Abstract
An increasing number of bacterial pathogens are acquiring resistance to the commonly used antibiotics. This has spurred a global threat leading to a resistance era and has penetrated the consciousness of the common people and the clinicians alike. The delay in discovering new antibiotics has exacerbated the resistance problem, forcing researchers to focus on unconventional antimicrobial therapeutics that differ from conventional antibiotics. Alternative therapies have emerged in recent years, including antimicrobial peptides, phage therapy, efflux pump inhibitors, antibodies, and immunomodulatory agents, which have produced impressive results in both laboratory and in clinical trials. Additionally, ultra-narrow-spectrum therapeutics such as CRISPR-Cas system and peptide nucleic acids aided in the development of sequence-specific antimicrobials. Moreover, combinatorial therapies that combine these new approaches have been efficient enough to get approval for clinical use and have accelerated the discovery of novel combination approaches that enhance the performance of already in-use antibiotics. In this review, we provide an overview of these approaches along with studies that focus on the uncharted microbial territories that have been able to deliver some of the important new antibiotics of recent times. It is hoped that the information gathered in this article will provide an update on the current antibiotic resistance threat and encourage profound research.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Since the discovery of penicillin in 1929, the “wonder drug” has been used indiscriminately to eliminate infection-causing organisms. The discovery of penicillin, its modifications, and the subsequent discovery of bioactive scaffolds derived from natural products served as thrusters to advance the antibiotic field into its golden age. Antibiotics were effective enough to treat bacterial infections caused by medical procedures that could otherwise would have been fatal. Because of the decrease in the prevalence of infectious diseases, clinicians have been focusing more on the improvement of medical procedures. But since the onset of the resistance era, life is being continuously threatened by microbes that can withstand every known antibiotic action resulting in the emergence of multidrug-resistant (MDR) bacteria. The category of multidrug-resistant bacteria has been segregated into extensively drug-resistant (XDR) bacteria and pan-drug resistance (PDR) bacteria depending on the degrees of resistance to varieties of drugs including the pathogens which are resistant to all the approved antimicrobial agents.
In a study that was published in 2011, researchers demonstrated that the phenomenon of antibiotic resistance is ancient and predates the current pressure to use antibiotics. When metagenomic DNA from 30,000-year-old permafrost sediments was analyzed, it was discovered to contain several genes for β-lactam, carbapenem, and vancomycin resistance and to be strikingly similar to their modern counterparts [1]. This suggests that the widespread use of antibiotics in modern times has favored the resistance determinants that were already present in the bacterial pan-genome. Thus, the antibiotic resistome, or the diversity of the genes responsible for antibacterial resistance must be thoroughly understood before new mechanisms for inhibiting bacterial growth can be developed [2]. In the year 2021, Lopatkin et al. discovered that antibiotic resistance may be conferred by mutations in the core metabolic genes which regulate innate pathways in bacteria [3]. Since antibiotic treatment affects metabolic pathways in bacteria in numerous complex ways, it is reasonable to investigate the plethora of metabolic changes that may confer resistance. As a result, metabolic mutations must be considered as a new mechanism of resistance in addition to the three major mechanisms of resistance: target modification, drug inactivation, and drug transportation. Pathogens, armed with these mechanisms, are constantly in conflict with researchers trying to render them inactive.
It has been proposed that antibacterial resistance is inevitable and likely to persist even if antibiotic use is reduced [4]. Moreover, the overuse of antibiotics through over-the-counter prescriptions, which generally do not follow clinical guidelines, contributes significantly to the spread of resistance. Developing countries use antibiotics more indiscriminately than developed countries, but it is evident that even the high-income countries face challenges of access and excess [5]. As a result, novel strategies for dealing with these new realities are desperately needed. Recent strategies being developed by researchers all over the world, as well as the current state of antibiotic resistance, have been highlighted in this review. Current research is focusing on finding new alternatives to antibiotics, compounds that target previously ignored unconventional mechanisms of bacteria, and revisiting previously described methods. The information covered in this article is intended to give readers some ideas for commencing research and development for introducing new techniques to strengthen our fight against drug-resistant pathogens.
The Current State of Antibiotic Resistance
Resistance to commonly used antibiotics has recently peaked resulting in the emergence of multidrug-resistant bacteria capable of resisting the action of last resort antibiotics such as colistin and tigecycline.
WHO recently published a report on the global shortage of novel antimicrobials in April 2021, concluding that the current antibiotic development pipelines and clinically approved antibiotics are insufficient to combat drug-resistant bacteria. [6]. The report analyzes the antibiotics in clinical development against pathogens outlined in February 2017 bacterial priority pathogens list. For the first time, WHO included non-traditional antibacterial medicines in its report, which examined 27 non-traditional antibacterial agents such as antibodies and bacteriophages [7]. Moreover, in the Indian scenario, the Indian Council of Medical Research (ICMR) in its annual report on Antimicrobial Resistance Research and Surveillance Network (from January 2020 to December 2020) represented recent resistance trends on the priority pathogens [8]. The report highlighted the lowering sensitivities of pathogens to commonly used antibiotics for drug-resistant pathogens such as cephalosporins, carbapenems, monobactams, and β-lactam—β-lactamase inhibitors based on data collected from tertiary hospitals and research labs. Because India was hit by the COVID-19 pandemic during the reporting period, the resistance pattern was also examined among isolates from secondary bacterial infections in COVID-19-positive patients. These data revealed that Klebsiella pneumoniae (K. pneumoniae) was the most commonly isolated pathogen from the respiratory tract of COVID-19-positive patients followed by Acinetobacter baumannii (A. baumannii) and Escherichia coli (E. coli). Additionally, it has been discovered that when these pathogens were isolated from COVID-19-positive patients, their antibiotic resistance increased noticeably. In addition, according to a recent report by the Center for Disease Control (CDC) in the United States, an estimated 2.8 million people are infected with antibiotic-resistant pathogens each year, with 35,000 deaths. Furthermore, $4.6 billion was required to treat infections caused by six multidrug-resistant bacteria. The CDC has categorized 18 antibiotic-resistant bacteria and fungi into three categories based on their level of concern for human health, which are “urgent, serious, and concerning” [9]. Considering these scenarios, there is an urgent need for the development of novel therapies to treat bacterial infections caused by multidrug-resistant pathogens.
Revisiting Natural Products as a Source of Novel Antibiotics
The current state of antibiotic resistance demands alternative strategies for dealing with the growing problem and revisiting the natural products derived from microorganisms is one such promising field. Various studies attempting to explore the natural environment with new techniques insights have resulted in the discovery of rare and novel antimicrobials. In an attempt to explore the vast universe of uncharted microbes, Omsland et al., were successful in cultivating Coxiella burnetti (C. burnetti) in a host-cell-free environment through optimization of culture media [10]. Since then, research on natural products from this rare organism has been on the rise. As evident from this study, the key to finding rare organisms is to find the optimal culture condition to grow the said organism. One of the ways to find this synchronized condition is to use microfluidics devices. In another study by Ma et al., a new bacterium called microfluidics I was isolated by developing a device with 3200 nanoliter-sized wells which, when allowed to process the sample, would contain only one cell at a time, allowing it to grow without any competition from other species for nutrients. The study’s most intriguing aspect was the use of samples targeting the human microbiota in search of novel species [11]. A research team from Northeastern University in Boston, along with other collaborators, attempted to incubate the microorganisms in their own natural habitats, i.e., soil, with a recently developed device called the iChip. This resulted in the identification of one of the most significant antibiotics in recent years, teixobactin [12]. The iChip allowed the growth of many different species which would not have been possible if it were incubated in conventional laboratory conditions leading to the discovery of Eleftheria terrae, a bacterium that produced the antibiotic teixobactin. This antibiotic, a macrocyclic depsipeptide, has distinct pharmacological properties along with interesting structural features such as D-amino acids and non-standard amino acids. It is a potent antimicrobial against Gram-positive bacteria where it interrupts cell wall synthesis by binding to lipid II and lipid III which are essential for peptidoglycan and wall teichoic acid synthesis, respectively [13]. Because it is a moderately complex depsipeptide, researchers have begun complete synthesis of the molecule to simplify the remaining structural complexities. This has resulted in the synthesis of various teixobactin analogues with enhanced and distinct modes of action, such as supramolecular assembly, which ultimately disrupts the cell membrane. [14].
Prospecting for genes that produce bioactive compounds is also prominent for locating new species and their compounds. A machine-learning algorithm was developed by Donia et al., that recognizes the pattern in these important genes and looks for similar hits when let loose on bacterial genomes of interest. Recently, the team employed an improved version of this algorithm for prospecting of genes from the human microbiota and revealed a substantial number of biosynthetic gene clusters for many important bioactive compounds. Further investigations led to the structure elucidation of a thiopeptide antibiotic, lactocillin, from a prominent member of the vaginal microbiota [15].
