Abstract
In general, males mate with multiple females to increase individual reproductive success. Whether or not, and under what circumstances, females benefit from multiple mating has been less clear. Our review of 154 studies covering 184 populations of amphibians and reptiles showed that polyandry was widespread and variable among and within taxonomic groups. We investigated whether amphibian and reptile females had greater reproductive output as the number of sires for offspring increased. Meta-analysis revealed significant heterogeneity in the dataset of all taxa. Expected heterozygosity was a significant moderator (covariate) of positive relationships between female reproductive output and the number of sires, but a sensitivity test showed the result was tenuous. Significant heterogeneity remained despite controlling for expected heterozygosity and other variables but was resolved for most taxonomic groups with subgroup meta-analyses. Subgroup meta-analyses showed that only female salamanders (Caudata) had significantly greater reproductive output with an increased number of sires. For many species of Caudata, males cannot coerce females into accepting spermatophores. We therefore suggest that if females control the number of matings, they can use polyandry to increase their fitness. Caudata offers ideal models with which to test this hypothesis and to explore factors enabling and maintaining the evolution of female choice. Outstanding problems may be addressed by expanding taxonomic coverage and data collection and improving data reporting.
Significance Statement
Many factors and combinations of factors drive polyandry. Whether or not females benefit from mating with more than one male remains equivocal. Focusing on amphibians and reptiles, our analyses demonstrate that female salamanders produced more offspring when mated with multiple males, whereas this was not the case for reptiles. Unlike many other species in our dataset, the polyandrous female salamanders fully control sperm intake and have chosen to mate multiple times. We further highlight problems and key directions for future research in the field.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mating is costly for the participants, so convention is that the more highly invested sex (usually the female) should restrict mating to the minimum required for fertilisation whereas the less constrained sex (usually the male) should benefit from mating with many individuals (Bateman 1948; Trivers 1972). However, the theory that polyandry (females mating with more than one male in the same breeding season; see Table 1) has no fitness benefit for females is questionable because it does not explain why polyandry occurs for so many species (Slatyer et al. 2012; Taylor et al. 2014). Polyandry can be simultaneous, when sperm from different males fertilise the same clutch or litter to result in multiple paternity, or sequential, in which offspring partitioned into separate clutches are fertilised by different males (Table 1). Molecular-based paternity testing within and between the clutches or litters of offspring is a means of detecting multiple sires. Incidence of multiple paternity in clutches or litters is used to indicate incidence of polyandry, but is not accurate if not all mating events resulted in fertilisation (Bretman and Tregenza 2005). Additionally, polyandry resulting in different sires between the clutches or litters of the same female will not be detected by assessing the paternity of only one clutch or litter. However, multiple paternity can only happen if the female had mated with at least two different males at some point in the past. Thus, even if the correspondence may not be one-to-one, in general, the incidence of multiple paternity should scale with the incidence of polyandry—there is, for example, evidence that multiple paternity increases with mate availability (Jensen et al. 2006; Sandrin et al. 2015). Multiple paternity detected for many species is evidence that polyandry is a common behaviour, even for socially monogamous species (see Supporting Information SI1, Table S1).
To explain widespread polyandry, mating with multiple males is suggested to benefit females despite the well-known costs of mating (Table 2 (adaptive explanations), also see Olsson and Madsen 2001; Uller and Olsson 2008; Parker and Birkhead 2013; Forstmeier et al. 2014). Having sperm from multiple males increases the diversity of offspring (Jennions and Petrie 2000) and allows for insurance against male infertility and/or bet-hedging against incompatible matings (Garcia-Gonzalez et al. 2015), sperm competition (Friesen et al. 2020) and post-copulatory mate choice by females (cryptic female choice) (Birkhead 2000; Firman et al. 2017). Genetically acquired benefits may further manifest as increased offspring viability (Olsson and Madsen 2001). Indeed, one view is that multiple mating is nearly always beneficial to females but is logistically constrained by mate acquisition (Avise and Liu 2011). Notwithstanding these hypotheses, the question of whether or not females benefit from polyandry still stands, because even with the focus in behavioural ecology shifting from the male to the female perspective (e.g. Lyons et al. 2021), there remains no consensus on the matter, with reviews of various taxonomic groups and topics reaching equivocal or different conclusions (SI1, Table S1).
