Abstract
Significant progress has been made in understanding the functional organisation of the cell nucleus. Still many questions remain to be answered about the relationship between the spatial organisation of the nucleus and the regulation of the genome function. There are many conflicting data in the field making it very difficult to merge published results on mammalian cells into one model on subnuclear chromatin organisation. The fission yeast, Schizosaccharomyces pombe, over the last decades has emerged as a valuable model organism in understanding basic biological mechanisms, especially the cell cycle and chromosome biology. In this review we describe and compare the nuclear organisation in mammalian and fission yeast cells. We believe that fission yeast is a good tool to resolve at least some of the contradictions and unanswered questions concerning functional nuclear architecture, since S. pombe has chromosomes structurally similar to that of human. S. pombe also has the advantage over higher eukaryotes in that the genome can easily be manipulated via homologous recombination making it possible to integrate the tools needed for visualisation of chromosomes using live-cell microscopy. Classical genetic experiments can be used to elucidate what factors are involved in a certain mechanism. The knowledge we have gained during the last few years indicates similarities between the genome organisation in fission yeast and mammalian cells. We therefore propose the use of fission yeast for further advancement of our understanding of functional nuclear organisation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
More and more experimental data have highlighted the importance of the organisation of the cell nucleus in gene regulation and chromosome maintenance (de Wit and van Steensel 2009; Misteli and Soutoglou 2009). The cell nucleus is a highly organised organelle with specific subnuclear structures such as the nuclear membrane containing the nuclear pores. In addition, there are several non-membrane enclosed nuclear structures such as the nucleoli, splicing speckles, cajal bodies and gems that are thought to be self-organising (Matera et al. 2009). Different types of chromatin, transcriptionally active euchromatin and silent heterochromatin are kept separated in the nuclear space. However, understanding the rules that govern this type of organisation that in turn influence gene activity has turned out to be a challenging task. There are conflicting experimental results that has lead to several theories on the functional organisation of the human cell nucleus. More experiments are necessary to make a more complete picture of the nucleus. The use of model organisms such as the fission yeast, Schizosaccharomyces pombe, can be a good complement to studies of mammalian cell culture cells for increasing our understanding of the functional nuclear landscape. Fission yeast has the advantage of being easily manipulated genetically and moreover, has a similar chromosome structure as human cells. The basic organisation unit of chromatin, the nucleosome, is similar between human and yeast cells and in addition fission yeast and human cells shares homologues proteins, such as HP1 proteins, bound to the nucleosomes in the heterochromatic regions.
In this review article we go through the basic knowledge of chromatin; how various histone modifications make up different types of chromatin and how these different types are kept separated in the cell nucleus. We will also discuss the similarities and differences between chromatin structure in fission yeast and human cells. We will end with a comparison between the current knowledge about the organisation of the human and fission yeast nuclei. With that as a starting point, we will discuss future directions for the study of nuclear organisation using fission yeast as a model system.
Chromatin in eukaryotic cells
The basic organisation unit of chromatin, the core nucleosome, is made up of histone proteins with 147 bp of DNA wrapped around it with 1.7 turns. The nucleosome is an octamer with two of each of four different histones, namely: H2A, H2B, H3 and H4. In higher eukaryotes histone H1 binds to the core nucleosome at the DNA exit, stabilising the nucleoprotein complex. Linker DNA connects the core histones with each other. The nucleosome repeat length in human chromatin is around 185 bp, while S. pombe has a shorter repeat length of 154 bp (Jiang and Pugh 2009; Lantermann et al. 2010). The histone proteins are rich in basic amino acids such as lysine and arginine and thus interact well with the negatively charged DNA. How the chromatin is organised into higher order structures above the nucleosome level is still not clear, but it is known that the chromatin forms a 30 nm fibre, although the exact nature of this fibre still remains to be resolved (Dorigo et al. 2004; Robinson et al. 2006). Most researchers seem to agree that the 30 nm fibre forms loop structures, but whether or not other higher order structures are formed above the 30 nm fibre is not well understood and it is under much debate. In mammalian cells each chromosome has its own space within the cell nucleus forming the so-called “Chromosome Territories” (CTs). However, the degree of intermingling between these CTs is under debate (Dehghani et al. 2005; Cremer and Cremer 2006; Branco and Pombo 2007). Moreover, the CTs are oriented with the active genes towards the nuclear centre, and regions with low gene activity are found at the nuclear periphery (Fedorova and Zink 2008; Guelen et al. 2008). The genes that are associated with the nuclear periphery changes as cells go through differentiation, and there is a strong correlation between cells leaving the nuclear periphery and subsequent activation of a gene (Peric-Hupkes et al. 2010).
Histone modifications
Different types of nucleosomes with different kinds of histone modifications and histone variants are crucial for setting up distinct types of chromatin of which there are two basic types in the cell nucleus: euchromatin and heterochromatin. Euchromatin is gene dense and many genes within euchromatin are transcriptionally active. Heterochromatin, on the other hand, is gene poor, with a low transcription level and it also contains many repeated sequences. Histone marks commonly associated with euchromatin are: acetylated histones and methylation of lysine 4 on histone H3 (H3K4Me2/3). Heterochromatin, is characterised by: low acetylation levels and methylation of histone H3 at lysine 9 (H3K9Me2/3).
It is mainly the unstructured N-terminal tails that protrude from the core nucleosome that is post-translationally modified on many different amino acids through the addition, removal or replacement of for example methyl- or acetyl-groups. Different combinations of histone modification patterns have been suggested to form the so-called ‘histone code’ (Jenuwein and Allis 2001). How to read this code is not fully understood, but advances over the last decade have increased our understanding of how it is deciphered. The modifications create binding platforms for different chromatin modifying enzymes as well as proteins with structural functions in chromatin formation. In addition, acetylation of lysine residues causes a change in the charge of the histone protein, from basic to neutral, thus weakening the interaction between the histone and DNA within the nucleosome. This is thought to lead to a more open chromatin that would promote transcription. Importantly, these covalent histone modifications are reversible thereby making it possible to switch between different chromatin states, for example, between active euchromatin and transcriptionally inactive heterochromatin (Ekwall 2005; Mosammaparast and Shi 2010). Different histone modifications are created by distinct groups of enzymes. The addition of an acetyl-group to the histones is performed by histone acetyltransferases (HATs), while histone deacetylases (HDACs) removes the acetyl mark (Table 1). There are HATs and HDACs with broad substrate specificity as well as those with more narrow targets. For example, the HDAC Clr3 in S. pombe is specific for histone H3K14, while the HDAC Clr6 from the same organism has a broad activity, being able to deacetylase all tested acetyl marks (Bjerling et al. 2002; Wiren et al. 2005). The HDACs can be divided into three different classes. Class I and II have a similar enzymatic activity, while class III or Sirtuins (from the founding member Sir2 from budding yeast), uses a different chemistry being nicotinamide adenine dinucleotide (NAD+) dependent. Histone methyltransferases (HMTs) and histone demethylases (HDMs) all have a more narrow activity for one or two specific histone lysine residues. HMTs transfer one, two or three methyl groups onto a lysine residue via their SET domain. There are two different families of HDM enzymes that remove the methyl groups. The first utilises a flavin adenine dinucleotide (FAD) dependent oxidative reaction where formaldehyde is generated during demethylation (Shi et al. 2004). The founding member of this family is LSD1 that has the capacity to remove mono-or dimethyl-group bound to H3K4 or H3K9 (Shi et al. 2004; Metzger et al. 2005). In S. pombe there are two such enzymes while the budding yeast is devoid of this enzyme family (Table 1) (Lan et al. 2007). These two fission yeast enzymes are important for setting up the boundaries between different types of chromatin (see below) (Lan et al. 2007). Moreover, another family of HDM proteins called the Jumonji family also possessess demethylase activity (Balciunas and Ronne 2000; Tsukada et al. 2006). This family of HDMs can in addition to demethylation of mono- and dimetylated-lysines also demethylate trimethylated histone marks (Mosammaparast and Shi 2010). In addition to modifications of the canonical core histones, nucleosomes can also contain histone variants contributing to the nucleosome property (Talbert and Henikoff 2010).