Natural product discovery is an ongoing search that has resulted in the discovery of some of the most important molecules in recent history. With the aid of cutting-edge sequencing platforms and prediction algorithms, ventures into previously uncharted natural habitats have gained popularity and shown promising results.
Non-Traditional Approaches to Combat Antibiotic Resistance
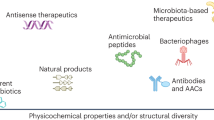
The vast knowledge on pathogen cellular mechanics has highlighted the various potential targets that can be exploited for antibiotic development. Important metabolic pathways in bacteria that can be inhibited with having a detrimental effect on their survival should be the targets of new drugs. Consequently, researchers have focused on targets for which resistance should be hard to develop. The arsenal for combatting resistance can be expanded to include nanomaterials and phytopharmaceuticals, which have recently gained global attention. Nanomaterials such as nanoparticles, nanorods, nanowires, and 2D materials can augment the action of antibiotics by acting as carriers or they can be used as standalone bactericidal agents [16, 17]. On the other hand, phytochemicals of various classes have demonstrated substantial inhibitory potential against drug-resistant pathogens by interfering with resistance pathways such as efflux pumps, biofilms, and bacterial cell communications [18]. But translating these phytochemicals into marketable product has been in a major standstill. The major non-traditional strategies that were outlined by WHO that have the potential to be translated into marketable product include antimicrobial peptides. These are short peptides with variable mode of actions that can cripple the pathogen defense systems, efflux pump inhibitors, which act to block one of the most efficient mechanisms developed by living beings to throw out unwanted chemicals, phage therapy, which uses one of nature’s menacing organisms such as the viruses that only targets bacteria in their own specific way, and combinatorial therapy, which attempts to blend the properties of some important compounds that increases the efficacy of antibiotics and sensitizes the pathogens to commonly used drugs. The WHO has recently acknowledged antibodies and immunomodulatory drugs as non-traditional therapeutics for treating infectious diseases. Also, sequence-specific antimicrobials including gene editing tools such as CRISPR-Cas system and peptide nucleic acids have also gained importance recently for the development of ultra-narrow-spectrum therapeutics. A schematic representation of how these approaches are being used to tackle antibiotic resistance is represented in Fig. 1. These emerging approaches can be the utilized to halt the reversal of our current situation to the pre-antibiotic era and to transform the current resistance era into a narrow-spectrum era in which antibiotic therapy will be more specific and personalized to each individual.
Schematic representation of the various approaches used for combating antibiotic resistance in the recent times. Approaches including unconventional targets of the pathogens, alternative therapies, ultra-narrow-spectrum antimicrobials, revamping the already in-use antibiotics and mining the uncharted microbial territories, work together to halt the progress of antibiotic resistance and transform it into a narrow-spectrum era where antibiotics act specifically to target the drug-resistant pathogens
Targeting the Bacterial Cell Membrane—Antimicrobial Peptides
Antimicrobial peptides (AMPs) are cationic host defense peptides with variable mechanism of actions that are produced ubiquitously among all the kingdoms including protozoa, bacteria, archaea, fungi, plants, and animals. These agents do not target proteins in the bacterial cell, but they act by associating themselves into the membrane leading to perturbation of its bilayer structure. These host defense peptides (HDPs) are major role players in the innate immune system, regulating key features such as the chemokine release from immune cells, wound healing enhancement, anti-apoptotic effect on immune cells, and acting as adjuvants to stimulate the adaptive immune response [19].
The prospect of using these agents for targeting pathogenic microorganisms raises hopes to circumvent the growing resistance against antimicrobial agents because of three major reasons: Firstly, AMP’s act through a physical mode of action attributed to their charged nature enabling membrane disruption. Resistance against this mechanism is rare among bacteria. Secondly, resistance is difficult to develop due to the diverse mode of action of a single peptide, which is dependent on peptide structure, peptide to lipid ratio, and lipid membrane properties. Resistance is hard to develop. Lastly, because of advancements in sequencing technologies and molecular dynamics platforms, these small peptides can be easily modified and developed in silico for use as adjuvants to other antibiotics to enhance their activities and break any developing resistance.
Till date, several different mechanisms of AMPs depicting their action have been reported. Kim et al. [20], evaluated the immunomodulatory activity of chicken NK (cNK) lysin, the chicken homologue of human granulysin. cNK is a cationic amphiphilic antimicrobial peptide (AMP) produced by cytotoxic T cells and natural killer cells and its derivatives, particularly the peptide cNK-2. They recognized the role of AMPs in modulating both innate and adaptive responses. They used the site-directed mutation approach to decipher the structure and function of the peptides. In a relatively recent study, a group from Lanzhou University, China tried the approach of chemical modifications through conjugation of fatty acid chains of varying lengths into peptides. Acetylation, cyclization, and several other reactions produced new and promising peptide derivatives with antimicrobial activity against diverse multidrug-resistant pathogens [21]. In various studies, including that of Malekkhaiat Häffner and Malmsten et al., [22], the impact of the introduction of dimers in AMPs for enhanced activity as well as reduced toxicity was demonstrated. In their study, they successfully showed the peptide self-assembly (micelles, vesicles, nanofibers, nanotubes, etc.) with an increased antimicrobial activity. In another study, Zhu et al. designed analogs of Mastoparan-C an antimicrobial peptide with excellent broad-spectrum activity but is highly toxic to mammalian cells due to high cell membrane perturbation capability. The screened analogs showed remarkably less toxicity and were shown to aid the antibiotics rifampicin and gentamicin in ceasing Gram-negative bacterial infections [23].
Antimicrobial peptides offer hopes for combating antimicrobial resistance, but they face challenges such as high cytotoxicity, high production costs compared to conventional antibiotics such as aminoglycosides, and non-selective membrane disruption. Antimicrobial peptides offer hope for combating antimicrobial resistance, but they face challenges such as high cytotoxicity, high production costs compared to conventional antibiotics such as aminoglycosides, and non-selective membrane disruption. Clinical trials of various antimicrobial peptides such as Nisin, Gramicidin, Polymixins, and others have revealed that in order to improve the efficacy, various strategies such as shortening the peptide length to reduce production costs, searching for optimal delivery systems where the peptides can be encapsulated for better bioavailability, and modifying the peptide sequence for improved bio selectivity must be implemented [24].
Targeting the Bacterial Efflux Mechanism—Efflux Pump Inhibitors
Antibiotic resistance is known to occur through three major processes including drug inactivation, alteration of the target molecules intrinsic to the bacteria, and efflux of drugs preventing their accumulation inside the cell to impede intracellular activity. The efflux pumps present in the cytoplasmic membrane aid in this process by selectively pumping out chemical compounds against its concentration gradient (active mechanism). Although, initially discovered in bacteria, these efflux pumps are prevalent in higher eukaryotes, where they form a major part of the genome sequence. The presence of drug transporters in eukaryotic cells can be explained by the fact that the majority of these transporters have broad specificities and select unrelated classes of compounds. Furthermore, transporters serve as a general barrier against amphiphilic compounds that may cross the cytoplasmic barrier and disturb the osmotic balance of the cell. Advances in structural biology have deciphered the structure of these transporter proteins and their mechanism of transportation which forms a tripartite system that allows the flow of molecules across the cytoplasmic barrier along the periplasmic space of Gram-negative bacteria [25]. Insights into the molecular mechanisms of these transporters have enlightened certain crucial points that can be manipulated in order to develop future drugs targeting these mechanisms. The development of efflux pump inhibitors (EPIs) is imperative when multidrug-resistant bacteria are concerned since efflux pumps form a major determinant for drug resistance. An optimal inhibitor is expected to reduce the intrinsic resistance of the bacteria against antibiotics and other drugs along with curbing the emerging frequency of multidrug-resistant strains. The inhibition of efflux can be accomplished in various ways including (i) downregulating the expression of efflux pump genes by manipulating the genetic regulation, (ii) redesigning previously used antibiotics, (iii) inhibiting the assembly of functional efflux pumps, (iv) avoiding substrate binding to the active site of the efflux pump, and (v) collapsing the energy mechanism responsible for energizing these pumps. The EPIs are molecules that are able to fulfill almost all of the inhibition criteria.
