Abstract
Measures of phylogenetic balance, such as the Colless and Sackin indices, play an important role in phylogenetics. Unfortunately, these indices are specifically designed for phylogenetic trees, and do not extend naturally to phylogenetic networks (which are increasingly used to describe reticulate evolution). This led us to consider a lesser-known balance index, whose definition is based on a probabilistic interpretation that is equally applicable to trees and to networks. This index, known as the \(B_2\) index, was first proposed by Shao and Sokal (Syst Zool 39(3): 266–276, 1990). Surprisingly, it does not seem to have been studied mathematically since. Likewise, it is used only sporadically in the biological literature, where it tends to be viewed as arcane. In this paper, we study mathematical properties of \(B_2\) such as its expectation and variance under the most common models of random trees and its extremal values over various classes of phylogenetic networks. We also assess its relevance in biological applications, and find it to be comparable to that of the Colless and Sackin indices. Altogether, our results call for a reevaluation of the status of this somewhat forgotten measure of phylogenetic balance.










Similar content being viewed by others
References
Agapow P-M, Purvis A (2002) Power of eight tree shape statistics to detect nonrandom diversification: a comparison by simulation of two models of cladogenesis. Syst Biol 51(6):866–872. https://doi.org/10.1080/10635150290102564
Aldous D (1996) Probability distributions on cladograms. Random discrete structures. Springer, New York, pp 1–18. https://doi.org/10.1007/978-1-4612-0719-1_1
Bapteste E, van Iersel L, Janke A, Kelchner S, Kelk S, McInerney JO, Morrison DA, Nakhleh L, Steel M, Stougie L, Whitfield J (2013) Networks: expanding evolutionary thinking. Trends Genet 29(8):439–441. https://doi.org/10.1016/j.tig.2013.05.007
Bienvenu F et al (2020) Data and code for revisiting Shao and Sokals \(B_2\) index of phylogenetic balance. Zenodo. https://doi.org/10.5281/zenodo.4088651
Bienvenu F, Lambert A, Steel M (2020) Combinatorial and stochastic properties of ranked tree-child networks. arXiv preprint arXiv:2007.09701
Blum MG, François O (2005) On statistical tests of phylogenetic tree imbalance: the Sackin and other indices revisited. Math Biosci 195(2):141–153. https://doi.org/10.1016/j.mbs.2005.03.003
Blum MG, François O (2006) Which random processes describe the tree of life? A large-scale study of phylogenetic tree imbalance. Syst Biol 55(4):685–691. https://doi.org/10.1080/10635150600889625
Blum MG, François O, Janson S (2006) The mean, variance and limiting distribution of two statistics sensitive to phylogenetic tree balance. Ann Appl Prob 16(4):2195–2214. https://doi.org/10.1214/105051606000000547
Cardona G, Zhang L (2020) Counting and enumerating tree-child networks and their subclasses. J Comput Syst Sci 114:84–104. https://doi.org/10.1016/j.jcss.2020.06.001
Cardona G, Rosselló F, Valiente G (2009) Comparison of tree-child phylogenetic networks. IEEE/ACM Trans Comput Biol Bioinform (TCBB) 6(4):552–569. https://doi.org/10.1109/TCBB.2007.70270
Cardona G, Mir A, Rosselló F (2013) Exact formulas for the variance of several balance indices under the Yule model. J Math Biol 67:6–7, 1833–1846. https://doi.org/10.1007/s00285-012-0615-9
Chazelle B (1985) On the convex layers of a planar set. IEEE Trans Inf Theory 31:509–517. https://doi.org/10.1109/TIT.1985.1057060
Colless DH (1982) Review of phylogenetics: the theory and practice of phylogenetic systematics. https://doi.org/10.2307/2413419
Coronado TM, Fischer M, Herbst L, Rosselló F, Wicke K (2020a) On the minimum value of the Colless index and the bifurcating trees that achieve it. J Math Biol 80(7):1993–2054. https://doi.org/10.1007/s00285-020-01488-9