DNA methylation
In mammalian cells as in many eukaryotic cells from many different kingdoms the DNA can be methylated. The methylation occurs on cytosine residues of DNA (5-methylcytosine). In mature mammalian cells the methylation is present where there is a cytosine followed by a guanosine, CpG, while in embryonic stem cells and other species also CpA, CpT or CpC can be methylated (Chen et al. 2008; Lister et al. 2009; Zemach et al. 2010). The extent of DNA methylation is also different between different species, 4% of all cytosines are methylated in human fibroblast cells, while in Drosophila melanogaster 0.1–0.05% of the cytosines are in the methylcytosine form (Gowher et al. 2000; Lister et al. 2009). DNA methylation is associated with transcriptional repression, but also occurs throughout the coding regions of active genes (Zemach et al. 2010). There is cross talk between DNA methylation and histone modifications (Chen et al. 2008). DNA methylation is associated with high levels of H3K9Me2/3 and low levels of H3K4Me2/3. Some fungi, like Neurospora crassa, have DNA methylation, while yeast lacks this modification (Antequera et al. 1984; Rountree and Selker 2010).
Heterochromatin
The H3K9Me2/3 methylation in heterochromatin is carried out by several types of SET domain proteins, among them are the founding member of HMTs, Su(var)3–9 in Drosophila, and its homologues SUVAR39H1 and SUVAR39H2 in human cells (Rea et al. 2000). In S. pombe the Su(var)3–9 homologue is named Clr4 (Table 1) (Ivanova et al. 1998). The H3K9Me2/3 mark is one of the best-understood histone modifications, since it creates a binding site for the structural component of heterochromatin, namely Heterochromatin Protein 1 (HP1). Both the H3K9Me2/3 mark and HP1 are well conserved among human, the fruitfly and fission yeast (Bannister et al. 2001). In fission yeast the HP1 protein is named Swi6 (Fig. 1). Budding yeast, Saccharomyces cerevisiae, on the other hand lacks H3K9Me2/3 as well as HP1 proteins (Table 1).
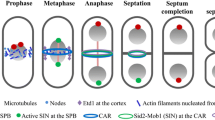
The mating-type region in fission yeast contains a region with heterochromatin. The nucleation of heterochromatin formation occurs at cenH (dotted box) a region with 96% identity to the centromeric dg and dh repeats (Grewal and Klar 1997). Transcription of the repeats (curved lines) recruits HMT Clr4 that methylates histone H3 on lysine 9 (Me). This in turn creates a binding site for HP1 proteins, mostly Swi6, but also Chp1 and Chp2. The heterochromatin spreads until it reaches a boundary element, IR-L (black arrow), centromere-proximal or IR-R (black arrow) centromere-distal of the mating-type region
The DNA in heterochromatin typically contains repeated sequences, for example the human pericentromeric heterochromatin is made up of tandem arrays of the 171 bp long α-satellite repeat (Bjerling and Ekwall 2002). In the fission yeast all the heterochromatic areas contain full or parts of the dg and dh repeats (Verdel and Moazed 2005). The environment in heterochromatin hampers transcription, although this is not absolute. In S. pombe a low level of transcription, especially during the S-phase of the cell cycle, is necessary for establishing the pericentromeric heterochromatin (Chen et al. 2008). Interestingly, it is Swi6, the structural component of heterochromatin, which recruits Jumonji protein Epe1 that in turn recruits RNA polII that transcribes the dg and dh repeats of the centromere (Zofall and Grewal 2006). Moreover, in the fruitfly, D. melanogaster, there are quite a large number of genes located in heterochromatin that remain active, and for many of these, the heterochromatic environment itself appears important for their activity (Fedorova and Zink 2008).
Facultative and constitutive heterochromatin
Heterochromatin can be further divided into two different subtypes, facultative and constitutive heterochromatin. Constitutive heterochromatin is stable and rarely changes epigenetic states. Facultative heterochromatin, on the other hand is unstable, can change expression states and will vary among different cell types. Constitutive heterochromatin is necessary in forming important chromosome structures. Around the centromeres, where the microtubuli binding kinetochore is formed, a function essential for pulling the sisterchromatids apart during cell division, the pericentromeric heterochromatin is found. Also the ends of the chromosomes, the telomeres, are accompanied by subtelomeric heterochromatin. Both human and fission yeast chromosomes contain pericentromeric heterochromatin, while this is lacking in the budding yeast. In S. pombe the pericentromeric heterochromatin is important for attracting cohesion that holds the two sister chromatids together and set up the bi-oriented kinetochores before entry into segregation during mitosis (Bernard et al. 2001; Sakuno et al. 2009). However, this function of the pericentromeric heterochromatin does not seem to be conserved in human cells (Serrano et al. 2009).
There are also additional parts of the genome that contain constitutive heterochromatin, for example S. pombe has a specialised region of chromosome 2 that contains the information required for mating (Fig. 1). Here the mat1 locus is expressed and contains either P or M information. The genetic information in the mat1 locus can switch due to exchange of information with one of the two silent storage cassettes: mat2-P containing the P cell information and mat3-M harbouring the M cell specificity. The cassettes, mat2-P and mat3-M, are found in a 16 kb heterochromatic region enriched in HP1 proteins and as a result these cassettes are transcriptionally repressed. In S. pombe the constitutive heterochromatin is found at the nuclear periphery. The centromeres are attached to the spindle pole body (SPB), and the mating-type region is closely associated with the centromeres (Chikashige et al. 1997; Alfredsson-Timmins et al. 2007) (Fig. 2). The telomeres are attached to the nuclear membrane at the opposite side of the cell nucleus, as compared to the SPB, in proximity to the nucleolus (Chikashige et al. 1997, 2009) (Fig. 2). This linear arrangement of the chromosomes in the interphase nucleus is different from the round CTs found in mammalian cells. Constitutive heterochromatin in human cells, in contrast to yeast, is found in the interior of the cell nucleus where it forms dense bodies (Fedorova and Zink 2008).
