Abstract
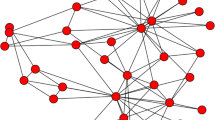
This paper investigates the effects of the community structure of a network on the spread of an epidemic. To this end, we first establish a susceptible–infected–recovered (SIR) model in a two-community network with an arbitrary joint degree distribution. The network is formulated as a probability generating function. We also obtain the sufficient conditions for disease outbreak and extinction, which involve the first-order and second-order moments of the degree distribution. As an example, we then study the effect of community structure on epidemic spread in a complex network with a Poisson joint degree distribution. The numerical solutions of the SIR model well agree with stochastic simulations based on the Monte Carlo method, confirming that the model is reliable and accurate. Finally, by strengthening the community structure in the simulation, i.e. fixing the total degree distribution and reducing the number ratio of the external edges, we can increase or decrease the final cumulative epidemic incidence depending on the transmissibility of the virus between humans and the community structure at that point. Why community structure can affect disease dynamics in a complicated way is also discussed. In any case, for large-scale epidemics, strengthening the community structure to reduce the size of disease is undoubtedly an effective way.















Similar content being viewed by others
References
Ball F (1983) The threshold behavior of epidemic models. J Appl Probab 20(2):227–241
Ball F, Neal P (2008) Network epidemic models with two levels of mixing. Math Biosci 212:69–87
Ball F, Mollison D, Scalia-Tomba G (1997) Epidemics in populations with two levels of mixing. Ann Appl Probab 7:46–89
Ball FG, Sirl DJ, Trapman P (2009) Threshold behaviour and final outcome of an epidemic on a random network with household structure. Adv Appl Probab 41:765–796
Ball FG, Sirl DJ, Trapman P (2010) Analysis of a stochastic SIR epidemic on a random network incorporating household structure. Math Biosci 224:53–73
Ball F, Britton T, Sirl D (2013) A network with tunable clustering, degree correlation and degree distribution and an epidemic thereon. J Math Biol 66:979–1019
Chris TB (2005) The spread of infectious diseases in spatially structured populations: an invasory pair approximation. Math Biosci 198:217–237
Eames KTD (2008) Modelling disease spread through random and regular contacts in clustered populations. Theor Popul Biol 73:104–111
Gleeson JP (2009) Bond percolation on a class of clustered random networks. Phys Rev E 80:036107
Gleeson JP, Melnik S, Adam H (2010) How clustering affects the bond percolation threshold in complex networks. Phys Rev E 81:066114
Gross T, DLima C, Blasius B (2006) Epidemic dynamics on an adaptive network. Phys Rev Lett 96(20):208701
House T, Keeling MJ (2010) Epidemic prediction and control in clustered populations. J Theor Biol 272(1):1–7
House T, Keeling MJ (2011) Insights from unifying modern approximations to infections on networks. J R Soc Interface 8:67–73
Huang W, Li C (2007) Epidemic spreading in scale-free networks with community structure. J Stat Mech 1(1):01014
Jin EM, Girvan M, Newman MEJ (2001) Structure of growing social networks. Phys Rev E 64(2):322–333
Kiss IZ, Miller JC, Simon PL (2017) Mathematics of epidemics on networks: from exact to approximate models. Springer, New York
Koch D, Illner R, Ma JL (2013) Edge removal in random contact networks and the basic reproduction number. J Math Biol 67:217–238
Liu Z, Hu B (2005) Epidemic spreading in community networks. EPL 72(2):315–321
Liu JZ, Wu JS, Yang ZR (2004) The spread of infectious disease with household-structure on the complex networks. Physica A 141:273–280
