Abstract.
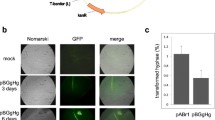
Conditions for the production of protoplasts and gene transfer in Pythium aphanidermatum were investigated. Efficient protoplast generation was possible after culture of mycelium in potato dextrose broth followed by digestion with 0.5% (w/v) each of cellulase and β-d-glucanase. Plasmid pHAMT35N/SK encoding the nptII gene under control of the Ham34 promoter from the oomycete Bremia lactucae was used to define electroporation parameters for gene transfer. A square-wave electroporation pulse of 2500 V/cm at 50 μF capacitance reproducibly produced transformants, albeit at low efficiency (0.1–0.4 transformants from ~105 regenerable protoplasts per microgram of DNA). Thirty-two independent transformants exhibited wild-type growth on potato dextrose agar amended with geneticin at 50 μg/ml, a concentration that near completely inhibited the growth of untransformed P. aphanidermatum. Southern blot analysis indicated that transforming DNA was integrated into the oomycete genome and that the DNA was stably inherited through sporogenesis. Growth on geneticin-free media, the ability to form zoospores or oospores, and the ability to cause disease in sugarbeet seedlings in the laboratory were indistinguishable between a subset of the transformed isolates and the progenitor isolate 898B. Co-electroporation of pHAMT35N/SK with plasmid pACT-GUS encoding the Escherichia coli gusA gene controlled by oomycete transcriptional promoter and terminator sequences or with pEGFP encoding enhanced green fluorescent protein under the control of the immediate early promoter from the mammalian cytomegalovirus produced, respectively, stable β-glucuronidase and transient expression of blue-green fluorescence. Application of the technique to studies on the biochemical basis for pathogenesis in this agriculturally important group of fungi is discussed.





Similar content being viewed by others
References
Agrios GN (1988) Plant pathology, 3rd edn. Academic Press, St. Paul, Minn.
Bae YS, Knudsen GR (2000) Cotransformation of Trichoderma harzianum with beta-glucuronidase and green fluorescent protein genes provides a useful tool for monitoring fungal growth and activity in natural soils. Appl Environ Microbiol 66:810–815
Bailey AM, Mena GL, Estrella-Herrera L (1991) Genetic transformation of the plant pathogens Phytophthora capsici and Phytophora parasitica. Nucleic Acids Res 19:4273–4278
Beck E, Ludwig G, Auerswald EA, Reiss B, Schaller H (1982) Nucleotide sequence and exact localization of the neomycin phosphotransferase gene from transposon Tn5. Gene 19:327–336
Bölker M, Böhnert HU, Braun K-H, Görl J, Kahmann R (1995) Tagging pathogenicity genes in Ustilago maydis by restriction enzyme mediated integration (REMI). Mol Gen Genet 248:547–552
Bottin A, Larche L, Villalba F, Gaulin E, Esquerre-Tugaye MT, Rickauer M (1999) Green fluorescent protein (GFP) as gene expression reporter and vital marker for studying development and microbe–plant interaction in the tobacco pathogen Phytophthora parasitica var nicotianae. FEMS Microbiol Lett 176:51–56
Chang DC, Saunders JA, Chassy BM, Sowers AE (1992) Overview of electroporation and electrofusion. In: Chang DC, Chassy BM, Saunders JA, Sowers AE (eds) Guide to electroporation and electrofusion. Academic Press, San Diego, pp 1–9
Chen WC, Hsieh H-J, Tseng T-C (1998) Purification and characterization of a pectin lyase from Pythium splendens-infected cucumber fruits. Bot Bull Acad Sin 39:181–186
Ciuffetti LM, Tuori RP, Gaventa JM (1997) A single gene encodes a selective toxin causal to the development of tan spot of wheat. Plant Cell 9:135–144
De Wit PJGM (1997) Pathogen avirulence and plant resistance: a key role for recognition. Trends Plant Sci 2:452–458
Hargreaves J, Turner G (1992) Gene transformation in plant pathogenic fungi. In: Gurr SJ, McPherson MJ, Bowles DJ (eds) Molecular plant pathology: a practical approach. IRL Press, Oxford, pp 79–98
Herskowitz I (1987) Functional inactivation of genes by dominant negative mutations. Nature 329:219–222
Hwang CS, Kolattukudy PE (1995) Isolation and characterization of genes expressed uniquely during appressorium formation by Colletotrichum gloeosporioides conidia induced by the host surface wax. Mol Gen Genet 247:282–294
Judelson HS (1993) Intermolecular ligation mediates efficient cotransformation in Phytophthora infestans. Mol Gen Genet 239:241–250
Judelson HS, Tyler BM, Michelmore RW (1991) Transformation of the oomycete pathogen, Phytophthora infestans. Mol Plant-Microbe Interact 4:602–607
Judelson HS, Tyler BM, Michelmore RW (1992) Regulatory sequences for expressing genes in oomycete fungi. Mol Gen Genet 234:138–146