Although there is a myriad of EPIs in the laboratory stage showing promising inhibitory activity, classifying them is a difficult task since the mode of action of many of them is unknown. Nonetheless, depending on the current data regarding the mode of action of EPIs, they can be classified into two broad categories (i) EPIs that work by decoupling the energy from efflux pumps, and (ii) EPIs that directly binds and inhibits the efflux pumps. On the other hand, EPIs can also be classified based on its their source which includes broad categories such as (i) Plant-based EPIs, (ii) Synthetic EPIs, and (iii) EPIs derived from microorganisms [26]. A few examples of the most commonly used EPIs that encompass this broad classification are listed in Table 1.
One of the ways of inhibiting the pumps is blocking the outer membrane factor (OMF) of the tripartite efflux pumps. Two indole derivatives were designed that were capable of blocking TolC, the OMF of different Enterobacteriaceae efflux pumps, resulting in strain sensitization to antibiotics such as chloramphenicol, tetracycline, erythromycin, and ciprofloxacin [27]. In a very recent study, the importance of norA efflux pumps to confer resistance among S. aureus isolates from ciprofloxacin was established, highlighting the importance of its inhibition to control the evolution of resistant strains overexpressing norA pumps, and it has been reported in many studies that plant-based natural products have promising inhibitory effect against norA pumps [28,29,30,31].
Inhibiting the expression of multidrug efflux pumps is also a substantial alternative that can be useful against efflux pumps whose basal expression can contribute to the intrinsic resistance of a bacterium [32]. Short anti-sense oligomers have been used to disrupt the expression of AcrAB-TolC efflux system in E. coli which hyper-sensitizes them to antibiotics [33]. The efficacy of the short oligomer is highly specific to the conserved target sequence of the bacteria and have low detectable toxicity against human cells. In another study, anti-sense peptide nucleic acids were designed to target the three components of the CmeABC efflux pump in Campylobacter jejuni which inhibited its expression and resulted in a sensitive strain against antibiotics [34]. Although, the use of these anti-sense nucleotides has been found to be effective in lab conditions, it is still unknown whether it will be effective in clinical settings or pass the clinical trials.
EPIs promise considerable efficacy in curbing antibiotic resistance primarily by aiding the antibiotics in their mode of action, or by weakening the bacteria before antibiotics take effect. Although experiments have suggested the clinical use of EPIs, a substantial understanding of toxicity is still required to effectively administer EPIs along with antibiotics. But hope has been shown by studies on approved drugs where more than 1200 drugs were screened and a tyrosine kinase inhibitor, nilotinib, was found to be effective in inhibiting the norA efflux pump which confers resistance to S. aureus against fluoroquinolones (e.g., ciprofloxacin) [35].
Phage Therapy
Bacteriophages or “phages” are viruses that can infect and replicate within bacterial cells and exclusively targets bacteria. They form the most abundant and ubiquitous group of organisms on earth that interferes and affects microbial physiology, evolution, population dynamics, and therapeutics [36]. The exclusivity of these phages toward bacteria has been the most important factor that renders them therapeutically important. Although the term bacteriophage coined by Felix d’Herelle consequently led to its use in treating bacterial infections in human patients a century ago, the rise of antibiotics in the mid of 20th century has side-lined the importance of bacteriophages as therapeutic agents. In a review about bacteriophages, Altamirano et al., have compared phage therapy to antibiotics regarding different aspects of its function [37]. This highlighted important points such as the high host specificity of the phages that can be used as an advantage over antibiotics because phages would not exhibit any off-target side effects due to disturbances in the microbiota. The host specificity can also turn disadvantageous in clinical terms, since it demands a thorough diagnosis of the infection and identification of the etiological agent to strain level. Another most important advantage of using phages therapeutically is that, unlike antibiotics, phages will always have the advantage over bacteria as they develop new infectivity and mutate alongside their host. Furthermore, because of their abundance and genetic versatility, there are an infinite number of sources from which they can be isolated and efficiently molded and modified. [37].
The therapeutic use of bacteriophages or “phage therapy” requires some key aspects for it to be used clinically. The regulatory requirement for therapeutic use constitutes strictly lytic phages, confirmed antimicrobial activity against the target pathogen, and removal of contaminating bacterial debris and endotoxins [38]. Consequently, phages are being extensively studied around the world for developing optimized viruses or cocktails of these viruses that can be used clinically. The last decade has witnessed numerous in vitro and preclinical studies that are being considered to enter phase I clinical trials. These trials were based on three types of administration, i.e., topical administration, oral administration, and intravenous administration. Phage therapy has been used to treat chronic bacterial prostatitis, a prolonged bacterial infection caused by many nosocomial pathogens. It was used against a patient with chronic bacterial prostatitis caused by Gram-positive bacteria that was resistant to antibiotic treatment. After the phage treatment, there was a substantial reduction in bacterial load and resolution of the infection [39]. Another interesting use of phages was represented in a recent work by Wang and Loh et al., where they used phages to make A. baumannii strains sensitive to previously resistant strains to colistin. This was done by co-incubating a lytic phage with the A. baumannii strain, resulting in the emergence of phage-resistant A. baumannii which were sensitive to colistin. This aspect where the bacterial pathogen chooses to become resistant to the phages by modifying its capsular material resulting in antibiotic sensitivity can be utilized in clinical settings [40].
It is well established that the phage genomes encode numerous proteins that aid in the breaching of bacterial cell membranes so that the virus can inject their nucleic acids into the host. Two important proteins or enzymes are noteworthy during the infection phase; the virion-associated peptidoglycan hydrolases (VAPGH) and the polysaccharide depolymerases [41]. VAPGH is found in the base plate of the virus that locally targets the peptidoglycan of the bacterial wall allowing the phage tail tube structure to embed and eject its genomic material. Another enzyme, which is the polysaccharide depolymerases, targets the polysaccharide components of the bacterial cell envelope such as the lipopolysaccharide (LPS), the extracellular matrix components in biofilms, bacterial capsule, etc. This exposes the phage-recognized receptors on the bacterial cell surface, allowing phages to identify specific strains [42]. The capsule and biofilm are important virulence components of the bacteria which are utilized either to escape the defense system of the host, or to resist the attack of antibiotics. Thus, using virion-based enzymes such as polysaccharide depolymerases along with antibiotics can give it an extra edge to penetrate the bacteria and exert its effect. Consequently, phage-based depolymerases produced through a monophage therapy along with amoxicillin were used for the treatment of K. pneumoniae infection, where the enzyme was attributed for its extracellular matrix-degrading properties allowing better penetration of the antibiotic as well as whole phage particles [42].
Altogether, the use of phage therapy as an efficient tool to fight against antibiotic resistance appears promising due to the multidimensional strategies provided by these phages. It has been recently pointed out that the interrelationship between bacteria and phages must be understood from an evolutionary perspective, as evidenced by studies conducted in Bangladesh on the endemic cholera caused by Vibrio cholerae [43].
Further research is required to establish phage therapy as a fore-front strategy due to the recurring resistance in bacteria attributed to evolutionary stand-offs. Phage therapy promises hope in these tiring times, when the scientific community values trying out new combinations and methods.
Combinatorial Therapy—Engaging Multiple Targets
Combination of therapies for treatment of diseases has been well established in the clinical practice where conditions such as HIV, tuberculosis, malaria are confronted with one or two combinations of drugs. The previous sections discussed about strategies that target unconventional mechanisms of the pathogens, but combining these strategies to attain a synergy to which the emergence of resistance may be negligible is an interesting approach. This approach has been followed to tackle infections by multidrug-resistant bacteria through either antibiotic–antibiotic combination, or through the combination of antibiotics with adjuvants that either directly target resistance mechanisms or indirectly enhances the effect of antibiotics by inhibiting its efflux or targets bacterial signaling mechanisms [44]. Figure 2 shows a schematic representation of a combination therapy that can be adopted using AMPs, EPIs, and bacteriophages as adjuvants along with commonly used antibiotics.
Schematic representation of the combinatorial approach using a combination of antibiotics or using antibiotics with appropriate adjuvants including antimicrobial peptides, efflux pump inhibitors, and bacteriophages, enhance the performance of the antibiotics for treatment of drug-resistant pathogens
This approach can greatly enhance the efficacy of already in-use antibiotics or can be used to sensitize the bacteria before the action of antibiotics. In the recent times, clinical practice has demonstrated success in the combination of two or more antibiotics or a combination of antibiotics with adjuvants. The combination of antibiotics or other drugs is focused on targeting multiple targets to effectively kill the pathogen leaving less or no chance for the pathogen to develop resistance. The combination of antibiotics can follow three strategies including targeting different pathways, for example, the treatment of Mycobacterium tuberculosis infections was done using four different antibiotics such as isoniazid, rifampicin, ethambutol, and pyrazinamide. Another strategy is to inhibit different targets in the same pathway such as inhibition of successive steps in folic acid synthesis using the combination of sulfamethoxazole and trimethoprim. A third strategy is to inhibit the same target in different ways such as using streptogramins [45].