Coronado TM, Mir A, Rosselló F, Rotger L (2020b) On Sackins original proposal: the variance of the leaves depths as a phylogenetic balance index. BMC Bioinform 21(1):1–17. https://doi.org/10.1186/s12859-020-3405-1
Curien N (2018) Random graphs: the local convergence point of view. Lecture notes. https://www.imo.universite-paris-saclay.fr/~curien/cours/cours-RG.pdf
Felsenstein J (2003) Inferring phylogenies, 2nd edn. Sinauer Associates, Sunderland
Fischer M (2018) Extremal values of the sackin balance index for rooted binary trees. arXiv preprint arXiv:1801.10418
Flajolet P, Prodinger H (1987) Level number sequences for trees. Discret Math 65(2):149–156. https://doi.org/10.1016/0012-365X(87)90137-3
Hayati M, Shadgar B, Chindelevitch L (2019) A new resolution function to evaluate tree shape statistics. PLoS ONE 14(11)
Heard SB (1818) Patterns in tree balance among cladistic, phenetic, and randomly generated phylogenetic trees. Evolution 46(6):1992. https://doi.org/10.2307/2410033
Huerta-Cepas J, Capella-Gutiérrez S, Pryszcz LP, Marcet-Houben M, Gabaldón T (2014) PhylomeDB v4: zooming into the plurality of evolutionary histories of a genome. Nucleic Acids Res 42(D1):D897–D902. https://doi.org/10.1093/nar/gkt1177
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23(2):254–267. https://doi.org/10.1093/molbev/msj030
Janson S (2012) Simply generated trees, conditioned Galton–Watson trees, random allocations and condensation. Probab Surv 9:103–252. https://doi.org/10.1214/11-PS188
Kingman JFC (1982) The coalescent. Stoch Process Appl 13(3):235–248. https://doi.org/10.1016/0304-4149(82)90011-4
Kirkpatrick M, Slatkin M (1993) Searching for evolutionary patterns in the shape of a phylogenetic tree. Evolution 47(4):1171. https://doi.org/10.2307/2409983
Knuth DE (1997) The art of computer programming: volume 1: fundamental algorithms. Addison-Wesley Professional, Boston
Lambert A (2017) Probabilistic models for the (sub)tree(s) of life. Brazil J Probab Stat 31(3):415–475. https://doi.org/10.1214/16-BJPS320
Maia LP, Colato A, Fontanari JF (2004) Effect of selection on the topology of genealogical trees. J Theor Biol 226(3):315–320
Matsen FA (2006) A geometric approach to tree shape statistics. Syst Biol 55(4):652–661. https://doi.org/10.1080/10635150600889617
McKenzie A, Steel M (2000) Distributions of cherries for two models of trees. Math Biosci 164(1):81–92. https://doi.org/10.1016/S0025-5564(99)00060-7
Moran PAP (1958) Random processes in genetics. Math Proc Cambridge Philos Soc 54(1):60–71. https://doi.org/10.1017/S0305004100033193
Penel S, Arigon A-M, Dufayard J-F, Sertier A-S, Daubin V, Duret L, Gouy M, Perrière G (2009) Databases of homologous gene families for comparative genomics. In: BMC bioinformatics, vol 10. https://doi.org/10.1186/1471-2105-10-S6-S3
Roesler U, Rüschendorf L (2001) The contraction method for recursive algorithms. Algorithmica 29(1):3–33
Rogers JS (1994) Central moments and probability distribution of Colless coefficient of tree imbalance. Evolution 48(6):2026–2036. https://doi.org/10.1111/j.1558-5646.1994.tb02230.x
Rogers JS (1996) Central moments and probability distributions of three measures of phylogenetic tree imbalance. Syst Biol 45(1):99. https://doi.org/10.2307/2413515
Rotger L (2019) New balance indices and metrics for phylogenetic trees. Universitat de les Illes Balears PhD thesis
Sackin MJ (1972) Good and bad phenograms. Syst Biol 21(2):225–226. https://doi.org/10.1093/sysbio/21.2.225
Scornavacca C, Belkhir K, Lopez J, Dernat R, Delsuc F, Douzery EJP, Ranwez V (2019) OrthoMaM v10: scaling-up orthologous coding sequence and exon alignments with more than one hundred mammalian genomes. Mol Biol Evol 36(4):861–862. https://doi.org/10.1093/molbev/msz015
Shao KT, Sokal RR (1990) Tree balance. Syst Zool 39(3):266–276. https://doi.org/10.2307/2992186
