evolutionary history and stress regulation of plant receptor-like kinase/pelle genes. Plant Physiol 150:12–26
Small J, Vierstra RD (2004) The Ubiquitin 26S proteasome proteolytic pathway. Annu Rev Plant Biol 55:555–590
Sreenivasulu N, Harshavardhan VT, Govind G, Seiler C, Kohli A (2012) Contrapuntal role of ABA: does it mediate stress tolerance or plant growth retardation under long-term drought stress? Gene 15:265–273
Suzuki R, Shindo H, Tase A, Kikuchi Y, Shimizu M, Yamazaki T (2009) Solution structures and DNA binding properties of the N-terminal SAP domains of SUMO E3 ligases from Saccharamyces cerevisiae and Oryza sativa. Protein Struct Funct Bioinforma 75:336–347
Suzuki R, Tsuchiya W, Shindo H, Yamazaki T (2011) NMR assignments of ubiquitin fold domain (UFD) in SUMO-activating enzyme subunit 2 from rice. Biomol NMR Assign 5:245–248
Tan X, Calderon-Villalobos LI, Sharon M, Zheng C, Robinson CV, Estelle M, Zheng N (2007) Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446:640–645
Tang N, Zhang H, Li X, Xiao J, Xiong L (2012) Constitutive activation of transcription factor OsbZIP46 improves drought tolerance in rice. Plant Physiol 158:1755–1768
Tardieu F (2012) Any trait or trait-related allele can confer drought tolerance: just design the right drought scenario. J Exp Bot 63:25–31
Teng S, Luo H, Wang L (2010) Predicting protein sumoylation sites from sequence features. Amino Acids 43:447–455
Testerink C, Munnik T (2005) Phosphatidic acid: a multifunctional stress signaling lipid in plants. Trends Plant Sci 10:368–375
Thangasamy S, Guo CL, Chuang MH, Lai MH, Chen JC, Jauh GY (2011) Rice SIZ1, a SUMO E3 ligase, controls spikelet fertility through regulation of anther dehiscence. New Phytol 189:869–882
Thines B, Katsir L, Melotto M, Niu Y, Mandaokar A, Liu G, Nomura K, He SY, Howe GH, Browse J (2007) JAZ repressor proteins are targets of the SCF(COI1) complex during jasmonate signaling. Nature 448:661–665
Tian M, Xie Q (2013) Non-26S proteasome proteolytic role of ubiquitin in plant endocytosis and endosomal trafficking. J Integr Plant Biol 55:54–63
van den Burg HA, Takken FLW (2010) SUMO-, MAPK- and resistance protein-signaling converge at transcription complexes that regulate plant innate immunity. Plant Signal Behav 5:1597–1601
Ueguchi-Tanaka M, Nakajima M, Katoh E, Ohmiya H, Asano K, Saji S, Hongyu X, Ashikari M, Kitano H, Yamaguchi I, Matsuoka M (2007) Molecular interactions of a soluble gibberellin receptor, GID1, with a rice DELLA protein, SLR1, and gibberellins. Plant Cell 19:2140–2155
Vierling E (1991) The roles of heat shock proteins in plants. Annu Rev Plant Phys 42:579–620
Wang W, Vinocur B, Shoseyou O, Altman A (2004) Role of plant heat-shock proteins and molecular chaperones in the abiotic stress response. Trends Plant Sci 9:244–252
Wang HD, Makeen K, Yan Y, Cao Y, Sun SB, Xu GH (2011) OsSIZ1 regulates the vegetative growth and reproductive development in rice. Plant Mol Biol Rep 29:411–417
Wurzinger B, Mair A, Pfister B, Teige M (2011) Cross-talk of calcium-dependent protein kinase and MAP kinase signaling. Plant Signal Behav 6:8–12
Xie Q, Guo HS, Dallman G, Fang S, Weissman AM, Chua NH (2002) SINAT5 promotes ubiquitin-related degradation of NAC1 to attenuate auxin signals. Nature 419:167–170
Xue Y, Zhou F, Fu C, Xu Y, Yao X (2006) SUMOsp: a web server for sumoylation site prediction. Nucleic Acids Res 34:W254–W257
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Yamanouchi U, Yano M, Lin H, Ashikari M, Yamada K (2002) A rice spotted leaf gene, Spl17, encodes a heat stress transcription factor protein. Proc Natl Acad Sci USA 99:7530–7535
Yang A, Dai Xiaoyan, Zhang Wen-Hao (2012) A R2R3-type MYB gene, OsMYB2, is involved in salt, cold, and dehydration tolerance in rice. J Exp Bot 63:2541–2556
Yokotani N, Ichikawa T, Kondou Y, Matsui M, Hirochika H, Iwabuchi M, Oda K (2008) Expression of rice heat stress transcription factor OsHsfA2e enhances tolerance to environmental stresses in transgenic Arabidopsis. Planta 227:957–967
Yoo CY, Miura K, Jin JB, Lee J, Park HC, Salt DE, Yun DJ, Bressan RA, Hasegawa PM (2006) SIZ1 small ubiquitin-like modifier E3 ligase facilitates basal thermotolerance in Arabidopsis independent of salicylic acid. Plant Physiol 142:1548–1558
Yu H, Chen X, Hong YY, Wang Y, Xu P, Ke SD, Liu HY, Zhu JK, Oliver DJ, Xian CB (2008) Activated expression of an Arabidopsis HD-START protein confers drought tolerance with improved root system and reduced stomatal density. Plant Cell 20:1134–1151
Yu S, Liao F, Wang F, Wen W, Li J, Mei H, Luo L (2012) Identification of rice transcription factors associated with drought tolerance using the ecotilling method. PLoS One 7:e30765
Yuan S, Lin HH (2008) Role of salicylic acid in plant abiotic stress. Z Naturforsch C 63:313–320
Zhang S, Qi Y, Liu M, Yang C (2013) SUMO E3 ligase AtMMS21 regulates drought tolerance in Arabidopsis thaliana. J Integr Plant Biol 55:83–95
Zheng Y, Schumaker KS, Guo Y (2012) Sumoylation of transcription factor MYB30 by the small ubiquitin-like modifier E3 ligase SIZ1 mediates abscisic acid response in Arabidopsis thaliana. Proc Natl Acad Sci USA 109:12822–12827
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. Kumar.
A contribution to the Special Issue: Plant Hormone Signaling.
Rights and permissions
About this article
Cite this article
Raorane, M.L., Mutte, S.K., Varadarajan, A.R. et al. Protein SUMOylation and plant abiotic stress signaling: in silico case study of rice RLKs, heat-shock and Ca2+-binding proteins. Plant Cell Rep 32, 1053–1065 (2013). https://doi.org/10.1007/s00299-013-1452-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-013-1452-z