Ultimately, to demonstrate that a female benefits from polyandry, we need to show that her fitness increases. Sensu stricto, an individual’s fitness is defined in terms of all of its descendants, but lifetime reproductive success (all progeny produced in her lifetime) is considered an appropriate, and a more readily quantifiable, proxy (Brommer et al. 2004). Still, collecting data on lifetime productivity is more suited for laboratory settings and difficult to obtain for individuals in wild populations, especially when they may be long lived. A more tractable indicator is the reproductive output of one breeding event, which contributes to individual fitness irrespective of the mechanisms involved in producing that output. A short-term measure of reproductive output may still provide a broad indication of total lifetime reproductive success (Nguyen and Moehring 2015). If polyandry has fitness benefits for females, then, regardless of the drivers, we may expect the number of offspring (reproductive output) of those females to increase with the number of sires.
Nonetheless, a key problem is that empirical evidence of females benefiting from polyandry has been hard to find (e.g. Olsson and Madsen 2001; Noble et al. 2013); therefore, alternative explanations have been proposed (Table 2 (neutral (null) explanations)). One is sexual conflict, whereby males coerce females into extra matings (Boulton et al. 2018). Male–male competition may be particularly influential in driving polyandry when density is high, and sex ratio is male-biased (Lyons et al. 2021). Females may also mate according to the mate encounter rate of the population (e.g. Lee et al. 2018), which may be likely where mates are limited or if females cannot predict whether they are likely to meet further mates (Gowaty 2013; Kokko and Mappes 2013).
Various factors may affect the incidence of polyandry. For example, with regard to female fecundity, species with larger brood sizes have greater numerical opportunity for multiple paternity compared with those with smaller brood sizes (Avise and Liu 2011; Correia et al. 2021; Lyons et al. 2021). Other factors may be environmental: mating incidence may vary with latitude because temperature influences longevity, breeding seasonality and mate encounter rates (Olsson et al. 2011; Taylor et al. 2014). Some studies have tested offspring and maternal phenotypic characteristics, offspring heterozygosity, rates of offspring survival or viability and population factors (e.g. density) as possible covariates (Table 2). However, given the variability of behavioural characteristics across species and how the optimum mating strategy at any given time is conditional on population and environmental and individual circumstances (Gowaty 2013; Lyons et al. 2021), it is unlikely for there to be only a single driving factor, even for the same species (Lyons et al. 2021). Variables potentially affecting polyandry become increasingly difficult to resolve as species have more complicated mating behaviours and ecologies.
Focusing on amphibians and reptiles, our aim was to address the question of whether or not females have greater reproductive output if they have more mates. The causes of polyandry may potentially be more easily teased apart for these taxa compared with vertebrates that have more complex and obligate social and mating behaviours (e.g. mammals and birds) (Uller and Olsson 2008). However, two reviews of anuran amphibians and reptiles published over a decade ago used descriptive approaches to conclude that there was little evidence that females of these species benefited from polyandry (Uller and Olsson 2008; Roberts and Byrne 2011). Here, we extended previous reviews with updated literature. Further, we extended previous approaches by taking an empirical approach in examining whether or not female reproductive output increased with polyandry. We used meta-analytical methods to summarise and organise information and to test the relationship between the number of offspring and the number of sires. To compare results from different studies, we used a standardised effect size (a statistic reflecting the strength of the relationship between the variables; Borenstein et al. 2009). We also tested a number of methodological, environmental and biological variables as possible moderators (covariates) of the effect sizes.
Methods
Assembling the global dataset and extraction of data
We searched Web of Science and Scopus through to August 2019 (SI1, Fig. S1). We limited searches to peer-reviewed journals and to studies that used molecular data for paternity analyses. We considered only studies based on amphibians or reptiles that could express natural mating behaviour. Experimental studies were excluded as they may not reflect natural patterns of polyandry and constitute a different body of literature (examples in SI1, Table S1, and see Hettyey et al. 2010). We accepted studies of reintroduced/translocated populations and of wild-caught individuals temporarily held in captivity, where individuals had mated naturally (details in SI1).