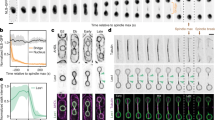
The organisation of the fission yeast nucleus. The spindle pole body (SPB) is inserted into the nuclear membrane and the centromeres (CEN) and the mating-type region (MAT) are found together at the SPB. The telomeres (TEL) cluster in 2–3 foci at the opposite end of the nucleus where the nucleolus with the ribosomal DNA (rDNA) is found. The telomeres are attached to the NM via the action of Clr3, Bqt3 and Bqt4 proteins. A cluster of nitrogen-repressed genes (Chr1) is found at the nuclear periphery together with the telomeres in a Clr3-dependent manner
Human cells also contain facultative heterochromatin, which is unstable and can change expression states. The regions that are contained in facultative heterochromatin can presumably differ between developmental stages and cell types. In human cells facultative heterochromatin is commonly found at the nuclear periphery (Fedorova and Zink 2008). Interestingly, S. pombe also contain limited amounts of facultative heterochromatin. A few genes, like ssm4 + and mei4 +, induced during meiosis are enriched in H3K9Me2/3 and HP1 protein (Cam et al. 2005). However, at least the ssm4 + gene does not associate with the other heterochromatic regions at the nuclear periphery but is found in the centre of the cell nucleus (Alfredsson-Timmins et al. 2009). To what degree is the heterochromatin responsible for setting up the organisation of the cell nucleus? Indeed, there are some studies indicating that heterochromatin is acting as an organiser of the chromatin. Regions of heterochromatin tend to aggregate. For example, the pericentromeric heterochromatin is found in one to a few clusters in the cell nucleus of organisms as diverse as yeast, Drosophila and human (Taddei et al. 2004). In Drosophila all the heterochromatic regions of the genome cluster together forming the chromocenter. Another striking example from the fruitfly is the somatic pairing of the homologous chromosomes leading to silencing of the brown gene by the dominant action of the brownD allele. The brownD allele contains a big insertion of heterochromatin that drags not only the brownD allele, but also the wild-type brown allele to the chromocenter and thus performs silencing in trans (Csink and Henikoff 1996).
Chromodomain proteins
An important component of heterochromatin is the HP1 proteins that are bound to nucleosomes with an H3K9Me2/3 mark via their N-terminal chromodomain (Bannister et al. 2001). Chromodomain containing proteins is a large group of chromatin-associated proteins with various functions, where the HP1 subfamily is distinguished by the presence of another conserved chromodomain-related domain at the C-terminal end known as the chromoshadow domain (Aasland and Stewart 1995). The HP1 proteins bind to each other via the chromoshadow domain (Cowieson et al. 2000). It is widely believed that further dimer interactions play a role in the spreading of heterochromatin. Moreover, the two chromoshadow domains together form a pit that in turn can interact with other proteins, such as transcriptional corepressors, chromatin remodelling factors and the lamin B receptor, that contain a hydrophobic P × V × L motif (Le Douarin et al. 1996; Ye et al. 1997; Smothers and Henikoff 2000; Thiru et al. 2004). In human cells there are three members of the HP1 family: HP1α (CBX5), HP1β (CBX1), and HP1γ (CBX3) (Lomberk et al. 2006). All three HP1 isoforms are concentrated at pericentromeric heterochromatin, but HP1β (CBX1), and HP1γ (CBX3) are also found in the euchromatic areas of the human genome (Minc et al. 2000; Vogel et al. 2006; Mateos-Langerak et al. 2007).
In S. pombe there are two members of the HP1 family: Swi6 and Chp2 (Thon and Verhein-Hansen 2000; Sadaie et al. 2004; Huisinga et al. 2006). In addition, fission yeast has two chromodomain protein outside the HP1 family that bind to H3K9Me2/3, namely Chp1 and the H3K9 methyltransferase, Clr4, (Zhang et al. 2008). Clr4 belongs to the Su(var)3–9 subfamily of histone methyltransferases, an enzyme family with chromo- and SET domains (Rea et al. 2000; Malik and Bhaumik 2010). The chromodomain proteins are concentrated at the heterochromatic regions of S. pombe (Sadaie et al. 2004; Zhang et al. 2008). Swi6 has functional similarities with HP1α in higher eukaryotes, being a structural component of heterochromatin, and is thus found in quite a large amounts (around 20,000 molecules/cell) (Sadaie et al. 2008). Chp1 is found together with Tas3 and Ago1 in the RITS complex. This complex is needed for the RNAi machinery, presumably by bringing Ago1 to the site where heterochromatin needs to be established or maintained (see below) (Verdel et al. 2004). Chp2 was recently found to associate with the SHREC complex that also contains: the HDAC Clr3, the Zn-finger protein Clr1, the nucleosome remodeller Mit1 and Clr2 (Sugiyama et al. 2007; Motamedi et al. 2008). However, it is unclear whether Chp2 is present in stoichiometric amounts with the other SHREC components, since in the first purification it was not detected (Sugiyama et al. 2007). Chp2 is found in limited amounts inside the cell nucleus, estimated to around 200–300 molecules/cell (Sadaie et al. 2008). Another component of the SHREC complex, Clr2, is also found in a very limited number in the S. pombe cell nucleus (Bjerling et al. 2004). ChIP-chip data for the SHREC complex clearly show a discrete association of Clr1 and Clr2 with only the heterochromatic regions while Clr3 binds throughout the genome (Sugiyama et al. 2007). These data indicate that Clr3 most likely performs additional functions involved in euchromatic gene regulation outside of the SHREC complex. Indeed, Clr3 regulates many genes (Hansen et al. 2005; Wiren et al. 2005).
RNAi
In S. pombe the RNAi machinery initiates and maintains the pericentromeric heterochromatin, while there are redundant pathways for heterochromatin establishment in the other heterochromatic parts of the genome. Transcription from repeated DNA sequences at the centromeres, mating-type region and telomeres especially during the S-phase of the cell cycle, triggers the RNAi machinery. This results in the recruitment of chromatin factors such as the H3K9 methyltransferase Clr4 and chromodomain proteins (Chen et al. 2008; Halic and Moazed 2010). The site of transcription and the subsequent accumulation of siRNA are the nucleation sites for heterochromatin formation. Once seeded, the heterochromatin continues to spread until it reaches a boundary element (Fig. 1) (Noma et al. 2004). In the centromeric regions, this pathway is essential for the initiation of heterochromatin while there are redundant establishment mechanisms in the mating-type region and the subtelomeric regions. In the mating-type region the recruitment of silencing factors can occur via Atf1/Pcr1 and in the subtelomeric regions Taz1 can initiate Swi6 binding (Jia et al. 2004; Kim et al. 2004; Kanoh et al. 2005). The RNAi machinery consists of the RNA-dependent RNA polymerase (Rdr1) that turns ssRNA into dsRNA. These are subsequently cleaved to 21–23 bp long siRNA by Dicer (Dcr1) and then loaded onto Argonaute (Ago1) in the RITS complex (Halic and Moazed 2010). The role of the small RNAs in setting up repressive chromatin structures have been described in fungi, plants, Drosophila and mammalian cell lines (Moazed 2009). Budding yeast, however, lacks the RNAi machinery; the genome does not contain homologues of Dicer or Argonaut (Table 1). Furthermore, it was recently shown that Dicer in S. pombe is localised to the nuclear periphery, coating the inside of the nuclear membrane although not colocalising with the heterochromatic regions (Emmerth et al. 2010). Clustering of the telomeres, but not attachment to the nuclear membrane is dependent on the RNAi machinery, indicating that the siRNA are needed for higher order chromatin structures to form in S. pombe (Hall et al. 2003). Interestingly, an unidentified RNA component is important for higher order chromatin structures to form in mouse cells (Maison et al. 2002). Moreover, several long non-coding RNA recruits remodelling factors to set up repressive chromatin structures, for example Kcnq1ot1 that is involved in imprinting of the Kcnq1 chromosomal domain (Pandey et al. 2008). In addition, non-coding RNAs are structural components of the X-chromosome dosage compensation mechanisms; Xist RNA is covering the inactivated X-chromosome in female mammals while roX RNA is coating the male X-chromosome in Drosophila (Ilik and Akhtar 2009; Senner and Brockdorff 2009).