Luo XF, Zhang X, Sun GQ, Jin Z (2014) Epidemical dynamics of SIS pair approximation models on regular and random networks. Physica A 410:144–153
Ma J, Van DP, Willeboordse FH (2013) Effective degree household network disease model. J Math Biol 66:75–94
Miller J (2009a) Percolation and epidemics in random clustered networks. Phys Rev E 80:020901R
Miller J (2009b) Spread of infectious disease through clustered populations. J R Soc Interface 6:1121–1134
Miller JC (2011) A note on a paper by Erik Volz: SIR dynamics in random networks. J Math Biol 62:349–358
Miller JC, Volz EM (2013) Incorporating disease and population structure into models of SIR disease in contact networks. PLoS One 8:1–14
Molloy M, Reed R (1995) A critical point for random graphs with a given degree sequence. Random Struct Algorithms 6(2–3):161–180
Neal P (2007) Coupling of two SIR epidemic models with variable susceptibility and infectivity. J Appl Probab 44:41–57
Newman MEJ (2002) Spread of epidemic disease on networks. Phys Rev E 66:016128
Newman MEJ (2003a) Mixing patterns in networks. Phys Rev E 67(2):241–251
Newman MEJ (2003b) Properties of highly clustered networks. Phys Rev E 68:026121
Newman MEJ, Girvan M (2004) Finding and evaluating community struture in networks. Phys Rev E 69(2):026113
Orman K, Labatut V, Cherifi H (2013) An empirical study of the relation between community structure and transitivity. Complex Netw SCI 424:99–110
Peng XL, Small M, Xu XJ, Fu X (2013) Temporal prediction of epidemic patterns in community networks. New J Phys 15(11):17161–17175
Rowthorn RE, Laxminarayan R, Gilligan CA (2009) Optimal control of epidemics in metapopulations. J R Soc Interface 6:1135–1144
Salathé M, Jones JH (2010) Dynamics and control of diseases in networks with community structure. PLOS Comput Biol 6(4):e1000736
Sun HJ, Gao ZY (2007) Dynamical behaviors of epidemics on scale-free networks with community structure. Physica A 381:491–496
Traud A, Kelsic E, Mucha P, Porter M (2009) Community structure in online collegiate social networks. APS March Meet 53(3):526–543
Tunc I, Shaw LB (2014) Effects of community structure on epidemic spread in an adaptive network. Phys Rev E 90(2):022801
Volz E (2008) SIR dynamics in random networks with heterogeneous connectivity. J Math Biol 56:293–310
Wang B, Cao L, Suzuki H, Aihara K (2010) Epidemic spread in adaptive networks with multitype agents. J Phys A Math Theor 44:035101
Wang B, Cao L, Suzuki H, Aihara K (2012) Impacts of clustering on interacting epidemic. J Theor Biol 304:121–130
Wu XY, Liu ZH (2008) How community structure influences epidemic spread in social networks. Physica A 387:623–630
Yan G, Fu ZQ, Ren J, Wang WX (2007) Collective synchronization induced by epidemic dynamics on complex networks with communities. Phys Rev E 75(1 Pt 2):112–118
Zachary W (1977) An information flow model for conflict and fission in small groups. J Anthropol Res 33(4):452–473
Zhou J, Liu ZH (2009) Epidemic spreading in communities with mobile agents. Physica A 388:1228–1236
Acknowledgements
This work is supported by the National Natural Science Foundations of China under Grant (Nos. 11571210, 11331009, 11501339, 11101251, 11001157, 11471197, 11701348) and the Youth Science Foundation of Shanxi Province (No. 2010021001-1).The authors wish to thank the anonymous referees for their helpful feedback and suggestions.
Author information
Authors and Affiliations
Corresponding author
Appendices
Appendix A: the dynamics of SIR epidemic model in a network with two communities
In this section, we develop an SIR-epidemic model involving the variables \(\theta _{mn}\), \(p^{I_m}_{S_n}\) and \(p^{S_m}_{S_n}\), \(m,n=1,2\). The method used is similar to that (Volz 2008).