As previously stated, AMPs are excellent antimicrobial molecules that act by disrupting the bilayer structure of the bacterial cell membrane. Combining AMPs with conventional antibiotics can be a proficient combination therapy because AMPs facilitate the entry of conventional antibiotics. Colistin, a cationic AMP, which is also one of the last resort antibiotics has shown synergy with antibiotics such as azithromycin and has potentiated its entry into bacterial cells to exert its effect against multidrug-resistant Gram-negative bacteria [46]. Another important AMP such as daptomycin has also demonstrated synergistic effects with ampicillin in vancomycin-resistant Enteroccocus faecium (VRE) clinical isolate that can mostly be attributed to the modification of surface charge by the AMP [47].
In case of EPIs, the central role would be to sensitize the resistant bacteria to antibiotics. Keeping this in mind, antibiotic hybrids such as EPI/antibiotic hybrids present a hopeful scenario posing as next-generation agents and adjuvants to control resistance among Gram-negative bacteria [48]. In a study that attempted to conjugate antibiotics with EPIs, tobramycin-EPI conjugates showed synergistic effect with tetracycline antibiotics to enhance its effect and prevent its subsequent sequestration out of the cell by disrupting the proton motif force, which cripples the efflux pumps resulting in sensitization of multidrug-resistant P. aeruginosa [49]. Bacteriophages and their associated enzymes also exhibit the potential to be used in combination therapy along with conventional antibiotics.
Studies have reported the use of antibiotic combinations to inhibit the cell wall formation of Gram-positive bacteria such as Methicilin-Resistant S. aureus MRSA. For example, a natural product, tunicamycin, inhibits the teichoic acid in the Gram-positive bacterial cell wall and demonstrates excellent synergy with β-lactam antibiotics which resulted in several folds decrease in minimum inhibitory concentration (MIC) of oxacillin against MRSA [50]. The target described here is the enzyme N-acetylglucosamine-1-phosphate transferase encoded by the gene tarO, which is the first enzyme in the biosynthesis pathway of wall teichoic acid. Another compound named ticlopidine, which is an antiplatelet drug has been shown to identify the enzyme N-acetylglucosamine-1-phosphate transferase as its target. Ticlopidine showed synergistic effects with cefuroxime against a range of MRSA strains which resulted in a decrease of the MIC concentrations to upto 64-folds [51]. Also, there are several known inhibitors of cell wall synthesis such as bacitracin, teicoplanin, vancomycin, and so on, which shows synergistic effects with β-lactam antibiotics against Gram-positive bacterial strains [52].
Recently, in a study, it was found that anti-virulence compounds such as gallium, a siderophore quencher, and furanone C-30, a quorum-sensing inhibitor, can aid a combination of antibiotics demonstrating synergistic effect against P. aeruginosa [53]. The team used four clinically relevant antibiotics such as ciprofloxacin, colistin, meropenem, and tobramycin to treat drug-resistant bacteria and found promising levels of synergies between certain drug pairs. Also, the anti-virulence compounds acted as potent adjuvants to the antibiotics rendering them as growth inhibitors against certain drug-resistant strains of P. aeruginosa. Similar studies conducted around the world on drug combinations as a potential source of antibiotic strategies gives us hope that in near future, approaches like these may lead to the alleviation of the resistance problem.
Antibodies as Antibiotics
In April 2021, WHO published a list of non-traditional antibacterials, which include alternative strategies to direct acting small molecules, and one of which is the use of antibodies as antibiotics. Before the advent of antibiotics, serum antibodies were the prevalent medication for treating infectious diseases. With the onset of antibiotic resistance, researchers are once again probing into this field and vaccination strategies to treat bacterial infectious diseases.
Antibodies as therapeutic agents utilize the potential of a human body’s immune system. Human monoclonal antibodies are less likely to be cleared by the body and show less toxicity with a higher longevity with a typical half-life of 21 days for IgG types. Another advantage of antibodies is their high specificity, which makes them useful as a therapeutic agent that does not disrupt normal bacterial flora. Taking these advantages into account, various monoclonal antibodies have been developed and are currently undergoing clinical trials. These antibodies have shown to be active against nosocomial pathogens such as S. aureus and P. aeruginosa, and also difficult-to-treat pathogens such as A. baumannii. A human monoclonal antibody developed as such by Trellis Biosciences disrupts the biofilm formation of multiple bacteria by targeting the DNABII protein required for biofilm formation [54]. Another product that is being developed by Roche is an antibody–antibiotic conjugate which is specific to S. aureus. The human monoclonal antibody is designed to bind to the surface of S. aureus putting the conjugated antibiotic in close proximity that enhances the effect of the antibiotic [54]. A bi-specific antibody was developed by the company AstraZeneca PLC (formerly MedImmune), MEDI3902, which targets a surface polysaccharide of P. aeruginosa and affects its biofilm forming capability [54]. Recently, a double-blind, placebo-controlled, phase 2 trial was conducted with suvratoxumab, a human monoclonal antibody capable of controlling ventilator-associated pneumonia caused by S. aureus for the patients in Intensive care units with mechanical ventilators [55]. This monoclonal antibody targets the pore-forming α-toxin and thus greatly reduces its toxic effect.
Although many of the monoclonal antibodies are in the Phase 1 or Phase 2 of the clinical trials, only a handful have the potential to be translated into a marketable product. Further research into the field of antibodies is required to strengthen the hold of monoclonal antibodies against drug resistance since it is less likely to contribute to the increasing antibiotic resistance than antibiotics themselves.
Immunomodulatory Agents as Anti-Bacterial Therapeutics
Harnessing the full potential of a body’s immune system is an interesting approach in the era of antibiotic resistance. Immunomodulating agents are a promising alternative to antibiotics in treating infectious diseases by utilizing the antimicrobial effector mechanisms of the immune cells such as macrophages [56]. The mechanisms through which macrophages eliminate intracellular bacterial infections include human antimicrobial peptides such as cathelicidin, LL-37, nitric oxide (NO) produced by inducible NO synthase (iNOS), and reactive oxygen species (ROS). These effector strategies used by macrophages destroy the cell membrane integrity and interfere with the membrane forming mechanism of the bacteria. For the treatment of tuberculosis, antimycobacterial immunity in the body can be enhanced by many immunomodulatory agents such as arginine, active vitamin D3, 1,25-dihydroxyvitamin D3 (vitD), or histone deacetylase (HDAC) inhibitors such as sodium phenylbutyrate (PBA). A combination of vitD + PBA has been shown to be effective against Mycobacterium tuberculosis through the human antimicrobial peptide (LL-37) pathway [56]. Repurposing existing drugs that modulates the immune system for anti-tuberculosis therapy is also an interesting approach that is being followed [57]. Immunomodulatory drugs that have been repurposed include statins, Non-Steroidal Anti-inflammatory Drugs (NSAIDs), Linezolid, Metformin, and many others.
The role of gut microbiota in modulating the immune response of a body is an attractive approach. The commensal microorganisms present in the digestive tract provide health benefit to the host by maintaining immune homeostasis [58]. This is achieved through mechanisms such as engaging the toll-like receptors and pathogen-specific receptors to the ligands of the commensal microorganisms, which maintains tolerance and immune response against the pathogens Furthermore, the metabolites of the gut microbiota have an immense role in modulating a wide range of responses in the human body by inducing the differentiation and maintenance of lymphocytes, which are effector immune cells. Short chain fatty acids (SCFAs) which includes acetate, propionate, and butyrate, are essential metabolites synthesized by intestinal microbiota and are formed during the bacterial growth process through carbohydrate utilization [59]. The concentration of SCFAs depends substantially on the diet and the state of the microbiota. The SCFAs play an important role in maintaining the proper balance in cases of microbial dysbiosis, i.e., the imbalance of the gut microbiota leading to disease conditions of the intestine.. In cases of microbial infections, SCFAs can directly act by diffusing through the membrane of the bacteria at low pH where it is lipid soluble and once it is inside it disrupts the ionic balance [60]. SCFAs like butyrate and acetate also have an effect on pathogen virulence by lowering the pH, which reduces flagellar motility and the ability to form biofilms. Also, it has been observed that in Salmonella strains, SCFAs impair the production of extracellular components that are required for biofilm formation. It was also observed that butyrate downregulates genes required for invasion and translocation from the intestine to the bloodstream of Salmonella strains [61].
In the era where infectious diseases are getting stronger and more resilient to conventional therapies, strengthening the immune system of the body seems to be a promising endeavor. Microbial metabolites of the gut such as SCFAs represent a hopeful scenario but a deeper understanding of the biological pathways is required for the development of concrete therapies. Meanwhile, prebiotics and probiotics that strengthen the gut microflora and its metabolites appear to be promising for tackling resilient infections.