The On-Line Encyclopedia of Integer Sequences (2020) Published electronically at https://urldefense.proofpoint.com/v2/urls?u=https-3A_oeis.org&d=DwIDaQ&c=vh6FgFnduejNhPPD0fl_yRaSfZy8CWbWnIf4XJhSqx8&r=JxLWRfjFp6vfB3IFaoebJ17aAJLUj5TpdLYeq8QKCxw&m=Ztg9MHwGTuar2preoVNEAMSrBxLjgsHNLKc4rjHH9jM&s=SDrn3nxnalS5qgTScck5RUEauNLjWtOc4mZSbz5S_s&e=
Vos RA, Balhoff JP, Caravas JA, Holder MT, Lapp H, Maddison WP, Midford PE, Priyam A, Sukumaran J, Xia X, Stoltzfus A (2012) NeXML: rich, extensible, and verifiable representation of comparative data and metadata. Syst Biol 61(4):675–689. https://doi.org/10.1093/sysbio/sys025
Wei C, Gong D, Wang Q (2013) Chu-Vandermonde convolution and harmonic number identities. Integral Transform Spec Funct 24(4):324–330. https://doi.org/10.1080/10652469.2012.689762
Acknowledgements
The authors thank Simon Penel for his help with the HOGENOM database and Roberto Bacilieri for helpful discussions. FB and CS were funded by grants ANR-16-CE27-0013 and ANR-19-CE45-0012 from the Agence Nationale de la Recherche, respectively; GC was funded by FEDER / Ministerio de Ciencia, Innovación y Universidades / Agencia Estatal de Investigación project PGC2018-096956-B-C43.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix
Appendix
1.1 A.1 Variance of \(B_2\) in binary Galton–Watson trees
In this section, we prove Proposition 3.3 concerning the variance of \(B_2\) in binary Galton–Watson trees. Let us start with a standard result, which we recall and prove for the sake of completeness.
Lemma A.1.1
Let \((Z_t)\) be a Galton–Watson process with offspring distribution \(Y \sim 2\, \mathrm {Bernoulli(1/2)}\). Then, \({\mathbb {E}}\!\left( {Z_t^2}\right) = t + 1\).
Proof
Since \(Z_{t + 1} = \sum _{i = 1}^{Z_t} Y_t(i)\), we have
Letting \({\mathcal {F}}_t\) be the natural filtration of the process, we thus have
and, as a result,
Since here \({\mathbb {E}}\!\left( {Y}\right) = 1\), \({\mathbb {E}}\!\left( {Y^2}\right) = 2\) and \({\mathbb {E}}\!\left( {Z_t}\right) = 1\), this simplifies to
The lemma then follows by induction, since \({\mathbb {E}}\!\left( {Z_0^2}\right) = 1\). \(\square \)
Let us now recall Proposition 3.3 and prove it.
Proposition 3.3
Let T be a critical binary Galton–Watson tree, that is, assume that the offspring distribution is \({\mathbb {P}}\!\left( {Y = 0}\right) = {\mathbb {P}}\!\left( {Y = 2}\right) = 1/2\). Then,
Proof
To alleviate the notation, let us write \(B_t = B_2(T_{[t]})\). With this notation, since in the case of a binary Galton–Watson tree \({\mathbb {1}}_{\{Y_t > 0\}} = Y_t / 2\), Equation (3) from the proof of Theorem 3.1 becomes
Therefore, letting \({\mathcal {F}}_t\) denote the natural filtration of the process and using that \({\mathbb {E}}\!\left( {Y}\right) = 1\), we have
As a result,
Let us now turn our attention to \({\mathbb {E}}\!\left( {B_t Z_t}\right) \). Using Equation (A.1), we get
and since \({\mathbb {E}}\!\left( {\left. B_t \sum _{i = 1}^{Z_t} Y_t(i) \right| {\mathcal {F}}_t}\right) = B_t Z_t\) this yields
By Lemma A.1.1, \({\mathbb {E}}\!\left( {Z_{t + 1}^2}\right) = t + 2\). Since \({\mathbb {E}}\!\left( {B_0 Z_0}\right) = 0\), we thus have
Plugging this and \({\mathbb {E}}\!\left( {Z_{t + 1}^2}\right) = t + 2\) in Equation (A.2), we get the following closed recurrence relation for \({\mathbb {E}}\!\left( {B_t^2}\right) \):
Solving this recurrence relation with the initial condition \({\mathbb {E}}\!\left( {B_0^2}\right) = 0\) then yields
Finally, since by Theorem 3.1, \({\mathbb {E}}\!\left( {B_t}\right) = 1 - 2^{-t}\), we have
concluding the proof. \(\square \)
1.2 A.2 Bounds on the number of distinct values of \(B_2\)
Proposition A.2.2
Let \({\mathscr {T}}_n\) denote the set of rooted binary trees with n leaves, labeled or unlabeled. Then,
where \(a_n\) is sequence A002572 in Online Encyclopedia of Integer Sequences (2020) and satisfies \(a_n \sim K \rho ^n\), with \(\rho \approx 1.7941\) and \(K \approx 0.2545\) the Flajolet–Prodinger constant.