Judelson HS, Dudler R, Pieterse CMJ, Unkles SE, Michelmore RW (1993) Expression and antisense inhibition of transgenes in Phytophthora infestans is modulated by choice of promoter and position effects. Gene 133:63–69
Kamoun S, Huitema E, Vleeshouwers VGAA (1999) Resistance to oomycetes: a general role for the hypersensitive response. Trends Plant Sci 4:196–200
Maier FJ, Schafer W (1999) Mutagenesis via insertional or restriction enzyme-mediated integration (REMI) as a tool to tag pathogenicity related genes in plant pathogenic fungi. Biol Chem 380:855–864
Martin FN (1990) Taxonomic classification of asexual isolates of Pythium ultimum based on cultural characteristics and mitochondrial DNA restriction patterns. Exp Mycol 14:47–56
Martin FN (1992) Pythium. In: Singleton LL, Mihail JD, Rush CM (eds) Methods for research on soilborne phytopathogenic fungi. APS Press, pp 39–49
Martin FN (1995) Pythium. In: Kohmoto K, Singh US, Singh RP (eds) Eukaryotes.(Pathogenesis and host specificity in plant diseases: histopathological, biochemical, genetic and molecular bases, vol 2) Elsevier, Oxford, pp 17–36
Martin FN, Hancock JG (1987) The use of Pythium oligandrum for biological control of preemergence damping off caused by P. ultimum. Phytopathology 77:1013–1020
Martin FN, Loper JE (1999) Soilborne plant diseases caused by Pythium spp: ecology, epidemiology, and prospects for biological control. Crit Rev Plant Sci 18:111–181
Martin FN, Semer CR IV (1997) Selection of drug-tolerant strains of Pythium sylvaticum using sublethal enrichment. Phytopathology. 87:685–692
McCarter SM, Littrell RH (1970) Comparative pathogenicity of Pythium aphanidermatum and Pythium myriotylum to twelve plant species and intraspecific variation in virulence. Phytopathology 60:264–268
Monke E, Schafer W (1993) Transient and stable gene expression in the fungal maize pathogen Cochliobolus heterostrophus after transformation with the β-glucuronidase (GUS) gene. Mol Gen Genet 241:73–80
Oliver R, Osbourn A (1995) Molecular dissection of fungal phytopathogenicity. Microbiology 141:1–9
Osburn RM, Schroth MN, Hancock JG, Hendson M (1989) Dynamics of sugarbeet seed colonization by Pythium ultimum and Pseudomonas species: effects on seed rot and damping off. Phytopathology 79:709–716
Pollock MA, Oppenheimer DG (1999) Inexpensive alternative to M&S medium for selection of Arabidopsis plants in culture. Biotechniques 26:254–257
Rochelle PA, Weightman AJ, Fry JC (1992) DNase I treatment of Taq DNA polymerase for complete PCR decontamination. Biotechniques 13:520
Ruben DM, Frank ZR, Chet I (1980) Factors affecting behavior and developmental synchrony of germinating oospores of Pythium aphanidermatum. Phytopathology 70:54–59
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
Spellig T, Bottin A, Kahmann R (1996) Green flourescent protein (GFP) as a new vital marker in the phytopathogenic fungus Ustlago maydis. Mol Gen Genet 252:503–509
Stomp A-M (1992) Histochemical localization of β-glucuronidase. In: Gallagher, SR (ed) GUS protocols. Academic Press, San Diego, pp 103–113
Sweigard JA, Carroll AM, Farrall L, Chumley FG, Valent B (1998) Magnaporthe grisea pathogenicity genes obtained through insertional mutagenesis. Mol Plant-Microbe Interact 11:404–412
Van der Plaats-Niterink AJ (1981) Monograph of the genus Pythium. (Studies in mycology 21) Cenraalbureau Voor Schimmelcutures, Baarn
Walton JD (1994) Deconstructing the cell wall. Plant Physiol 104:1113–1118
Weaver JC (1995) Electroporation theory: concepts and mechanisms. In: Nicloloff JA (ed) Electroporation protocols for microorganisms. Humana Press, Totowa, N.J., pp 1–27
Weiland JJ (1997) Rapid procedure for the extraction of DNA from fungal spores and mycelia. Fungal Genet Newsl 44:60–63
West P van, Kamoun S, Klooster JW van't, Govers F (1999) Internuclear gene silencing in Phytophora infestans. Mol Cell 3:339–348
Acknowledgements.
The author wishes to acknowledge the efforts of Brant B. Bigger and Lisa M. Voll for technical expertise and creative input in the study and contributions from Gary E. Nielsen, Jeff Miller, and Roger Effertz. We are indebted to David Johnson (Michigan State University–East Lansing) for taxonomic appraisal of the P. aphanidermatum used in the study. This project was supported by USDA-ARS CRIS 0015442090. Mention of a trademark or proprietary product does not constitute a guarantee or warranty of the product by the USDA or imply approval to the exclusion of other products that may also be suitable.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Hohmann
Rights and permissions
About this article
Cite this article
Weiland, J.J. Transformation of Pythium aphanidermatum to geneticin resistance. Curr Genet 42, 344–352 (2003). https://doi.org/10.1007/s00294-002-0359-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-002-0359-y