Abstract
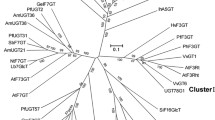
Secondary plant metabolites undergo several modification reactions, including glycosylation. Glycosylation, which is mediated by UDP-glycosyltransferase (UGT), plays a role in the storage of secondary metabolites and in defending plants against stress. In this study, we cloned one of the glycosyltransferases from rice, RUGT-5 resulting in 40–42% sequence homology with UGTs from other plants. RUGT-5 was functionally expressed as a glutathione S-transferase fusion protein in Escherichia coli and was then purified. Eight different flavonoids were used as tentative substrates. HPLC profiling of reaction products displayed at least two peaks. Glycosylation positions were located at the hydroxyl groups at C-3, C-7 or C-4′ flavonoid positions. The most efficient substrate was kaempferol, followed by apigenin, genistein and luteolin, in that order. According to in vitro results and the composition of rice flavonoids the in vivo substrate of RUGT-5 was predicted to be kaempferol or apigenin. To our knowledge, this is the first time that the function of a rice UGT has been characterized.



Similar content being viewed by others
Abbreviations
- IPTG::
-
Isopropyl-β-d-thiogalactopyran oside
- PCR::
-
Polymerase chain reaction
- UGT::
-
Uridine diphosphate dependent glycosyltransferase
References
Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408:796–815
Arend J, Waszecha H, Hefner T, Stockigt J (2001) Utilizing genetically engineered bacteria to produce plant-specific glucosides. Biotechnol Bioeng 76:125–131
Aziz AA, Edwards CAM, Lean MEJ, Crozier A (1998) Absorption and excretion of conjugated flavonols, including quercetin-4′-O-β-glucoside and isorhamnetin-4′-O-β-glucosides by human volunteers after the consumption of onions. Free Radic Res 29:257–269
Bowles D, Lsayenkova J, Lim E-K, Poppenberger B (2005) Glycosyltransferases: managers of small molecules. Curr Opin Plant Biol 8:254–263
Cornwell T, Cohick W, Raskin I (2004) Dietary phytoestrogens and health. Phytochemistry 65:995–1016
Hughes J, Hughes MA (1994) Multiple secondary plant product UDP-glucose glucosyltransferase genes expressed in cassava (Manihot esculenta Crantz) cotyledons. DNA Seq 5:41–49
Jones P, Vogt T (2001) Glycosyltransferases in secondary plant metabolism: tranquilizers and stimulant controllers. Planta 213:164–174
Kim BG, Kim SY, Song HS, Lee C, Hur HG, Kim SI, Ahn J-H (2003) Cloning and expression of the isoflavone synthase gene from Trifolium pratense. Mol Cells 15:301–306
Kim BG, Ko JH, Min SY, Ahn J-H (2005) Classification and expression profiling of putative R2R3 MYB genes in rice. Agric Chem Biotechnol 48:127–132
Kramer CM, Prata RTN, Willits MG, De Luca V, Steffens JC, Graser G (2003) Cloning and regiospecificity studies of two flavonoid glucosyltransferases from Allium cepa. Phytochemistry 64:1069–1076.
Li Y, Baldauf S, Lim E-K, Bowles DJ (2001) Phylogenetic analysis of the UDP-glycosyltransferase multigene family of Arabidopsis thaliana. J Biol Chem 276:4338–4343
Mackenzie PI, Owens IS, Burchell B, Bock KW, Bairoch A, Belanger A, Fournel-Gigleux S, Green M, Hum DW, Iyanagi T, Lancet D, Louisor P, Magdalow J, Chowdhury JR, Ritter JK, Schachter H, Tephly TR, Tipton KF, Nebert DW (1997) The UDP glycosyltransferase gene superfamily: recommended nomenclature update based on evolutionary divergence. Pharmacogenetics 7:255–269
Meβner B, Thulke O, Schäffner AR (2003) Arabidopsis glucosyltransferases with activities toward both endogenous and xenobiotic substrates. Planta 217:138–146
Ross J, Li Y, Lim E-K, Bowles D (2001) Higher plant glycosyltransferases. Genome Biol 2:3004.1–3004.6
Schwab W (2003) Metabolome diversity: too few genes, too many metabolites? Phytochemistry 62:837–849
Tohge T, Nishiyama Y, Hirai MY, Yano M, Nakajima J, Awazuhara M, Inoue E, Takahashi H, Goodenowe DB, Kitayama M, Noji M, Yamazaki M, Saito K (2005) Functional genomics by integrated analysis of metabolome and transcriptome of Arabiopsis plants over-expressing an MYB transcription factor. Plant J 42:218–235
Vogt T, Zimmermann E, Grimm R, Meyer M, Stack D (1997) Are the characteristics of betanidin glucosyltransferases from cell-suspension cultures of Dorotheanthus bellidiformis indicative of their phylogenetic relationship with flavonoid glucosyltransferases? Planta 203:349–361
Vogt T, Jones P (2000) Glycosyltransferases in plant natural product synthesis: characterization of a supergene family. Trends Plant Sci 5:380–386
Wink N (1999) Biochemistry of plant secondary metabolism. In: Annual plant reviews, vol 2. CRC Press, Boca Raton
Acknowledgements
This work was supported by a grant of Biogreen 21 Program, Rural Development Administration, Republic of Korea and in part by KRF2004-F00019 and the R&D Program for NBT Fusion Strategy of Advanced Technologies (Korea Ministry of Commerce, Industry and Energy).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by I. S. Chung
Rights and permissions
About this article
Cite this article
Ko, J.H., Kim, B.G., Hur, HG. et al. Molecular cloning, expression and characterization of a glycosyltransferase from rice. Plant Cell Rep 25, 741–746 (2006). https://doi.org/10.1007/s00299-006-0119-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-006-0119-4