Boundary elements
The border between euchromatin and heterochromatin is often very sharp due to boundary elements. These elements have the capacity to act as shields protecting genes from both spreading of nearby heterochromatin, as well as preventing the spread of euchromatin into heterochromatin regions disrupting important structures. Boundary elements frequently contain binding sites for different factors, for example tRNA transcription factors in yeast and mammals (Lunyak 2008) (see below). One example of a well-studied boundary element is in the chicken β–globin locus, where the 5’HS4 element is a sharp transition point between H3K4Me and H3K9Me histone modifications (Litt et al. 2001). In S. pombe, boundary elements surrounds both the pericentromeric and mating-type heterochromatin, and specific histone modifications are important for the boundary function. In fission yeast strains lacking the demethylase Lsd1 or the Jumonji domain protein Epe1, the pericentromeric heterochromatin spreads outside of the boundary elements (Zofall and Grewal 2006; Lan et al. 2007). The mechanism by which Epe1 aids in boundary formation is not fully understood, since the Jumonji domain in the protein lacks demethylase activity (Zofall and Grewal 2006). The boundary elements at the centromeres are clusters of tRNA genes and in two of the three centromeres there are also two inverted repeats surrounding the heterochromatin (Partridge et al. 2000; Cam et al. 2005). In the mating-type region (mat2/3) of fission yeast, the boundary elements consist of two inverted repeats, IR-L and IR-R, which separate the heterochromatin from the surrounding euchromatin (Fig. 1). These boundary elements are 100% identical over a 2.1 kb region. Upon deletion of these elements euchromatin can spread into the normally silent region and hence reporter genes inserted in this area are expressed (Thon et al. 2002). In a situation when deletion of IR-L and IR-R is combined with the simultaneous overexpression of the Swi6 protein, the heterochromatin spreads out into the normally euchromatic area (Noma et al. 2001). These two investigations demonstrate the importance of these boundary elements in separating the euchromatin from heterochromatin. Deletions of the boundary elements result in a blended chromatin at the edges of the mat2/3 region instead of the sharp borders seen in wild-type cells.
The boundary elements, IR-L and IR-R, might be a part of a redundant mechanism involved in tethering the mating-type region at the nuclear periphery. They contain a binding site for the polIII transcription factor TFIIIC. The boundary function is dependent on TFIIIC and this transcription factor does cluster together in foci at the nuclear membrane (Noma et al. 2006). In addition, a recent study also reveals that the genes transcribed by polIII: the tRNA and 5S rRNA genes are frequently located at the nuclear periphery together with heterochromatin (Iwasaki et al. 2010). However, there are redundant factors retaining the mating-type region at the nuclear periphery, since upon deletion of the boundary elements the mat2/3 region remains in the proximity of the nuclear membrane although slightly delocalised from its normal position next to the SPB (Alfredsson-Timmins et al. 2007). In contrast, heterochromatin formation is essential for positioning the mating-type region at the SPB, since upon deletion of the HMT Clr4, the mating-type region is completely delocalised and frequently found in the interior of the cell nucleus (Alfredsson-Timmins et al. 2007).
Functional genome organisation in budding yeast
Budding yeast lacks pericentromeric heterochromatin and has its own kind of stable subtelomeric chromatin. In budding yeast this resembles heterochromatin in human and fission yeast cells, but contains other components instead of HP1 proteins, namely the Sir proteins (Taddei et al. 2004). Budding yeast lacks the H3K9Me2/3 mark, but has a similar pattern of acetylated histones in actively transcribed chromatin and less acetylated histones in silent chromatin as in other eukaryotes. Budding yeast also has the histone methylation mark associated with active chromatin, H3K4Me2/3 (Taddei et al. 2004). The basic nuclear organisation of the chromatin is similar in fission and budding yeast with the centromeres attached to the SPB and the telomeres to the nuclear membrane at the opposite end of the nucleus where also the nucleolus is found. In budding yeast many induced genes move towards the nuclear pores in the nuclear membrane (Akhtar and Gasser 2007). The pores are thought to be transcriptionally active areas of the nuclear periphery whereas the spaces between the pores are repressed regions. A general movement of genes towards the nuclear pores upon activation is presumably restricted to smaller cell nuclei like the budding yeast cell. No studies indicate that such a movement would occur in human cells or in S. pombe. For fission yeast, though, we are not able to draw any final conclusions, since too few studies have been made on the movement of genes upon activation.
Functional genome organisation in human cells
The spatial organisation of the genome inside the cell nucleus is highly controlled, but the significance of this organisation and the rules for how it is set up and maintained is still poorly understood. Many studies indicate that in the human cell nuclei the chromosomes are confined within CTs where each chromosome has its own limited space, although this view has recently been challenged (see below). Chromosomes that are gene dense often have their CT in the centre of the nucleus, while gene poor chromosomes are found at the nuclear periphery. In addition, chromosomes at the vicinity of the nuclear periphery have their active genes towards the nuclear centre. In human cells a network of lamin proteins coat the inner nuclear membrane (Fedorova and Zink 2008). In both human cells and in D. melanogaster the chromatin at the nuclear periphery, the so-called “lamin associated domains (LADs)”, are not necessarily heterochromatic but is distinguished by: low gene density, low transcriptional activation and having low acetylation levels of the histones (Guelen et al. 2008; de Wit and van Steensel 2009). Consistent with this finding, the human HDAC3 is attached to both chromatin and the nuclear envelope by interaction with LAP2β (Somech et al. 2005). There are also several genes in human cells that are moved away from the nuclear periphery towards the nuclear centre upon induction (Kosak et al. 2002; Zink et al. 2004; Morey et al. 2008). Recently a genome-wide mapping of the changes in association with the nuclear membrane during differentiation for mouse stem cells to astrocytes was performed (Peric-Hupkes et al. 2010). This study revealed a high correlation between genes that were activated and moved away from the nuclear membrane. New imaging techniques have challenged the view of chromosomes being confined to a strict territory. For example, in mouse erythroid cells genes loop out of the CT to transcription factories shared by genes from different chromosomes in the interchromatin space (Osborne et al. 2004). The human cell nucleus can either be viewed as quite rigid with everything in its special place or as a highly dynamic entity with frequent contact between different regions within the nuclear space (Cremer and Cremer 2006; Branco and Pombo 2007). Currently, there are several different models on how chromosomes are organised inside the human cell nuclei and they are difficult to unify. In the oldest model, active transcription occurs in specialised interchromosome compartments named interchromosome domains (ICD) (Zirbel et al. 1993) (Fig. 3a). After transcription had been detected within CTs the chromosome territory-interchromatin compartment (CT-IC) model was launched (Fig. 3b) (Cremer and Cremer 2001). Here, the interchromatin compartment, were transcription is taking place, is described as a network between CTs but also within CT. These two early models on nuclear organisation also include the formation of higher order chromatin structures. Later on, using electron spectroscopic imaging (ESI), no higher order structures above the 30 nm structure were detected, leading to the proposal of the lattice model of chromatin organisation (Dehghani et al. 2005). This model describes the interphase nucleus as a meshwork of chromatin fibres of 10 and 30 nm with an extensive interchromosomal space (Fig. 3c). Moreover, techniques such as the genome-wide chromosome conformational capture (4C) and cryo-FISH have provided evidence for a greater degree of intermingling of chromosomes than it was previously envisioned (Branco and Pombo 2006; Simonis et al. 2006; Zhao et al. 2006). The intermingling of chromosomes and the colocalisation of these intermingling-sites with active transcription has lead to a fourth interchromosome network (ICN) model (Branco and Pombo 2007). The ICN model proposes that attachment of chromatin to different nuclear landmarks such as the nucleolus, and the peripheral lamins, as well as interchromosome interactions presumably via transcription factories. These interactions will lead to networks shaping the functional organisation of the mammalian cell nucleus (Fig. 3d).