Together, researchers around the world along with policy makers of numerous countries are acknowledging the dangers of antibiotic resistance and are trying their best to come up with innovative solutions that can mitigate the problem. Targeting these unconventional internal pathogen targets, such as the cell membrane integrity and the efflux pumps, demonstrates efficacy that might lead to victory in the battle against drug-resistant strains proper schemes and trials are devised. Case studies and clinical trials in the case of phage therapy have highlighted the importance of evolutionary advantages that can be exploited for developing efficient therapies and build up on the idea of combination therapies with important phages aiding the already in-use antibiotics.
CRISPR-Cas System for Combating Antibiotic Resistance
Gene editing tools such as the clustered regularly interspaced short palindromic repeats—CRISPR-associated (CRISPR-Cas) system have been recognized as one of the important strategies to combat antibiotic-resistant strains. The programmable nuclease protein (Cas) in the systems allows targeted and sequence-specific DNA cuts, guided by RNA-based spacers (guide RNA) that are flanked by partial repeats. This strategy has been used as a sequence-specific therapeutic to target antibiotic resistance genes in pathogenic strains that act by sensitizing the strains to antibiotics preventing the spread plasmids harboring of resistance genes [62]. The mechanism of action of CRISPR-Cas system can be understood in three stages that include adaptation, expression, and interference [63, 64]. The integration of the invading DNA into the CRISPR locus of the host genome occurs during the adaption stage and then followed by expression of the pre-crRNA and processing of the crRNA from the flanking spacer sequences during the expression stage. During the interference stage, crRNA along with the Cas proteins, detect the invading DNA sequences and produce double-stranded breaks. Based on the molecular characteristics and evolutionary relationships, the CRISPR-Cas system is classified into two classes containing six types and 33 subtypes [63]. The class 1 contains multiple Cas proteins which mediate the interference, compared to the single, large, multidomain crRNA-binding Cas proteins in class 2 (Cas9, Cas12, Cas13, etc.).
The different ways in which CRISPR-Cas can be used to combat antibiotic resistance include the following: (i) by targeting the Cas proteins to species-specific sequences so that only infectious bacteria can be targeted, reducing the loss of normal microbiota, (ii) through direction of the system to cleave drug-resistant genes which eliminate the bacteria harboring them, thus protecting the wild and susceptible ones, and (iii) the system can be used to modify or silence resistance gene harboring plasmids to re-sensitize the bacteria to the antibiotics [65, 66]. These various ways of using CRISPR-Cas in preclinical studies demonstrate the potential of the technology for extensive clinical use after passing through rigorous and transparent clinical testing. The main concern for developing this gene-editing tool as an antimicrobial therapy is its delivery into the bacterial system. One of the ways this can be achieved is through combination with bacteriophages which further potentiates specificity of the strategy [67]. CRISPR-Cas9 system was integrated into a temperate phage specific to S. aureus that has its genome trimmed off of virulence genes but maintaining the host specificity. This greatly increased the specificity of the CRISPR-Cas9 antimicrobial against S. aureus [68]. Phage therapeutics along with CRISPR-Cas system have shown potential in laboratory studies but need to be validated through animal model studies and clinical trials before they can be used as functional clinical therapies. In a study by Cobb et al., engineered temperate phages with integrated CRISPR-Cas9 machinery were helpful in reducing soft tissue infection by S. aureus in mouse models with osteomyelitis and soft tissue infection [69]. In an insightful study, Selle et al., repurposed bacterial endogenous type I-B CRISPR-Cas (CRISPR-Cas3) in Clostridium defficile (C. defficile) into self-targeting CRISPR that targets the bacterial chromosomal genome [70]. This was possible due to an engineered bacteriophage carrying a bacterial genome targeting CRISPR RNAs. This technique was useful in reducing C. defficile infection in both in vitro and in mouse models. In an aim to deliver CRISPR antimicrobials efficiently, Rodrigues et al., used pheromone-responsive conjugative plasmids incorporating type II CRISPR-Cas system to successfully transform Enterococcus faecalis (E. faecalis) targeting specific antibiotic resistance genes [71]. The efficacy of the plasmid containing the CRISPR-Cas system was maintained in animal models where the occurrence of E. faecalis was reduced in the murine intestine.
The future of deploying CRISPR-Cas systems as antimicrobials seems to be promising, but as any other technology, it comes with its own hurdles. The major limitation faced by this technology is the specificity of the guide RNA or crRNA that ultimately defines the action of the system. Thus, further clinical trials are required focusing on the specificity of the system to a particular population of bacteria in a complex microflora environment, to prove the efficacy of this system in actual healthcare settings.
Peptide Nucleic Acids as Sequence-Specific Antimicrobials
Targeted and specific action against drug-resistant pathogens has been the goal for the development of an efficient antimicrobial therapy. Although, this has been achieved to some extent using gene editing tools such as Zinc Finger nucleases, CRIPSR, and engineered bacteriophages, the efficiency of these techniques is yet to reach its clinical potential. Keeping this in mind, another narrow-spectrum therapy has been developed that uses peptide nucleic acids (PNAs) as bactericidal agents [72]. PNAs, although introduced in 1996, has found tremendous applications in the era of antibiotic resistance. PNAs are a group of compounds that interact with its complementary nucleic acids more strongly than a natural nucleic acid and have a modified sugar-phosphate backbone where the sugar and the phosphate bonds have been replaced with repeating N-(2-amino- ethyl) glycine units that are connected through methylene carbonyl linkers. They act as a programmable mimic of DNA that can be introduced in the bacterial cells to specifically silence genes through mechanisms such as transcriptional arrest, P-loop formation, inhibition of replication, translational arrest, etc.
In one of the recent examples, targeting the efaA gene and ftsZ gene controlled the biofilm formation and bacterial growth in E. faecalis using anti-sense PNAs [73]. The PNA was designed to target the start codons of the respective genes for translational arrest. The RNA binding was confirmed through northern-blotting and the PNA was delivered inside the bacteria by electroporation method. In another example, the gene adeB, controlling the RND type efflux pumps, was targeted using the anti-sense PNA technology that controls the RND type efflux pumps which provide resistance to ciprofloxacin [74]. PNA treatment significantly increased the susceptibility to ciprofloxacin where the anti-sense PNA was responsible for the translational arrest of the mRNA transcript of the gene. The engineered PNA was delivered through electroporation.
One of the main constraints of using PNAs as antimicrobial agents is their delivery into the bacterial cells due to its low solubility in water and high molecular weight. This limitation is mostly overcome by modifying the oligonucleotide backbone to increase hydrophilicity or by integrating with molecules that can penetrate cells that can act as transporters. Most studies have integrated cell-penetrating peptides or cationic peptides with PNA to deliver it into the bacterial cells. Barkowski et al., studied the efficiency of various cell-penetrating peptides linked to anti-gyrA PNA on the virulence of Streptococcus pyogenes where it was found that HIV-1 TAT, oligolysine (K8), and (RXR)4XB peptide-coupled anti-gyrA PNAs efficiently abolished bacterial growth in vitro [75]. Abushahba et al., also tested five different cell-penetrating peptides for their efficiency in delivering PNAs into Listeria monocytogenes and in Caenorhabditis elegans infection model [76]. Moreover, the antimicrobial peptides and other cell-penetrating peptides have been shown to be very effective in crossing the cell membrane and delivering the antimicrobials to exert its effect on intracellular pathogens [77].
Although PNAs deliver promising scenarios in developing narrow-spectrum antimicrobials, limitations such as intracellular delivery, low solubility, physiological stability, and clearance from the body still need to be addressed in the coming future to develop this technology as an ultra-narrow-spectrum antimicrobial therapeutics.
Conclusion
Since the emergence of resistance, the focus has been to sensitize the pathogens to antibiotics and stop the emergence of new resistant strains. This course has led to the identification of numerous targets in the pathogens that were not considered in the golden age of antibiotics such as the cell membrane integrity, efflux pumps, and subsequent targets in a single or multiple metabolic pathways. Viruses such as the bacteriophages are being extensively studied so that their inherent ability to specifically infect and kill a bacterium can be harnessed and turned into an efficient therapy. Interesting studies have been conducted on the field of antibodies as therapeutic agents along with immunomodulatory agents which have highlighted the importance of the body’s immune system in combatting serious bacterial infections. Ultra-narrow-spectrum antimicrobials have also been developed using the gene editing tools such as CRISPR-Cas system to specifically silence the resistance genes in the pathogens. PNAs which are oligomers like DNA with a high binding affinity toward complementary DNA have also been used to specifically arrest the expression of virulent genes and genes conferring resistance in pathogens. The clinical trials and case studies supporting these new approaches have also been the source of hope among scientist in the resistance era. Moreover, the researchers are also keen on unearthing novel species from the previously unseen and uncharted territories on land and sea. They have also tried out the various modifications of the existing approaches that have led to the discovery of some of the most important compounds in recent times. Hurdles in antibiotic discovery will continue to trouble researchers in the time to come but approaching the problem with strict policies and frameworks might be able to lend a helping hand and curtail the problem of antibiotic resistance to some extent.
