Abstract
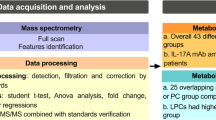
Sorafenib is a multi-kinase inhibitor for treatment of advanced hepatocellular carcinoma (HCC). Beyond its clinical benefit against advanced HCC, the efficacy and safety of sorafenib chemotherapy are critical concerns. In this study, we addressed the lipid profiles associated with the efficacy and safety of sorafenib chemotherapy. Plasma samples from HCC patients before sorafenib chemotherapy (N = 44) were collected and subjected to lipidomic analysis. We measured the levels of 176 lipids belonging to 8 classes of phosphoglycerolipids, 2 classes of sphingolipids, 3 classes of neutral lipids, and 4 other classes of lipids. To characterize lipids associated with efficacy, we compared the responder group (N = 21; partial response and stable disease) with non-responder group (N = 22; progressive disease). To characterize lipids associated with hand–foot skin reaction (HFSR), we compared the susceptible group (N = 12; grade 2 and 3) with non-susceptible group (N = 32; grade 0 and 1). The levels of 8 lipids, including phosphatidylcholine (PC)[34:2], PC[34:3]a, PC[35:2], PC[36:4]a, PC[34:3e], acylcarnitine (Car)[18:0], cholesterol ester[20:2], and diacylglycerol (DG)[34:2], were significantly lower in the responder group, and 6 out of 8 these lipids contained FA(18:2). In addition, the levels of 7 lipids (Car[12:0], Car[18:0], Car[18:1], Car[20:1] and fatty acid amides (FAA[16:0], FAA[18:0], and FAA[18:1]b)) were significantly lower in the group susceptible to HFSR. Our comprehensive lipidomics study using samples from sorafenib-treated patients with HCC revealed that significant differences in the lipid profiles of pre-treatment plasma were associated with sorafenib efficacy and sorafenib-induced HFSR. Validation using another set of patient plasma samples and elucidating the molecular basis of these changes will lead to better treatment with sorafenib chemotherapy.

Similar content being viewed by others
References
Llovet JM, Ricci S, Mazzaferro V, Hilgard P, Gane E, Blanc JF, de Oliveira AC, Santoro A, Raoul JL, Forner A, Schwartz M, Porta C, Zeuzem S et al (2008) Sorafenib in advanced hepatocellular carcinoma. N Engl J Med 359:378–390
Escudier B, Eisen T, Stadler WM, Szczylik C, Oudard S, Siebels M, Negrier S, Chevreau C, Solska E, Desai AA, Rolland F, Demkow T, Hutson TE et al (2007) Sorafenib in advanced clear-cell renal-cell carcinoma. N Engl J Med 356:125–134
Brose MS, Nutting CM, Jarzab B, Elisei R, Siena S, Bastholt L, de la Fouchardiere C, Pacini F, Paschke R, Shong YK, Sherman SI, Smit JW, Chung J et al (2014) Sorafenib in radioactive iodine-refractory, locally advanced or metastatic differentiated thyroid cancer: a randomised, double-blind, phase 3 trial. Lancet 384:319–328
Furuse J, Ishii H, Nakachi K, Suzuki E, Shimizu S, Nakajima K (2008) Phase I study of sorafenib in Japanese patients with hepatocellular carcinoma. Cancer Sci 99:159–165
Nakazawa T, Hidaka H, Takada J, Okuwaki Y, Tanaka Y, Watanabe M, Shibuya A, Minamino T, Kokubu S, Koizumi W (2013) Early increase in α-fetoprotein for predicting unfavorable clinical outcomes in patients with advanced hepatocellular carcinoma treated with sorafenib. Eur J Gastroenterol Hepatol 25:683–699
Lipworth AD, Robert C, Zhu AX (2009) Hand–foot syndrome (hand-foot skin reaction, palmar-plantar erythrodysesthesia): focus on sorafenib and sunitinib. Oncology 77:257–271
Miller KK, Gorcey L, McLellan BN (2014) Chemotherapy-induced hand-foot syndrome and nail changes: a review of clinical presentation, etiology, pathogenesis, and management. J Am Acad Dermatol 71:787–794
Li Y, Gao ZH, Qu XJ (2015) The adverse effects of sorafenib in patients with advanced cancers. Basic Clin Pharmacol Toxicol 116:216–221