Firstly, we introduce the dynamics of \(\theta _{mn}\), \(m,n=1,2.\) Consider a susceptible node ego in community l at time t with a joint degree (k, j), there are a set of k internal arcs \((\text{ ego }^l,\text{ alter }^{in}_{1})\), \((\text{ ego }^l,\text{ alter }^{in}_{2})\), \(\cdots \), \((\text{ ego }^l, \text{ alter }^{in}_{k})\) and a set of j external arcs \((\text{ ego }^l,\text{ alter }^{out}_{1})\), \((\text{ ego }^l,\text{ alter }^{out}_{2})\), \(\cdots \), \((\text{ ego }^l,\text{ alter }^{out}_{j})\) corresponding to the ego. We assume that for each arc \((\text{ ego }^l,\text{ alter }^{in}_{m})\) for \(m=1,2,\cdots , k\) and \((\text{ ego }^l,\text{ alter }^{out}_{n})\) for \(n=1,2,\cdots , j\), there will be uniform probabilities \(p^{I_l}_{S_l}=M^{I_l}_{S_l}/M^{in}_{S_l}\) and \(p^{I_n}_{S_l}=M^{I_n}_{S_l}/M^{out}_{S_l}\) for \(n\ne l\) that \(alter^{in}_{m}\) and \(alter^{out}_{n}\) are infectious, respectively. In a time dt, an expected number \(\gamma (k p^{I_l}_{S_l}+j p^{I_n}_{S_l})dt\) for \(n\ne l\) of these will be such that the infectious alter transmits to ego. Consequently, the hazard for ego becoming infected at time t is
Now let \(u^l_{(k,j)}(t)\) represent the fraction of nodes with joint degree (k, j) in community l which remain susceptible at time t. Using Eq. (42), we have
Subsequently, let
Then
Thus, the fraction of nodes which remain susceptible at time t in community l for \(l=1, 2\) are
respectively and the fraction of nodes which remain susceptible in the whole network at time t is
Similarly, for \(l,n=1,2\) and \(n\ne l\), we have
In terms of \(A^{in}_l=NG_l{(1,1)}\) and Eq. (46), we have
According to Eq. (44), the dynamics of \(\theta _{ln}\), \(l, n=1, 2\) is
which depends on the variable \(p^{I_n}_{S_l}\). To close the above system, we have to calculate the dynamics of \(p^{I_n}_{S_l}\).
Second, we introduce the dynamics of \(p^{I_n}_{S_l}\) for \(l,n=1,2\). The definition of \(p^{I_n}_{S_l}\) is
Hence,

From Eq. (47), we easily get
and
where \(l,n=1,2\) and \(n\ne l\).
The dynamics of \(M^{I_n}_{S_l}\) for l, \(n=1,2\) depends on the state of excessive neighbors (not counting the alter \(I_n\) in the chosen arc \((S_l, I_n)\)) of the ego node \(S_l\) for an random chosen arc \((S_l, I_n)\) in \(A^{I_n}_{S_l}\) for l, \(n=1,2\). The following notations in Table 2 are needed to clarify subsequent calculations.
The PGF for the excessive degree distribution of the ego node \(S_l\) in the selected arc \((S_l, I_n)\) are
The derivation of the Eq. (53) is presented in the “Appendix B” as an example.
Because arcs are distributed polynomially to nodes in sets S, I, R, we have \(g_{S_1I_1}{(X_{S_1},X_{I_1},X_{R_1},X_{S_2},X_{I_2},X_{R_2})}=g_{S_1S_1}{(X_{S_1},X_{I_1},X_{R_1},X_{S_2},X_{I_2},X_{R_2})}\), which is easy to verify by repeating the calculation in Eq. (53).
By Eqs. (53) and (54), we have
We next obtain the dynamics of \(M_{S_l}^{I_l}\). Volz (2008) introduces the approach to model SIR dynamics on networks. Here, we briefly present this approach.