Proof
The upper bound is obtained by noting that \(B_2(T)\) is a function of the multiset of the depths of the leaves of the binary tree T. Therefore, \(B_2\) cannot take more values than the number \(a_n\) of such multisets, whose asymptotics were characterized by Flajolet and Prodinger in Flajolet and Prodinger (1987).
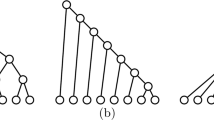
To obtain the lower bound, we exhibit \(2^{\lfloor {n/2}\rfloor - 1}\) rooted binary trees with n leaves whose \(B_2\) indices are different. For any integer m and any \({\mathbf {x}} \in \{0, 1\}^m\), let \(T({\mathbf {x}})\) denote the ordered (that is, embedded in the plane) rooted binary tree obtained by the following sequential construction: starting from the binary tree with two leaves, for \(k = 1, \ldots , m\),
-
If \(x_k = 0\), graft a cherry on the left-most leaf with depth k.
-
If \(x_k = 1\), graft a cherry on each of the two left-most leaves with depth k.
This construction is illustrated in Fig. 11.
Clearly, \(T({\mathbf {x}})\) has \(2 + \sum _{k = 1}^{m} (x_k + 1)\) leaves and, by Corollary 1.11,
As a result,
(to see this, note that \(B_2(T({\mathbf {x}})) = u_m + \underline{{\mathbf {x}}}_{(2)}\), where \(\underline{{\mathbf {x}}}_{(2)} = \sum _{k} x_k 2^{-k}\) denotes the dyadic rational of \([{0, 1}[\) whose binary expansion is \({\mathbf {x}}\)). Thus, this construction generates \(2^m\) trees whose \(B_2\) indices differ by at least \(2^{-m}\). However, these trees do not have the same number of leaves.
Let us first assume that n is even. Then, with \(m = n/2 - 1\), \(T(1\cdots 1)\) has n leaves and every other tree \(T({\mathbf {x}})\) with \({\mathbf {x}} \in \{0, 1\}^m\) has:
-
(i)
\(n - k_{\mathbf {x}}\) leaves, with \(1 \le k_{\mathbf {x}} \le m\);
-
(ii)
its left-most leaf at depth \(m + 1\).
Now, if we graft a caterpillar with \(k_{\mathbf {x}} + 1\) leaves on the left-most leaf at depth \(m + 1\) and let \(T'({\mathbf {x}})\) denote the resulting tree, then:
- (\(\mathrm {i}^\prime \)):
-
\(T'({\mathbf {x}})\) has n leaves;
- (\(\mathrm {ii}^\prime \)):
-
by Proposition 1.10, \(B_2(T'({\mathbf {x}})) - B_2(T({\mathbf {x}})) = 2^{-(m+1)}(2 - 2^{-k+1}) \in ]{0, 2^{-m}}[\).
Since the \(B_2\) indices of the trees \(T({\mathbf {x}})\) differ by at least \(2^{-m}\), by point (\(\mathrm {ii}^\prime \)) the \(B_2\) indices of the trees \(T'({\mathbf {x}})\) are all different, thereby proving the proposition in the case where n is even.
If n is odd, do the same construction, again with \(m = \lfloor n / 2\rfloor - 1\), to get \(2^m\) trees with \(n - 1\) leaves. Then, for each of these trees, graft a cherry on the sibling of the left-most vertex at depth \(m + 1\) (which exists and is always a leaf). This extra step increases \(B_2\) by the same amount \(2^{-(m + 1)}\) for every tree, concluding the proof. \(\square \)
Rights and permissions
About this article
Cite this article
Bienvenu, F., Cardona, G. & Scornavacca, C. Revisiting Shao and Sokal’s B2 index of phylogenetic balance. J. Math. Biol. 83, 52 (2021). https://doi.org/10.1007/s00285-021-01662-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00285-021-01662-7