Data Availability
Not applicable.
Code Availability
Not applicable.
References
Dcosta VM, King CE, Kalan L, Morar M, Sung WWL, Schwarz C, Froese D, Zazula G, Calmels F, Debruyne R, Golding GB, Poinar HN, Wright GD (2011) Antibiotic resistance is ancient. Nature 477:457–461
Nesme J, Simonet P (2015) The soil resistome: a critical review on antibiotic resistance origins, ecology and dissemination potential in telluric bacteria. Environ Microbiol 17(4):913–930. https://doi.org/10.1111/1462-2920.12631
Lopatkin AJ, Bening SC, Manson AL, Stokes JM, Kohanski MA, Badran AH, Earl AM, Cheney NJ, Yang JH, Collins JJ (2021) Clinically relevant mutations in core metabolic genes confer antibiotic resistance. Science. https://doi.org/10.1126/SCIENCE.ABA0862
Andersson DI, Hughes D (2011) Persistence of antibiotic resistance in bacterial populations. FEMS Microbiol Rev 35(5):901–911
Laxminarayan R, Duse A, Wattal C, Zaidi AKM, Wertheim HFL, Sumpradit N, Vlieghe E, Hara GL, Gould IM, Goossens H, Greko C, So AD, Bigdeli M, Tomson G, Woodhouse W, Ombaka E, Peralta AQ, Qamar FN, Mir F, Kariuki S, Bhutta ZA, Coates A, Bergstrom R, Wright GD, Brown ED, Cars O (2013) Antibiotic resistance-the need for global solutions. Lancet Infect Dis 13(12):1057–1098
Global shortage of innovative antibiotics fuels emergence and spread of drug-resistance. https://www.who.int/news/item/15-04-2021-global-shortage-of-innovative-antibiotics-fuels-emergence-and-spread-of-drug-resistance. Accessed 20 Oct 2021
2020 antibacterial agents in clinical and preclinical development: an overview and analysis. https://www.who.int/publications/i/item/9789240021303. Accessed 12 Oct 2021
ICMR—Antimicrobial Resistance Research and Surveillance Network Annual Report
CDC (2019) Biggest threats and data|antibiotic/antimicrobial resistance|CDC. 2019 1
Omsland A, Cockrell DC, Howe D, Fischer ER, Virtaneva K, Sturdevant DE, Porcella SF, Heinzen RA (2009) Host cell-free growth of the Q fever bacterium Coxiella burnetii. Proc Natl Acad Sci USA 106(11):4430–4434
Ma L, Kim J, Hatzenpichler R, Karymov MA, Hubert N, Hanan IM, Chang EB, Ismagilov RF (2014) Gene-targeted microfluidic cultivation validated by isolation of a gut bacterium listed in human microbiome project’s most wanted taxa. Proc Natl Acad Sci USA 111(27):9768–9773
Ling LL, Schneider T, Peoples AJ, Spoering AL, Engels I, Conlon BP, Mueller A, Schäberle TF, Hughes DE, Epstein S, Jones M, Lazarides L, Steadman VA, Cohen DR, Felix CR, Fetterman KA, Millett WP, Nitti AG, Zullo AM, Chen C, Lewis K (2015) A new antibiotic kills pathogens without detectable resistance. Nature 517(7535):455–459
Gunjal VB, Thakare R, Chopra S, Reddy DS (2020) Teixobactin: a paving stone toward a new class of antibiotics. J Med Chem 63(21):12171–12195. https://doi.org/10.1021/acs.jmedchem.0c00173
Morris MA, Vallmitjana A, Grein F, Schneider T, Arts M, Jones CR, Nguyen BT, Hashemian MH, Malek M, Gratton E, Nowick JS (2022) Visualizing the mode of action and supramolecular assembly of Teixobactin analogues in Bacillus subtilis. Chem Sci Adv Article. https://doi.org/10.1039/D2SC01388F
Donia MS, Cimermancic P, Schulze CJ, Wieland Brown LC, Martin J, Mitreva M, Clardy J, Linington RG, Fischbach MA (2014) A systematic analysis of biosynthetic gene clusters in the human microbiome reveals a common family of antibiotics. Cell 158(6):1402–1414. https://doi.org/10.1016/j.cell.2014.08.032
Gao W, Zhang L (2020) Nanomaterials arising amid antibiotic resistance. Nat Rev Microbiol 19(1):5–6. https://doi.org/10.1038/s41579-020-00469-5
Makabenta MJ, Nabawy A, Li HC, Schmidt-Malan S, Patel R, Rotello VM (2020) Nanomaterial-based therapeutics for antibiotic-resistant bacterial infections. Nat Rev Microbiol 19(1):23–36. https://doi.org/10.1038/s41579-020-0420-1
Khare T, Anand U, Dey A, Assaraf YG, Chen ZS, Liu Z, Kumar V (2021) Exploring phytochemicals for combating antibiotic resistance in microbial pathogens. Front Pharmacol 12:720726. https://doi.org/10.3389/fphar.2021.720726
Fjell CD, Hiss JA, Hancock REW, Schneider G (2012) Designing antimicrobial peptides: form follows function. Nat Rev Drug Discov 11(1):37–51. https://doi.org/10.1038/nrd3591
Kim WH, Lillehoj HS, Min W (2017) Evaluation of the immunomodulatory activity of the chicken NK-Lysin-derived peptide cNK-2. Sci Rep 7(1):1–11. https://doi.org/10.1038/srep45099
Zhong C, Zhu N, Zhu Y, Liu T, Gou S, Xie J, Yao J, Ni J (2020) Antimicrobial peptides conjugated with fatty acids on the side chain of D-amino acid promises antimicrobial potency against multidrug-resistant bacteria. Eur J Pharm Sci 141:105123. https://doi.org/10.1016/j.ejps.2019.105123
Malekkhaiat Häffner S, Malmsten M (2018) Influence of self-assembly on the performance of antimicrobial peptides. Curr Opin Colloid Interface Sci 38:56–79. https://doi.org/10.1016/j.cocis.2018.09.002
Zhu N, Zhong C, Liu T, Zhu Y, Gou S, Bao H, Yao J, Ni J (2021) Newly designed antimicrobial peptides with potent bioactivity and enhanced cell selectivity prevent and reverse rifampin resistance in Gram-negative bacteria. Eur J Pharm Sci 158:105665. https://doi.org/10.1016/j.ejps.2020.105665
Dijksteel GS, Ulrich MMW, Middelkoop E, Boekema BKHL (2021) Review: lessons learned from clinical trials using antimicrobial peptides (AMPs). Front Microbiol 12:616979. https://doi.org/10.3389/fmicb.2021.616979
Du D, Wang-Kan X, Neuberger A, van Veen HW, Pos KM, Piddock LJV, Luisi BF (2018) Multidrug efflux pumps: structure, function and regulation. Nat Rev Microbiol 16:523–539. https://doi.org/10.1038/s41579-018-0048-6
Sharma A, Gupta VK, Pathania R (2019) Efflux pump inhibitors for bacterial pathogens: from bench to bedside. Indian J Med Res 149(2):129–145. https://doi.org/10.4103/ijmr.IJMR_2079_17
Zeng B, Wang H, Zou L, Zhang A, Yang X, Guan Z (2010) Evaluation and target validation of indole derivatives as inhibitors of the AcrAB-TolC efflux pump. Biosci Biotechnol Biochem 74(11):2237–2241. https://doi.org/10.1271/bbb.100433
Papkou A, Hedge J, Kapel N, Young B, MacLean RC (2020) Efflux pump activity potentiates the evolution of antibiotic resistance across S. aureus isolates. Nat Commun 11(1):1–15. https://doi.org/10.1038/s41467-020-17735-y
Pereira da Cruz R, Sampaio de Freitas T, do Socorro Costa M, Lucas dos Santos AT, Ferreira Campina F, Pereira RLS, Bezerra JWA, Quintans-Júnior LJ, De Souza Araújo AA, De Siqueira Júnior JP, Iriti M, Varoni EM, De Menezes IRA, Melo Coutinho HD, Bezerra Morais-Braga MF (2020) Effect of α-Bisabolol and Its β-Cyclodextrin complex as TetK and NorA efflux pump inhibitors in Staphylococcus aureus strains. Antibiotics 9(1): 28. DOI: https://doi.org/10.3390/antibiotics9010028
Singh S, Kalia NP, Joshi P, Kumar A, Sharma PR, Kumar A, Bharate SB, Khan IA (2017) Boeravinone B, a novel dual inhibitor of NorA bacterial efflux pump of Staphylococcus aureus and human P-glycoprotein, reduces the biofilm formation and intracellular invasion of bacteria. Front Microbiol 8:1868. https://doi.org/10.3389/fmicb.2017.01868