There are three reasons for the reduction of the fraction of arcs between \(S_l\) and \(I_l\). One reason is the newly infectious nodes in community l. In a time infinitesimal dt, the fraction of newly infectious nodes is \(-\dot{s}_l\). According to Eq. (45), we can obtain
in which \(-\frac{N}{N_l}G_l(\theta _{ll},\theta _{ln})\dot{\theta }_{ll}\) describes a fraction nodes infected by infectious alters in community l and \(-\frac{N}{N_l}G^l(\theta _{ll},\theta _{ln})\dot{\theta }_{ln}\) describes a fraction nodes infected by infectious alters in community n for \(l\ne n\). Therefore, \(M^{I_l}_{S_l}\) decreases at rate
Because \(\delta _{S_lI_l}(I_l)\) does not count the arc along which a node was infected, another reason is the transmission from the infectious alter to the susceptible ego in this arc, which results in a reduction at a rate
The third reason is the recovery of the infectious alter in the arcs, which brings about a reduction at a rate
On the other hand, the increase of \(M^{I_l}_{S_l}\) is only due to the infection of the fraction of the susceptible egos which have excessive susceptible neighbors. So \({M^{I_l}_{S_l}}\) increases at the rate
From Eqs. (48), (59)–(60), we have
Now applying Eqs. (47), (48), (51) and (61) to Eq. (50), we obtain the dynamics of \( p^{I_l}_{S_l}\), i.e.
Similarly, we can obtain the dynamics of \(p^{I_n}_{S_l}\), \(p^{S_l}_{S_l}\) and \(p^{S_n}_{S_l}\) for \(l\ne n\) and \(l, n=1, 2\).
Appendix B: the PGF for the excessive degree distribution of the ego node in an arc
We give PGF for the excessive degree distribution of the ego node \(S_l\) in an arc \((S_l, I_n)\), which is a conditional PGF involving the state of the ego node.
For an arbitrary chosen arc \(\xi \), define some random variables. Easy to find, we list them in Table 3. Let A, B and C represent the state of a node and they may be S, I or R. Let \(P(i_1, j_1, l_1, i_2, j_2, l_2\mid \xi )\) be the conditional probability that \(\xi _{S_1}=i_1\), \(\xi _{I_1}=j_1\), \(\xi _{R_1}=l_1\), \(\xi _{S_2}=i_2\), \(\xi _{I_2}=j_2\) and \(\xi _{R_2}=l_2\) for an arbitrary chosen arc \(\xi \). The amount of excessive neighbors in state S, I and R and in community l, \(l=1,2\), of ego node in state A and in the chosen arc are distributed polynomially with probability \(p^{I_l}_{A}\), \(p^{S_l}_{A}\) and \(p^{R_l}_{A}=1-p^{I_l}_{A}-p^{S_l}_{A}\), \(l=1,2\), respectively.
The PGF for the excessive degree distribution of the ego node \(S_1\) in the chosen arc \((S_1, I_1)\), is
From the definition of the conditional probability, we have
Grounded on total probability formula, we have
and
where
Since the amount of excessive neighbors of ego node in state S and in the chosen arc \((S_1,I_1)\) are distributed polynomially with probability \(p_{S_1}^{S_l}\), \(p_{S_1}^{I_l}\), \(p^{R_l}_{S_1}=1-p^{S_l}_{S_1}-p_{S_1}^{I_l}\), \(l=1,2\), respectively, we have
From Eqs. (63), (64), (65), (66) and (67), we have
Furthermore, Eq. (68) leads to
where
and
Thus, from Eqs. (70), (71) and (72), we have
where \(\alpha =p^{S_1}_{S_1}X_{S_1}+p^{I_1}_{S_1}X_{I_1}+p^{R_1}_{S_1}X_{R_1}\) and \(\beta =p^{S_2}_{S_1}X_{S_2}+p^{I_2}_{S_1}X_{I_2}+p^{R_2}_{S_1}X_{R_2}\). As in the above analysis, we have
From Eqs. (69), (73) and (74), we have
Rights and permissions
About this article
Cite this article
Li, J., Wang, J. & Jin, Z. SIR dynamics in random networks with communities. J. Math. Biol. 77, 1117–1151 (2018). https://doi.org/10.1007/s00285-018-1247-5
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00285-018-1247-5