Lee YS, Kim BH, Kim BC, Shin A, Kim JS, Hong SH, Hwang JA, Lee JA, Nam S, Lee SH, Bhak J, Park JW (2015) SLC15A2 genomic variation is associated with the extraordinary response of sorafenib treatment: whole-genome analysis in patients with hepatocellular carcinoma. Oncotarget 6:16449–16460
Sakai K, Takeda H, Nishijima N, Orito E, Joko K, Uchida Y, Izumi N, Nishio K, Osaki Y (2015) Targeted DNA and RNA sequencing of fine-needle biopsy FFPE specimens in patients with unresectable hepatocellular carcinoma treated with sorafenib. Oncotarget 6:21636–21644
Han X, Gross RW (2003) Global analyses of cellular lipidomes directly from crude extracts of biological samples by ESI mass spectrometry: a bridge to lipidomics. J Lipid Res 44:1071–1079
Watson AD (2006) Thematic review series: systems biology approaches to metabolic and cardiovascular disorders. Lipidomics: a global approach to lipid analysis in biological systems. J Lipid Res 47:2101–2111
Taguchi R, Nishijima M, Shimizu T (2007) Basic analytical systems for lipidomics by mass spectrometry in Japan. Methods Enzymol 432:185–211
Mené P, Simonson MS, Dunn MJ (1989) Phospholipids in signal transduction of mesangial cells. Am J Physiol 256:F375–F386
Hannun YA, Linardic CM (1993) Sphingolipid breakdown products: anti-proliferative and tumor-suppressor lipids. Biochim Biophys Acta 1154:223–236
Kolesnick RN, Haimovitz-Friedman A, Fuks Z (1994) The sphingomyelin signal transduction pathway mediates apoptosis for tumor necrosis factor, Fas, and ionizing radiation. Biochem Cell Biol 72:471–474
Chen S, Yin P, Zhao X, Xing W, Hu C, Zhou L, Xu G (2013) Serum lipid profiling of patients with chronic hepatitis B, cirrhosis, and hepatocellular carcinoma by ultra fast LC/IT-TOF MS. Electrophoresis 34:2848–2856
Qiu JF, Zhang KL, Zhang XJ, Hu YJ, Li P, Shang CZ, Wan JB (2015) Abnormalities in plasma phospholipid fatty acid profiles of patients with hepatocellular carcinoma. Lipids 50:977–985
Li J, Hu C, Zhao X, Dai W, Chen S, Lu X, Xu G (2013) Large-scaled human serum sphingolipid profiling by using reversed-phase liquid chromatography coupled with dynamic multiple reaction monitoring of mass spectrometry: method development and application in hepatocellular carcinoma. J Chromatogr A 1320:103–110
Huang X, Zeng J, Zhou L, Hu C, Yin P, Lin X (2016) A new strategy for analyzing time-series data using dynamic networks: identifying prospective biomarkers of hepatocellular carcinoma. Sci Rep 6:32448
Saito K, Ohno Y, Saito Y (2017) Enrichment of resolving power improves ion-peak quantification on a lipidomics platform. J Chromatogr B Anal Technol Biomed Life Sci 1055–1056:20–28
Fabris C, Federico E, Soardo G, Falleti E, Pirisi M (1997) Blood lipids of patients with chronic hepatitis: differences related to viral etiology. Clin Chim Acta 261:159–165
Qu F, Zheng SJ, Wu CS, Jia ZX, Zhang JL, Duan ZP (2014) Lipidomic profiling of plasma in patients with chronic hepatitis C infection. Anal Bioanal Chem 406:555–564
Zheng SJ, Qu F, Li JF, Zhao J, Zhang JY, Liu M, Ren F, Chen Y, Zhang JL, Duan ZP (2015) Serum sphingomyelin has potential to reflect hepatic injury in chronic hepatitis B virus infection. Int J Infect Dis 33:149–155
Wu T, Zheng X, Yang M, Zhao A, Li M, Chen T, Panee J, Jia W, Ji G (2017) Serum lipid alterations identified in chronic hepatitis B, hepatitis B virus-associated cirrhosis and carcinoma patients. Sci Rep 7:42710
Sari ME, Yalcin İ, Sahin H, Meydanli MM, Gungor T (2017) Risk factors for paraaortic lymph node metastasis in endometrial cancer. Int J Clin Oncol 22:937–944
Matsuoka T, Adair JE, Lih FB, Hsi LC, Rubino M, Eling TE, Tomer KB, Yashiro M, Hirakawa K, Olden K, Roberts JD (2010) Elevated dietary linoleic acid increases gastric carcinoma cell invasion and metastasis in mice. Br J Cancer 103:1182–1191
Liu L, Cao Y, Chen C, Zhang X, McNabola A, Wilkie D, Wilhelm S, Lynch M, Carter C (2006) Sorafenib blocks the RAF/MEK/ERK pathway, inhibits tumor angiogenesis, and induces tumor cell apoptosis in hepatocellular carcinoma model PLC/PRF/5. Cancer Res 66:11851–11858