Rezende-Júnior LM, de Andrade LM, S, Leal ALAB, Mesquita AB de S, Santos ALP de A dos, Neto J de SL, Siqueira-Júnior JP, Nogueira CES, Kaatz GW, Coutinho HDM, Martins N, da Rocha CQ, Barreto HM (2020) Chalcones isolated from Arrabidaea brachypoda flowers as inhibitors of NorA and MepA multidrug efflux pumps of Staphylococcus aureus. Antibiotics 9(6):351. https://doi.org/10.3390/antibiotics9060351
Blanco P, Sanz-García F, Hernando-Amado S, Martínez JL, Alcalde-Rico M (2018) The development of efflux pump inhibitors to treat Gram-negative infections. Expert Opin Drug Discov 13:919–931. https://doi.org/10.1080/17460441.2018.1514386
Ayhan DH, Tamer YT, Akbar M, Bailey SM, Wong M, Daly SM, Greenberg DE, Toprak E (2016) Sequence-specific targeting of bacterial resistance genes increases antibiotic efficacy. PLOS Biol 14(9):e1002552. https://doi.org/10.1371/journal.pbio.1002552
Mu Y, Shen Z, Jeon B, Dai L, Zhang Q (2013) Synergistic effects of anti-CmeA and anti-CmeB peptide nucleic acids on sensitizing Campylobacter jejuni to antibiotics. Antimicrob Agents Chemother 57(9):4575–4577. https://doi.org/10.1128/AAC.00605-13
Zimmermann S, Klinger-Strobel M, Bohnert JA, Wendler S, Rödel J, Pletz MW, Löffler B, Tuchscherr L (2019) Clinically approved drugs inhibit the staphylococcus aureus multidrug NorA efflux pump and reduce biofilm formation. Front Microbiol 10:2762. https://doi.org/10.3389/fmicb.2019.02762
Clokie MRJ, Millard AD, Letarov AV, Heaphy S (2011) Phages in nature. Bacteriophage 1(1):31–45. https://doi.org/10.4161/bact.1.1.14942
Gordillo Altamirano FL, Barr JJ (2019) Phage therapy in the postantibiotic era. Clin Microbiol Rev. https://doi.org/10.1128/CMR.00066-18
Young R, Gill JJ (2015) Phage therapy redux–What is to be done? Science 350(6265):1163–1164. https://doi.org/10.1126/science.aad6791
Johri AV, Johri P, Hoyle N, Pipia L, Nadareishvili L, Nizharadze D (2021) Case report: chronic bacterial prostatitis treated with phage therapy after multiple failed antibiotic treatments. Front pharmacol 12:1424. https://doi.org/10.3389/FPHAR.2021.692614/BIBTEX
Wang X, Loh B, Gordillo Altamirano F, Yu Y, Hua X, Leptihn S (2021) Colistin-phage combinations decrease antibiotic resistance in Acinetobacter baumannii via changes in envelope architecture. Emerg Microbes Infect 10(1):2205–2219. https://doi.org/10.1080/22221751.2021.2002671/SUPPL_FILE/TEMI_A_2002671_SM8370.EPS
Roach DR, Leung CY, Henry M, Morello E, Singh D, Di Santo JP, Weitz JS, Debarbieux L (2017) Synergy between the host immune system and bacteriophage is essential for successful phage therapy against an acute respiratory pathogen. Cell Host Microbe 22(1):38-47.e4. https://doi.org/10.1016/j.chom.2017.06.018
Maciejewska B, Olszak T, Drulis-Kawa Z (2018) Applications of bacteriophages versus phage enzymes to combat and cure bacterial infections: an ambitious and also a realistic application? Appl Microbiol Biotechnol 102:2563–2581
LeGault KN, Hays SG, Angermeyer A, McKitterick AC, Johura FT, Sultana M, Ahmed T, Alam M, Seed KD (2021) Temporal shifts in antibiotic resistance elements govern phage-pathogen conflicts. Science 373(6554):eabg2166. https://doi.org/10.1126/science.abg2166
Worthington RJ, Melander C (2013) Combination approaches to combat multidrug-resistant bacteria. Trends Biotechnol 31:177–184. https://doi.org/10.1016/j.tibtech.2012.12.006
Fischbach MA (2011) Combination therapies for combating antimicrobial resistance. Curr Opin Microbiol 14:519–523. https://doi.org/10.1016/j.mib.2011.08.003
Lin L, Nonejuie P, Munguia J, Hollands A, Olson J, Dam Q, Kumaraswamy M, Rivera H, Corriden R, Rohde M, Hensler ME, Burkart MD, Pogliano J, Sakoulas G, Nizet V (2015) Azithromycin synergizes with cationic antimicrobial peptides to exert bactericidal and therapeutic activity against highly multidrug-resistant gram-negative bacterial pathogens. EBioMedicine 2(7):690–698. https://doi.org/10.1016/j.ebiom.2015.05.021
Sakoulas G, Bayer AS, Pogliano J, Tsuji BT, Yang SJ, Mishra NN, Nizet V, Yeaman MR, Moise PA (2012) Ampicillin enhances daptomycin- and cationic host defense peptide-mediated killing of ampicillin- and vancomycin-resistant Enterococcus faecium. Antimicrob Agents Chemother 56(2):838–844. https://doi.org/10.1128/AAC.05551-11
Domalaon R, Idowu T, Zhanel GG, Schweizer F (2018) Antibiotic hybrids: the next generation of agents and adjuvants against gram-negative pathogens? Clin Microbiol Rev. https://doi.org/10.1128/CMR.00077-17
Yang X, Goswami S, Gorityala BK, Domalaon R, Lyu Y, Kumar A, Zhanel GG, Schweizer F (2017) A tobramycin vector enhances synergy and efficacy of efflux pump inhibitors against multidrug-resistant gram-negative bacteria. J Med Chem 60(9):3913–3932. https://doi.org/10.1021/acs.jmedchem.7b00156
Campbell J, Singh AK, Santa Maria JP, Kim Y, Brown S, Swoboda JG, Mylonakis E, Wilkinson BJ, Walker S (2011) Synthetic lethal compound combinations reveal a fundamental connection between wall teichoic acid and peptidoglycan biosyntheses in staphylococcus aureus. ACS Chem Biol 6(1):106–116. https://doi.org/10.1021/cb100269f
Farha MA, Leung A, Sewell EW, D’Elia MA, Allison SE, Ejim L, Pereira PM, Pinho MG, Wright GD, Brown ED (2013) Inhibition of WTA synthesis blocks the cooperative action of PBPS and sensitizes MRSA to β-lactams. ACS Chem Biol 8(1):226–233. https://doi.org/10.1021/cb300413m
Sieradzki K, Tomasz A (1997) Suppression of β-lactam antibiotic resistance in a methicillin-resistant Staphylococcus aureus through synergic action of early cell wall inhibitors and some other antibiotics. J Antimicrob Chemother 39:47–51. https://doi.org/10.1093/jac/39.suppl_1.47
Rezzoagli C, Archetti M, Mignot I, Baumgartner M, Kümmerli R (2020) Combining antibiotics with antivirulence compounds can have synergistic effects and reverse selection for antibiotic resistance in Pseudomonas aeruginosa. PLoS Biol 18(8):e3000805. https://doi.org/10.1371/JOURNAL.PBIO.3000805
Zurawski DV, McLendon MK (2020) Monoclonal antibodies as an antibacterial approach against bacterial pathogens. Antibiotics (Basel) 9(4):155. https://doi.org/10.3390/antibiotics9040155
François B, Jafri HS, Chastre J, Sánchez-García M, Eggimann P, Dequin PF, Huberlant V, Viña Soria L, Boulain T, Bretonnière C, Pugin J, Trenado J, Hernandez Padilla AC, Ali O, Shoemaker K, Ren P, Coenjaerts FE, Ruzin A, Barraud O, Timbermont L, Lammens C, Pierre V, Wu Y, Vignaud J, Colbert S, Bellamy T, Esser MT, Dubovsky F, Bonten MJ, Goossens H, Laterre PF et al (2021) Efficacy and safety of suvratoxumab for prevention of Staphylococcus aureus ventilator-associated pneumonia (SAATELLITE): a multicentre, randomised, double-blind, placebo-controlled, parallel-group, phase 2 pilot trial. Lancet Infect Dis 21(9):1313–1323. https://doi.org/10.1016/S1473-3099(20)30995-6