The human cell nucleus according to different models. a The inter chromosome domain (ICD) model. The chromosomes are organised in distinct chromosome territories (grey) with higher order chromatin structures (black structures). Transcription by RNA polII (black dots) occurs in the interchromatin space. b The chromosome territory-interchromatin compartment (CT-IC) model. Also here the chromosomes are restricted to chromosome territories (grey) with higher order chromatin structures (black structures) with transcription by RNA polII (black dots) in the interchromatin space as well as in channels within the chromosome territories. c The lattice model with a mesh of chromatin fibres of 10 or 30 nm with extensive overlap between the chromosome territories (grey) and no distinct interchromosomal space. RNA polII (black dots) can freely diffuse in the chromatin mesh. d The ICN model where functional interaction between chromosomes (grey), probably by transcription factories (white oval marked TF), and attachment to fixed structures in the nucleus such as the nucleolus and the nuclear envelope forms the nuclear organisation
Functional genome organisation in fission yeast
Labelling of two of the three chromosomes by chromosome paint in S. pombe revealed distinct territories for the chromosomes, much like what is seen in human cells (Fig. 2) (Scherthan et al. 1994). As mentioned above the heterochromatic regions in fission yeast localise in foci at the nuclear periphery, frequently together with the genes transcribed by polIII: the tRNA and 5S rRNA genes (Chikashige et al. 1997; Alfredsson-Timmins et al. 2007; Iwasaki et al. 2010) (Fig. 2). The centromeres are attached to the SPB, and the mating-type region is closely associated with the centromeres (Chikashige et al. 1997; Alfredsson-Timmins et al. 2007) (Fig. 2). The telomeres are attached to the nuclear membrane via Bqt3 and Bqt4 at the opposite side of the cell nucleus as compared to the SPB in proximity to the nucleolus (Chikashige et al. 2009) (Fig. 2). A chromosomal arrangement like this, where the chromosomes are linearly packed side by side in the interphase nucleus, is called the Rabl formation (Cremer and Cremer 2006). In cells lacking Bqt4 the telomeres are slightly detached from the nuclear periphery without affecting the chromatin at the telomeres (Chikashige et al. 2009). Moreover, the Tf2 retrotransposon elements cluster in 1–3 foci in the cell nucleus and this clustering is dependent on CENP-B proteins (Cam et al. 2008).
During normal growth, gene clusters that are repressed by nitrogen and thus having low-acetylation levels of the histones are found at the nuclear periphery (Alfredsson-Timmins et al. 2009) (Fig. 2). This localisation is dependent on the HDAC Clr3, since upon deletion of Clr3 the clusters move to the centre of the cell nucleus. Moreover, it is not only the enzymatic activity of Clr3 that is responsible for the attachment to the nuclear membrane, since an enzymatically inactive protein has an intermediate phenotype between wild type and clr3Δ (Alfredsson-Timmins et al. 2009). Interestingly, the HDAC Clr3 is highly concentrated at the nuclear periphery (Bjerling et al. 2002) which leads to the speculation that Clr3 might also have a structural role in keeping these gene clusters at the periphery in addition to its enzymatic role. It is noticeable that these gene clusters have properties similar to the LADs identified in human cells (de Wit and van Steensel 2009). There are no lamins in S. pombe, but there might be integral proteins of the nuclear membrane with similar functions in human and fission yeast cells. The gene clusters that are repressed by nitrogen and reside at the nuclear periphery shift in localisation away from the periphery towards the nucleus interior upon induction by nitrogen starvation (Alfredsson-Timmins et al. 2009). Moreover, the shift in localisation is accompanied by drastic changes in the chromatin with massive nucleosome loss (Kristell et al. 2010). One interesting possibility is that there might be other gene clusters that also undergo a subnuclear repositioning during activation. In mammalian cells, movements away from the nuclear periphery during gene activation has been detected (Peric-Hupkes et al. 2010). Another interesting aspect to consider is; what are the driving forces behind the movement of genes during transcriptional activation. In several cases, transcription itself is the driving force, since treating the cells with transcriptional inhibitors prevent the movement upon induction, as seen for example for both the CTCF locus in human cells and the genes induced by nitrogen starvation in S. pombe (Zink et al. 2004; Alfredsson-Timmins et al. 2009). Since actin and myosin has been implicated in transcription, it would be fascinating to investigate whether the repositioning during transcriptional activation is motor driven (Gieni and Hendzel 2009). Perhaps, the actual movement of one gene locus that becomes activated contributes to subsequent activation of other genes nearby.
Conclusion
The past decade has provided us with many insights into the functional organisation of the 3D nucleus, but many questions still remain unsolved. For instance, how rigid is the organisation of the genome within the cell nucleus? Does everything have their own fixed place where the chromosomes are strictly confined within their own territories, or is the nucleus a highly dynamic organelle with contact between many different structures and factors within the nuclear space? Is the content of the genome packaged into facultative heterochromatin changing throughout differentiation as genes are selectively turned on or off, or is the movement of chromatin a driving force in promoting cell-type identity? Perhaps, it is a combination of both? Furthermore, what function do the heterochromatic areas play in setting up cell structures that influence gene expression? Is the physical movement during gene activation driven by motor proteins or are the genes just stochastically shifted to another place in the nucleus and stay there due to high concentrations of transcription components? Some of these questions can be answered by using the genetic power of the fission yeast system. For example, questions relating to how heterochromatin organisation influences genes that are tethered to a particular compartment of the cell nucleus can be closely investigated in the S. pombe system. Indeed, also the influence of facultative heterochromatin on gene regulation can be studied in the fission yeast system. For example, is the facultative heterochromatin aggregating inside the nucleus in foci, and is this colocalisation lost upon activation of the genes during meiosis and does this then have an influence on the expression of nearby genes? Other questions are; how the repressed regions: the LADs in human cells and the gene clusters induced by nitrogen starvation in fission yeast are kept at the nuclear periphery, how do they dissociate upon transcriptional activation? Is this movement of activated genes affecting the activity of other genes and thus is a part of inducing a whole transcriptional programme? Is the movement necessary for changes in chromatin and transcriptional activation?
