Abstract
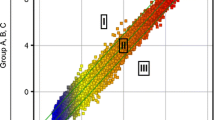
The goal of this study was to identify genes consistently differentially expressed in multiple pairs of isogenic cisplatin (DDP)-sensitive and resistant human ovarian carcinoma cell lines using microarray-based expression profiling. Expression profiling was carried out on six pairs of ovarian carcinoma cells lines growing under identical conditions; each cell expression profile was independently replicated six times. No genes were differentially expressed in all six pairs of cells or even in even in any five of the six pairs. Eighteen genes and 1 EST were upregulated, and four genes and 1 EST were downregulated, in at least four cell pairs. Of these, only metallothionein 2A has previously been implicated in DDP resistance. Among the genes identified on the basis of six replicates, an average of 24.8% would have been missed if only five replicates had been performed, and 38.3% would have been missed with only four replicates. The genes did not identify a dominant biochemical pathway or ontology category as being linked to DDP resistance; however, hierarchical clustering provided evidence for two classes DDP-resistant phenotypes within which there are additional cell pair-specific alterations. Many of the genes identified in this study play important roles in cell surface interactions and trafficking pathways not previously linked to DDP resistance. The genes discovered by this extensively replicated analysis are candidates for prediction of DDP responsiveness in ovarian cancer patients.



Similar content being viewed by others
Abbreviations
- SAM:
-
Significance analysis of microarrays
- DDP:
-
Cisplatin
References
Alldridge LC, Bryant CE (2003) Annexin 1 regulates cell proliferation by disruption of cell morphology and inhibition of cyclin D1 expression through sustained activation of the ERK1/2 MAPK signal. Exp Cell Res 290:93–107
Beffert U, Stolt PC, Herz J (2004) Functions of lipoprotein receptors in neurons. J Lipid Res 45:403–409
Croix BS, Rak JW, Kapitain S, Sheehan C, Graham CH, Kerbel RS (1996) Reversal by hyaluronidase of adhesion-dependent multicellular drug resistance in mammary carcinoma cells. J Natl Cancer Inst 88:1285–1296
Frisch SM, Francis H (1994) Disruption of epithelial cell–matrix interactions induces apoptosis. J Cell Biol 124:619–626
Holford J, Beale PJ, Boxall FE, Sharp SY, Kelland LR (2000) Mechanisms of drug resistance to the platinum complex ZD0473 in ovarian cancer cell lines. Eur J Cancer 36:1984–1990
Ihaka R, Gentleman R (1996) R: a language for data analysis and graphics. J Comput Graph Stats 5:299–314
Jensen R, Glazer PM (2004) Cell-interdependent cisplatin killing by Ku/DNA-dependent protein kinase signaling transduced through gap junctions. Proc Natl Acad Sci USA 101:6134–6139
Lee SH, Zhang W, Choi JJ, Cho YS, Oh SH, Kim JW, Hu L, Xu J, Liu J, Lee JH (2001) Overexpression of the thymosin beta-10 gene in human ovarian cancer cells disrupts F-actin stress fiber and leads to apoptosis. Oncogene 20:6700–6706
Liang XJ, Shen DW, Gottesman MM (2004) Down-regulation and altered localization of gamma-catenin in cisplatin-resistant adenocarcinoma cells. Mol Pharmacol 65:1217–1224
Mishima M, Samimi G, Kondo A, Lin X, Howell SB (2002) The cellular pharmacology of oxaliplatin resistance. Eur J Cancer 38:1405–1412
Rozen S, Skaletsky HJ (2000) Bioinformatics methods and protocols: methods in molecular biology. In: Krawetz S (ed) Primer3 on the WWW for general users and for biologist programmers. Humana, NJ
Safaei R, Howell SB (2005) Copper transporters regulate the cellular pharmacology and sensitivity to Pt drugs. Crit Rev Oncol Hematol 53:13–23
Shain KH, Dalton WS (2001) Cell adhesion is a key determinant in de novo multidrug resistance (MDR): new targets for the prevention of acquired MDR. Mol Cancer Ther 1:69–78
Siddik ZH (2003) Cisplatin: mode of cytotoxic action and molecular basis of resistance. Oncogene 22:7265–7279
Soccio RE, Adams RM, Romanowski MJ, Sehayek E, Burley SK, Breslow JL (2002) The cholesterol-regulated StarD4 gene encodes a Star-related lipid transfer protein with two closely related homologues, StarD5 and StarD6. Proc Natl Acad Sci USA 99:6943–6948
Storey JD, Tibshirani R (2003) SAM thresholding and false discovery rates for detecting differential gene expression in DNA microarrays. In: Parmigiani G, Garrett ES, Irizarry RA, Zeber SL (eds) The analysis of gene expression data: methods and software. Springer, Berlin Heidelberg New York
Tusher VG, Tibshirani R, Chu G (2001) Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA 98:5116–5121
Van Itallie CM, Fanning AS, Anderson JM (2003) Reversal of charge selectivity in cation or anion-selective epithelial lines by expression of different claudins. Am J Physiol Renal Physiol 285:F1078–1084
Wang Y, Serfass L, Roy MO, Wong J, Bonneau AM, Georges E (2004) Annexin-I expression modulates drug resistance in tumor cells. Biochem Biophys Res Commun 314:565–570
Yang CP, Galbiati F, Volonte D, Horwitz SB, Lisanti MP (1998) Upregulation of caveolin-1 and caveolae organelles in Taxol-resistant A549 cells. FEBS Lett 439:368–372
Yang Y, Dudoit S, Luu P, Lin D, Peng V, Ngai J, Speed T (2002) Normalization for cDNA microarray data: a robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res 30:e15–200
Acknowledgement
Supported by grants CA78648 and CA95298 from the National Institutes of Health, a post-doctoral fellowship grant from the Marsha Rivkin Foundation, and a grant from the Research Foundation. The authors express their appreciation to Dr. Bernard Palsson for use of the scanner and to Michael Fero and the staff of the Stanford Functional Genomics Facility for supplying the human cDNA microarrays used for this study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cheng, T.C., Manorek, G., Samimi, G. et al. Identification of genes whose expression is associated with cisplatin resistance in human ovarian carcinoma cells. Cancer Chemother Pharmacol 58, 384–395 (2006). https://doi.org/10.1007/s00280-005-0171-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00280-005-0171-8