Rao Muvva J, Ahmed S, Rekha RS, Kalsum S, Groenheit R, Schön T, Agerberth B, Bergman P, Brighenti S (2021) Immunomodulatory agents combat multidrug-resistant tuberculosis by improving antimicrobial immunity. J Infect Dis 224(2):332–344. https://doi.org/10.1093/infdis/jiab100
Fatima S, Bhaskar A, Dwivedi VP (2021) Repurposing immunomodulatory drugs to combat tuberculosis. Front Immunol 12:645485. https://doi.org/10.3389/fimmu.2021.645485
Ranjbar R, Vahdati SN, Tavakoli S, Khodaie R, Behboudi H (2021) Immunomodulatory roles of microbiota-derived short-chain fatty acids in bacterial infections. Biomed Pharmacother 141:111817. https://doi.org/10.1016/j.biopha.2021.111817
Gill PA, van Zelm MC, Muir JG, Gibson PR (2018) Review article: short chain fatty acids as potential therapeutic agents in human gastrointestinal and inflammatory disorders. Aliment Pharmacol Ther 48(1):15–34. https://doi.org/10.1111/apt.14689
Machado MG, Sencio V, Trottein F (2021) Short-chain fatty acids as a potential treatment for infections: a closer look at the lungs. Infect Immun 89(9):e0018821. https://doi.org/10.1128/IAI.00188-21
Lamas A, Regal P, Vázquez B, Cepeda A, Franco CM (2019) Short chain fatty acids commonly produced by gut microbiota influence salmonellaenterica motility, biofilm formation, and gene expression. Antibiotics (Basel) 8(4):265. https://doi.org/10.3390/antibiotics8040265
Bikard D, Euler CW, Jiang W, Nussenzweig PM, Goldberg GW, Duportet X, Fischetti VA, Marraffini LA (2014) Exploiting CRISPR-Cas nucleases to produce sequence-specific antimicrobials. Nat biotechnol 32(11):1146–1150. https://doi.org/10.1038/nbt.3043
Makarova KS, Haft DH, Barrangou R, Brouns SJ, Charpentier E, Horvath P, Moineau S, Mojica FJ, Wolf YI, Yakunin AF, Van Der Oost J (2011) Evolution and classification of the CRISPR–Cas systems. Nat Rev Microbiol 9(6):467–477. https://doi.org/10.1038/nrmicro2577
Gholizadeh P, Aghazadeh M, Asgharzadeh M, Kafil HS (2017) Suppressing the CRISPR/Cas adaptive immune system in bacterial infections. Eur J Clin Microbiol Infect Dis 36(11):2043–2051. https://doi.org/10.1007/s10096-017-3036-2
González de Aledo M, González-Bardanca M, Blasco L, Pacios O, Bleriot I, Fernández-García L, Fernández-Quejo M, López M, Bou G, Tomás M (2021) CRISPR-Cas, a revolution in the treatment and study of ESKAPE infections: pre-clinical studies. Antibiotics 10(7):756. https://doi.org/10.3390/antibiotics10070756
Valderrama JA, Kulkarni SS, Nizet V, Bier E (2019) A bacterial gene-drive system efficiently edits and inactivates a high copy number antibiotic resistance locus. Nat comm 10(1):1–8. https://doi.org/10.1038/s41467-019-13649-6
Gholizadeh P, Köse Ş, Dao S, Ganbarov K, Tanomand A, Dal T, Aghazadeh M, Ghotaslou R, Rezaee MA, Yousefi B, Kafil HS (2020) How CRISPR-Cas system could be used to combat antimicrobial resistance. Infect Drug Resist 13:1111. https://doi.org/10.2147/IDR.S247271
Park JY, Moon BY, Park JW, Thornton JA, Park YH, Seo KS (2017) Genetic engineering of a temperate phage-based delivery system for CRISPR/Cas9 antimicrobials against Staphylococcus aureus. Sci Rep 7(1):1–3. https://doi.org/10.1038/srep44929
Cobb LH, Park J, Swanson EA, Beard MC, McCabe EM, Rourke AS (2019) CRISPR-Cas9 modified bacteriophage for treatment of Staphylococcus aureus induced osteomyelitis and soft tissue infection. PLoS ONE 14(11):e0220421. https://doi.org/10.1371/journal.pone.0220421
Selle K, Fletcher JR, Tuson H, Schmitt DS, McMillan L, Vridhambal GS, Rivera AJ, Montgomery SA, Fortier LC, Barrangou R, Theriot CM (2020) In vivo targeting of Clostridioides difficile using phage-delivered CRISPR-Cas3 antimicrobials. MBio 11(2):e00019-20. https://doi.org/10.1128/mBio.00019-20
Rodrigues M, McBride SW, Hullahalli K, Palmer KL, Duerkop BA (2019) Conjugative delivery of CRISPR-Cas9 for the selective depletion of antibiotic-resistant enterococci. Antimicrob Agents Chemother 63(11):e01454-e1519. https://doi.org/10.1128/AAC.01454-19
Narenji H, Gholizadeh P, Aghazadeh M, Rezaee MA, Asgharzadeh M, Kafil HS (2017) Peptide nucleic acids (PNAs): currently potential bactericidal agents. Biomed Pharmacother 93:580–588. https://doi.org/10.1016/j.biopha.2017.06.092
Narenji H, Teymournejad O, Rezaee MA, Taghizadeh S, Mehramuz B, Aghazadeh M, Asgharzadeh M, Madhi M, Gholizadeh P, Ganbarov K, Yousefi M (2020) Antisense peptide nucleic acids againstftsZ andefaA genes inhibit growth and biofilm formation of Enterococcus faecalis. Microb pathog 139:103907. https://doi.org/10.1016/j.micpath.2019.103907
Abdi SN, Ghotaslou R, Asgharzadeh M, Mehramouz B, Hasani A, Baghi HB, Tanomand A, Narenji H, Yousefi B, Gholizadeh P, Yousefi M (2020) AdeB efflux pump gene knockdown by mRNA mediated peptide nucleic acid in multidrug resistance Acinetobacter baumannii. Microb pathog 139:103825. https://doi.org/10.1016/j.micpath.2019.103825
Barkowsky G, Lemster AL, Pappesch R, Jacob A, Krüger S, Schröder A, Kreikemeyer B, Patenge N (2019) Influence of different cell-penetrating peptides on the antimicrobial efficiency of PNAs in Streptococcus pyogenes. Mol Ther Nucleic Acids 18:444–454. https://doi.org/10.1016/j.omtn.2019.09.010
Abushahba MF, Mohammad H, Thangamani S, Hussein AA, Seleem MN (2016) Impact of different cell penetrating peptides on the efficacy of antisense therapeutics for targeting intracellular pathogens. Sci Rep 6(1):1–2. https://doi.org/10.1038/srep20832
Buccini DF, Cardoso MH, Franco OL (2021) Antimicrobial peptides and cell-penetrating peptides for treating intracellular bacterial infections. Front Cell Infect Microbiol 10:612931. https://doi.org/10.3389/fcimb.2020.612931
Dewanjee S, Dua TK, Bhattacharjee N, Das A, Gangopadhyay M, Khanra R, Joardar S, Riaz M, Feo V, Zia-Ul-Haq M (2017) Natural products as alternative choices for P-glycoprotein (P-gp) inhibition. Molecules 22(6):871. https://doi.org/10.3390/molecules22060871
Acknowledgements
I would like to acknowledge the Department of Biotechnology, Government of India, and the Director of Institute of Advanced Study in Science and Technology, Assam, India (an autonomous institute under Department of Science and Technology, Government of India) for providing the necessary facilities for the work.
Funding
This work was supported by the Department of Biotechnology, Government of India (Grant number BT/PR25029/NER/95/967/2017).
Author information
Authors and Affiliations
Contributions
ANK had the idea for the article. SNH, PB, and ANK performed the literature search and data analysis. and SNH and DT critically revised the work.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical Approval
Not applicable.
Consent for Publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Konwar, A.N., Hazarika, S.N., Bharadwaj, P. et al. Emerging Non-Traditional Approaches to Combat Antibiotic Resistance. Curr Microbiol 79, 330 (2022). https://doi.org/10.1007/s00284-022-03029-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-022-03029-7