References
Aasland R, Stewart AF (1995) The chromo shadow domain, a second chromo domain in heterochromatin-binding protein 1, HP1. Nucleic Acids Res 23:3168–3173
Akhtar A, Gasser SM (2007) The nuclear envelope and transcriptional control. Nat Rev Genet 8:507–517
Alfredsson-Timmins J, Henningson F, Bjerling P (2007) The Clr4 methyltransferase determines the subnuclear localization of the mating-type region in fission yeast. J Cell Sci 120:1935–1943
Alfredsson-Timmins J, Kristell C, Henningson F, Lyckman S, Bjerling P (2009) Reorganization of chromatin is an early response to nitrogen starvation in Schizosaccharomyces pombe. Chromosoma 118:99–112
Antequera F, Tamame M, Villanueva JR, Santos T (1984) DNA methylation in the fungi. J Biol Chem 259:8033–8036
Balciunas D, Ronne H (2000) Evidence of domain swapping within the jumonji family of transcription factors. Trends Biochem Sci 25:274–276
Bannister AJ, Zegerman P, Partridge JF, Miska EA, Thomas JO, Allshire RC, Kouzarides T (2001) Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410:120–124
Bernard P, Maure JF, Partridge JF, Genier S, Javerzat JP, Allshire RC (2001) Requirement of heterochromatin for cohesion at centromeres. Science 294:2539–2542
Bjerling P, Ekwall K (2002) Centromere domain organization and histone modifications. Braz J Med Biol Res 35:499–507
Bjerling P, Silverstein RA, Thon G, Caudy A, Grewal S, Ekwall K (2002) Functional divergence between histone deacetylases in fission yeast by distinct cellular localization and in vivo specificity. Mol Cell Biol 22:2170–2181
Bjerling P, Ekwall K, Egel R, Thon G (2004) A novel type of silencing factor, Clr2, is necessary for transcriptional silencing at various chromosomal locations in the fission yeast Schizosaccharomyces pombe. Nucleic Acids Res 32:4421–4428
Branco MR, Pombo A (2006) Intermingling of chromosome territories in interphase suggests role in translocations and transcription-dependent associations. PLoS Biol 4:e138
Branco MR, Pombo A (2007) Chromosome organization: new facts, new models. Trends Cell Biol 17:127–134
Cam HP, Sugiyama T, Chen ES, Chen X, FitzGerald PC, Grewal SI (2005) Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nat Genet 37:809–819
Cam HP, Noma K, Ebina H, Levin HL, Grewal SI (2008) Host genome surveillance for retrotransposons by transposon-derived proteins. Nature 451:431–436
Chen ES, Zhang K, Nicolas E, Cam HP, Zofall M, Grewal SI (2008) Cell cycle control of centromeric repeat transcription and heterochromatin assembly. Nature 451:734–737
Chikashige Y, Ding DQ, Imai Y, Yamamoto M, Haraguchi T, Hiraoka Y (1997) Meiotic nuclear reorganization: switching the position of centromeres and telomeres in the fission yeast Schizosaccharomyces pombe. EMBO J 16:193–202
Chikashige Y, Yamane M, Okamasa K, Tsutsumi C, Kojidani T, Sato M, Haraguchi T, Hiraoka Y (2009) Membrane proteins Bqt3 and -4 anchor telomeres to the nuclear envelope to ensure chromosomal bouquet formation. J Cell Biol 187:413–427
Cowieson NP, Partridge JF, Allshire RC, McLaughlin PJ (2000) Dimerisation of a chromo shadow domain and distinctions from the chromodomain as revealed by structural analysis. Curr Biol 10:517–525
Cremer T, Cremer C (2001) Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat Rev Genet 2:292–301
Cremer T, Cremer C (2006) Rise, fall, resurrection of chromosome territories: a historical perspective. Part II. Fall, resurrection of chromosome territories during the 1950 s to 1980 s. Part III. Chromosome territories, the functional nuclear architecture: experiments, models from the (1990) to the present. Eur J Histochem 50:223–272
Csink AK, Henikoff S (1996) Genetic modification of heterochromatic association and nuclear organization in Drosophila. Nature 381:529–531
de Wit E, van Steensel B (2009) Chromatin domains in higher eukaryotes: insights from genome-wide mapping studies. Chromosoma 118:25–36
Dehghani H, Dellaire G, Bazett-Jones DP (2005) Organization of chromatin in the interphase mammalian cell. Micron 36:95–108
Dorigo B, Schalch T, Kulangara A, Duda S, Schroeder RR, Richmond TJ (2004) Nucleosome arrays reveal the two-start organization of the chromatin fiber. Science 306:1571–1573
Ekwall K (2005) Genome-wide analysis of HDAC function. Trends Genet 21:608–615
Emmerth S, Schober H, Gaidatzis D, Roloff T, Jacobeit K, Buhler M (2010) Nuclear retention of fission yeast dicer is a prerequisite for RNAi-mediated heterochromatin assembly. Dev Cell 18:102–113
Fedorova E, Zink D (2008) Nuclear architecture and gene regulation. Biochim Biophys Acta 1783:2174–2184
Gieni RS, Hendzel MJ (2009) Actin dynamics and functions in the interphase nucleus: moving toward an understanding of nuclear polymeric actin. Biochem Cell Biol 87:283–306
Gowher H, Leismann O, Jeltsch A (2000) DNA of Drosophila melanogaster contains 5-methylcytosine. EMBO J 19:6918–6923
Grewal SI, Klar AJ (1997) A recombinationally repressed region between mat2 and mat3 loci shares homology to centromeric repeats and regulates directionality of mating-type switching in fission yeast. Genetics 146:1221–1238
Guelen L, Pagie L, Brasset E, Meuleman W, Faza MB, Talhout W, Eussen BH, de Klein A, Wessels L, de Laat W, van Steensel B (2008) Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453:948–951
Halic M, Moazed D (2010) Dicer-independent primal RNAs trigger RNAi and heterochromatin formation. Cell 140:504–516
Hall IM, Noma K, Grewal SI (2003) RNA interference machinery regulates chromosome dynamics during mitosis and meiosis in fission yeast. Proc Natl Acad Sci USA 100:193–198
Hansen KR, Burns G, Mata J, Volpe TA, Martienssen RA, Bahler J, Thon G (2005) Global effects on gene expression in fission yeast by silencing and RNA interference machineries. Mol Cell Biol 25:590–601
Huisinga KL, Brower-Toland B, Elgin SC (2006) The contradictory definitions of heterochromatin: transcription and silencing. Chromosoma 115:110–122
Ilik I, Akhtar A (2009) roX RNAs: non-coding regulators of the male X chromosome in flies. RNA Biol 6:113–121
Ivanova AV, Bonaduce MJ, Ivanov SV, Klar AJ (1998) The chromo and SET domains of the Clr4 protein are essential for silencing in fission yeast. Nat Genet 19:192–195