Liang Y, Chen J, Yu Q, Ji T, Zhang B, Xu J, Dai Y, Xie Y, Lin H, Liang X, Cai X (2017) Phosphorylated ERK is a potential prognostic biomarker for sorafenib response in hepatocellular carcinoma. Cancer Med 6:2787–2795
Oláh A, Ambrus L, Nicolussi S, Gertsch J, Tubak V, Kemény L, Soeberdt M, Abels C, Bíró T (2016) Inhibition of fatty acid amide hydrolase exerts cutaneous anti-inflammatory effects both in vitro and in vivo. Exp Dermatol 25:328–330
Di Marzo V (2008) Targeting the endocannabinoid system: to enhance or reduce? Nat Rev Drug Discov 7:438–455
Dai CY, Yeh ML, Huang CF, Hou CH, Hsieh MY, Huang JF, Lin IL, Lin ZY, Chen SC, Wang LY, Chuang WL, Yu ML, Tung HD (2015) Chronic hepatitis C infection is associated with insulin resistance and lipid profiles. J Gastroenterol Hepatol 30:879–884
Cassol E, Misra V, Holman A, Kamat A, Morgello S, Gabuzda D (2013) Plasma metabolomics identifies lipid abnormalities linked to markers of inflammation, microbial translocation, and hepatic function in HIV patients receiving protease inhibitors. BMC Infect Dis 13:203
Oberbach A, Blüher M, Wirth H, Till H, Kovacs P, Kullnick Y, Schlichting N, Tomm JM, Rolle-Kampczyk U, Murugaiyan J, Binder H, Dietrich A, von Bergen M (2011) Combined proteomic and metabolomic profiling of serum reveals association of the complement system with obesity and identifies novel markers of body fat mass changes. J Proteome Res 10:4769–4788
Miolo G, Muraro E, Caruso D, Crivellari D, Ash A, Scalone S, Lombardi D, Rizzolio F, Giordano A, Corona G (2016) Pharmacometabolomics study identifies circulating spermidine and tryptophan as potential biomarkers associated with the complete pathological response to trastuzumab-paclitaxel neoadjuvant therapy in HER-2 positive breast cancer. Oncotarget 7:39809–39822
Yu Z, Zhai G, Singmann P, He Y, Xu T, Prehn C, Römisch-Margl W, Lattka E, Gieger C, Soranzo N, Heinrich J, Standl M, Thiering E et al (2012) Human serum metabolic profiles are age dependent. Aging Cell 11:960–967
Menni C, Zhai G, Macgregor A, Prehn C, Römisch-Margl W, Suhre K, Adamski J, Cassidy A, Illig T, Spector TD, Valdes AM (2013) Targeted metabolomics profiles are strongly correlated with nutritional patterns in women. Metabolomics 9:506–514
Ishikawa M, Maekawa K, Saito K, Senoo Y, Urata M, Murayama M, Tajima Y, Kumagai Y, Saito Y (2014) Plasma and serum lipidomics of healthy white adults shows characteristic profiles by subjects’ gender and age. PLoS One 9:e91806
Acknowledgements
The authors would like to thank Ms. Katsuko Toyoshima, Ms. Mai Kojima, and Mr. Ryota Iiji (National Institute of Health Sciences) for experimental assistance and Ms. Chie Sudo for secretarial assistance. This work was financially supported by the Japan Agency for Medical Research and Development (AMED) (Grant Numbers 17mk0101045j0303 and 17ak0101043j0603). This research was also supported by funding from Bayer Yakuhin, Ltd., under a research contract.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
Shunsuke Kondo has received research funding from AstraZeneca, Eli Lilly Japan K.K., ASLAN, Pfizer, Takeda Yakuhin, Ltd., and Bayer Yakuhin, Ltd. None of the other authors have potential conflicts of interest to declare.
Additional information
Corresponding authors for lipidomics (Y. Saito) and clinical significance (S. Kondo).
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Saito, K., Ikeda, M., Kojima, Y. et al. Lipid profiling of pre-treatment plasma reveals biomarker candidates associated with response rates and hand–foot skin reactions in sorafenib-treated patients. Cancer Chemother Pharmacol 82, 677–684 (2018). https://doi.org/10.1007/s00280-018-3655-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00280-018-3655-z