Iwasaki O, Tanaka A, Tanizawa H, Grewal SI, Noma K (2010) Centromeric localization of dispersed Pol III genes in fission yeast. Mol Biol Cell 21:254–265
Jenuwein T, Allis CD (2001) Translating the histone code. Science 293:1074–1080
Jia S, Noma K, Grewal SI (2004) RNAi-independent heterochromatin nucleation by the stress-activated ATF/CREB family proteins. Science 304:1971–1976
Jiang C, Pugh BF (2009) Nucleosome positioning and gene regulation: advances through genomics. Nat Rev Genet 10:161–172
Kanoh J, Sadaie M, Urano T, Ishikawa F (2005) Telomere binding protein Taz1 establishes Swi6 heterochromatin independently of RNAi at telomeres. Curr Biol 15:1808–1819
Kim HS, Choi ES, Shin JA, Jang YK, Park SD (2004) Regulation of Swi6/HP1-dependent heterochromatin assembly by cooperation of components of the mitogen-activated protein kinase pathway and a histone deacetylase Clr6. J Biol Chem 279:42850–42859
Klose RJ, Kallin EM, Zhang Y (2006) JmjC-domain-containing proteins and histone demethylation. Nat Rev Genet 7:715–727
Kosak ST, Skok JA, Medina KL, Riblet R, Le Beau MM, Fisher AG, Singh H (2002) Subnuclear compartmentalization of immunoglobulin loci during lymphocyte development. Science 296:158–162
Kristell C, Orzechowski Westholm J, Olsson I, Ronne H, Komorowski J, Bjerling P (2010) Nitrogen depletion in the fission yeast Schizosaccharomyces pombe causes nucleosome loss in both promoters and coding regions of activated genes. Genome Res 20:361–371
Lan F, Zaratiegui M, Villen J, Vaughn MW, Verdel A, Huarte M, Shi Y, Gygi SP, Moazed D, Martienssen RA, Shi Y (2007) S. pombe LSD1 homologs regulate heterochromatin propagation and euchromatic gene transcription. Mol Cell 26:89–101
Lantermann AB, Straub T, Stralfors A, Yuan GC, Ekwall K, Korber P (2010) Schizosaccharomyces pombe genome-wide nucleosome mapping reveals positioning mechanisms distinct from those of Saccharomyces cerevisiae. Nat Struct Mol Biol 17:251–257
Le Douarin B, Nielsen AL, Garnier JM, Ichinose H, Jeanmougin F, Losson R, Chambon P (1996) A possible involvement of TIF1 alpha and TIF1 beta in the epigenetic control of transcription by nuclear receptors. EMBO J 15:6701–6715
Lister R, Pelizzola M, Dowen RH, Hawkins RD, Hon G, Tonti-Filippini J, Nery JR, Lee L, Ye Z, Ngo QM, Edsall L, Antosiewicz-Bourget J, Stewart R, Ruotti V, Millar AH, Thomson JA, Ren B, Ecker JR (2009) Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462:315–322
Litt MD, Simpson M, Gaszner M, Allis CD, Felsenfeld G (2001) Correlation between histone lysine methylation and developmental changes at the chicken beta-globin locus. Science 293:2453–2455
Lomberk G, Wallrath L, Urrutia R (2006) The heterochromatin protein 1 family. Genome Biol 7:228
Lunyak VV (2008) Boundaries. Boundaries…Boundaries??? Curr Opin Cell Biol 20:281–287
Maison C, Bailly D, Peters AH, Quivy JP, Roche D, Taddei A, Lachner M, Jenuwein T, Almouzni G (2002) Higher-order structure in pericentric heterochromatin involves a distinct pattern of histone modification and an RNA component. Nat Genet 30:329–334
Malik S, Bhaumik SR (2010) Mixed lineage leukemia: histone H3 lysine 4 methyltransferases from yeast to human. FEBS J 277:1805–1821
Mateos-Langerak J, Brink MC, Luijsterburg MS, van der Kraan I, van Driel R, Verschure PJ (2007) Pericentromeric heterochromatin domains are maintained without accumulation of HP1. Mol Biol Cell 18:1464–1471
Matera AG, Izaguire-Sierra M, Praveen K, Rajendra TK (2009) Nuclear bodies: random aggregates of sticky proteins or crucibles of macromolecular assembly? Dev Cell 17:639–647
Metzger E, Wissmann M, Yin N, Muller JM, Schneider R, Peters AH, Gunther T, Buettner R, Schule R (2005) LSD1 demethylates repressive histone marks to promote androgen-receptor-dependent transcription. Nature 437:436–439
Minc E, Courvalin JC, Buendia B (2000) HP1gamma associates with euchromatin and heterochromatin in mammalian nuclei and chromosomes. Cytogenet Cell Genet 90:279–284
Misteli T, Soutoglou E (2009) The emerging role of nuclear architecture in DNA repair and genome maintenance. Nat Rev Mol Cell Biol 10:243–254
Moazed D (2009) Small RNAs in transcriptional gene silencing and genome defence. Nature 457:413–420
Morey C, Da Silva NR, Kmita M, Duboule D, Bickmore WA (2008) Ectopic nuclear reorganisation driven by a Hoxb1 transgene transposed into Hoxd. J Cell Sci 121:571–577
Mosammaparast N, Shi Y (2010) Reversal of histone methylation: biochemical and molecular mechanisms of histone demethylases. Annu Rev Biochem 79:155–179
Motamedi MR, Hong EJ, Li X, Gerber S, Denison C, Gygi S, Moazed D (2008) HP1 proteins form distinct complexes and mediate heterochromatic gene silencing by nonoverlapping mechanisms. Mol Cell 32:778–790
Noma K, Allis CD, Grewal SI (2001) Transitions in distinct histone H3 methylation patterns at the heterochromatin domain boundaries. Science 293:1150–1155
Noma K, Sugiyama T, Cam H, Verdel A, Zofall M, Jia S, Moazed D, Grewal SI (2004) RITS acts in cis to promote RNA interference-mediated transcriptional and post-transcriptional silencing. Nat Genet 36:1174–1180
Noma K, Cam HP, Maraia RJ, Grewal SI (2006) A role for TFIIIC transcription factor complex in genome organization. Cell 125:859–872
Osborne CS, Chakalova L, Brown KE, Carter D, Horton A, Debrand E, Goyenechea B, Mitchell JA, Lopes S, Reik W, Fraser P (2004) Active genes dynamically colocalize to shared sites of ongoing transcription. Nat Genet 36:1065–1071
Pandey RR, Mondal T, Mohammad F, Enroth S, Redrup L, Komorowski J, Nagano T, Mancini-Dinardo D, Kanduri C (2008) Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol Cell 32:232–246
Partridge JF, Borgstrom B, Allshire RC (2000) Distinct protein interaction domains and protein spreading in a complex centromere. Genes Dev 14:783–791
Peric-Hupkes D, Meuleman W, Pagie L, Bruggeman SW, Solovei I, Brugman W, Graf S, Flicek P, Kerkhoven RM, van Lohuizen M, Reinders M, Wessels L, van Steensel B (2010) Molecular maps of the reorganization of genome-nuclear lamina interactions during differentiation. Mol Cell 38:603–613
Rea S, Eisenhaber F, O’Carroll D, Strahl BD, Sun ZW, Schmid M, Opravil S, Mechtler K, Ponting CP, Allis CD, Jenuwein T (2000) Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406:593–599
Robinson PJ, Fairall L, Huynh VA, Rhodes D (2006) EM measurements define the dimensions of the “30-nm” chromatin fiber: evidence for a compact, interdigitated structure. Proc Natl Acad Sci USA 103:6506–6511
Rountree MR, Selker EU (2010) DNA methylation and the formation of heterochromatin in Neurospora crassa. Heredity 105:38–44
Sadaie M, Iida T, Urano T, Nakayama J (2004) A chromodomain protein, Chp1, is required for the establishment of heterochromatin in fission yeast. EMBO J 23:3825–3835
Sadaie M, Kawaguchi R, Ohtani Y, Arisaka F, Tanaka K, Shirahige K, Nakayama J (2008) Balance between distinct HP1 family proteins controls heterochromatin assembly in fission yeast. Mol Cell Biol 28:6973–6988
Sakuno T, Tada K, Watanabe Y (2009) Kinetochore geometry defined by cohesion within the centromere. Nature 458:852–858
Scherthan H, Bahler J, Kohli J (1994) Dynamics of chromosome organization and pairing during meiotic prophase in fission yeast. J Cell Biol 127:273–285
Senner CE, Brockdorff N (2009) Xist gene regulation at the onset of X inactivation. Curr Opin Genet Dev 19:122–126
Serrano A, Rodriguez-Corsino M, Losada A (2009) Heterochromatin protein 1 (HP1) proteins do not drive pericentromeric cohesin enrichment in human cells. PLoS One 4:e5118
Shi Y, Lan F, Matson C, Mulligan P, Whetstine JR, Cole PA, Casero RA (2004) Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119:941–953
Simonis M, Klous P, Splinter E, Moshkin Y, Willemsen R, de Wit E, van Steensel B, de Laat W (2006) Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C). Nat Genet 38:1348–1354
Smothers JF, Henikoff S (2000) The HP1 chromo shadow domain binds a consensus peptide pentamer. Curr Biol 10:27–30
Somech R, Shaklai S, Geller O, Amariglio N, Simon AJ, Rechavi G, Gal-Yam EN (2005) The nuclear-envelope protein and transcriptional repressor LAP2beta interacts with HDAC3 at the nuclear periphery, and induces histone H4 deacetylation. J Cell Sci 118:4017–4025
Sugiyama T, Cam HP, Sugiyama R, Noma K, Zofall M, Kobayashi R, Grewal SI (2007) SHREC, an effector complex for heterochromatic transcriptional silencing. Cell 128:491–504
Taddei A, Hediger F, Neumann FR, Gasser SM (2004) The function of nuclear architecture: a genetic approach. Annu Rev Genet 38:305–345
Talbert PB, Henikoff S (2010) Histone variants–ancient wrap artists of the epigenome. Nat Rev Mol Cell Biol 11:264–275
Thiru A, Nietlispach D, Mott HR, Okuwaki M, Lyon D, Nielsen PR, Hirshberg M, Verreault A, Murzina NV, Laue ED (2004) Structural basis of HP1/PXVXL motif peptide interactions and HP1 localisation to heterochromatin. EMBO J 23:489–499
Thon G, Verhein-Hansen J (2000) Four chromo-domain proteins of Schizosaccharomyces pombe differentially repress transcription at various chromosomal locations. Genetics 155:551–568
Thon G, Bjerling P, Bunner CM, Verhein-Hansen J (2002) Expression-state boundaries in the mating-type region of fission yeast. Genetics 161:611–622
Tsukada Y, Fang J, Erdjument-Bromage H, Warren ME, Borchers CH, Tempst P, Zhang Y (2006) Histone demethylation by a family of JmjC domain-containing proteins. Nature 439:811–816
Verdel A, Moazed D (2005) RNAi-directed assembly of heterochromatin in fission yeast. FEBS Lett 579:5872–5878
Verdel A, Jia S, Gerber S, Sugiyama T, Gygi S, Grewal SI, Moazed D (2004) RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303:672–676
Vogel MJ, Guelen L, de Wit E, Peric-Hupkes D, Loden M, Talhout W, Feenstra M, Abbas B, Classen AK, van Steensel B (2006) Human heterochromatin proteins form large domains containing KRAB-ZNF genes. Genome Res 16:1493–1504
Wiren M, Silverstein RA, Sinha I, Walfridsson J, Lee HM, Laurenson P, Pillus L, Robyr D, Grunstein M, Ekwall K (2005) Genomewide analysis of nucleosome density histone acetylation and HDAC function in fission yeast. EMBO J 24:2906–2918
Ye Q, Callebaut I, Pezhman A, Courvalin JC, Worman HJ (1997) Domain-specific interactions of human HP1-type chromodomain proteins and inner nuclear membrane protein LBR. J Biol Chem 272:14983–14989
Zemach A, McDaniel IE, Silva P, Zilberman D (2010) Genome-wide evolutionary analysis of eukaryotic DNA methylation. Science 328:916–919
Zhang K, Mosch K, Fischle W, Grewal SI (2008) Roles of the Clr4 methyltransferase complex in nucleation, spreading and maintenance of heterochromatin. Nat Struct Mol Biol 15:381–388
Zhao Z, Tavoosidana G, Sjolinder M, Gondor A, Mariano P, Wang S, Kanduri C, Lezcano M, Sandhu KS, Singh U, Pant V, Tiwari V, Kurukuti S, Ohlsson R (2006) Circular chromosome conformation capture (4C) uncovers extensive networks of epigenetically regulated intra- and interchromosomal interactions. Nat Genet 38:1341–1347
Zink D, Amaral MD, Englmann A, Lang S, Clarke LA, Rudolph C, Alt F, Luther K, Braz C, Sadoni N, Rosenecker J, Schindelhauer D (2004) Transcription-dependent spatial arrangements of CFTR and adjacent genes in human cell nuclei. J Cell Biol 166:815–825
Zirbel RM, Mathieu UR, Kurz A, Cremer T, Lichter P (1993) Evidence for a nuclear compartment of transcription and splicing located at chromosome domain boundaries. Chromosome Res 1:93–106
Zofall M, Grewal SI (2006) Swi6/HP1 recruits a JmjC domain protein to facilitate transcription of heterochromatic repeats. Mol Cell 22:681–692
Acknowledgments
We thank Dr Jun-ichi Nakayama as well as past and present members of the Bjerling group for reading and commenting on this manuscript. We acknowledge support from the Göran Gustafssons Foundation, the Swedish Cancer Society and Carl Tryggers Foundation.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. Sunnerhagen.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Olsson, I., Bjerling, P. Advancing our understanding of functional genome organisation through studies in the fission yeast. Curr Genet 57, 1–12 (2011). https://doi.org/10.1007/s00294-010-0327-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-010-0327-x