We extracted data on multiple paternity and on numbers of sires and offspring in clutches or litters from the text or tables of studies. If presented in graphs, values were extracted using DataThief (Tummers 2006) or the R package metaDigitise (Pick et al. 2019). As often as possible, two people independently extracted data. Otherwise, the same person re-extracted data at least one other time, but on a different occasion (i.e. not immediately after the first extraction).
We used the multiple paternity data to estimate the percentage of females in the population with evidence of polyandry. A sample for the prevalence of polyandry (hereafter ‘PA’) was one group of offspring produced in the same reproductive season by a female. We pooled multiple paternity data for cases where multiple clutches or litters in the same breeding season were sampled for the same female (see SI1 for worked examples). However, offspring produced in different breeding seasons were considered independent samples, because many factors such as maternal body condition, environmental conditions and mate or resource availability could have changed between seasons, with the differences likely more pronounced between rather than within the same season.
To avoid over-representation of well-studied populations, we only had a single PA estimate for any one population (called a ‘case’, K). Data across consecutive breeding seasons were combined. For example, if polyandry was found for 2 of 10 samples in one season and 4 of 14 samples in another season, PA would be estimated as 25% (6 out of 24 samples with evidence of polyandry). However, if different data were collected with an interval of several years, if the studies used different methods or if some of the same data was used in different studies, we chose the data from the study that was more recent, reported more samples and/or used more advanced methods (see Supplementary Data, SI2).
Preliminary analyses led to exclusion of cases with fewer than five samples for analyses involving the PA variable. Cases with small sample sizes inflated the frequency of either 0% or 100% values for PA in the dataset (see SI1, Fig. S2; note that SI2 lists all cases, including those with fewer than five samples).
We also extracted data on a number of variables previously suggested to affect polyandry: latitude, average heterozygosity (observed and expected), average clutch or litter sizes and indicators of the sampling effort (e.g. number of loci, average number and average percentage of offspring genotyped). Data on average heterozygosity (observed and expected) were those estimated for the adult population and limited to cases using microsatellite loci for parentage and/or population genetic analysis. We tested sampling effort because methodological differences between studies may introduce error (Borenstein et al. 2009). For example, evidence of polyandry may become easier to detect as sampling effort increases (Griffith et al. 2002). In some cases, the data for variables were from associated literature (see SI1 for details) or missing. Cases with a missing variable were excluded from analyses involving the missing variable.
Datasets
Summary statistics were estimated for all data and for data in major taxonomic groups: Anura (frogs and toads), Caudata (newts and salamanders), Crocodylia (crocodiles and alligators), Lacertilia (lizards, skinks, geckos, iguanas and dragons), Serpentes (snakes) and Testudines (tortoises and turtles). We also obtained summary statistics for Cheloniidae (marine turtles, a family of Testudines) because this was the only family with sufficient cases to summarise the data for genera. Lacertilia and Serpentes are squamates, but we treated them separately as they are groups with distinct biological differences and large membership. Included in the summary statistics of the full dataset were data for single representatives of their respective taxonomic groups: the caecilian Boulengerula taitanus (Gymnophiona, Herpelidae) and the tuatara Sphenodon punctatus (Rhynchocephalia, Sphenodontidae).
Effect size data
We estimated the standardised effect size, Fisher’s z-transformation of the correlation coefficient (Zr). We calculated the correlation coefficient r (Pearson’s r) between the number of offspring and the number of sires. If there was only summary information (means, variances and sample sizes), we used the escalc() function of the R package metafor (Viechtbauer 2010) to calculate the bias-corrected standardised mean difference (Hedges’ g) between singly sired and multiply sired groups of offspring, which was then converted to correlation coefficient (r). We used reported r values if raw or summary data were not available. We then converted r values and their sample sizes to Zr and its variance (VarZr) (Borenstein et al. 2009) (see SI1 for formulae).
Some studies tested for relationships between the number of offspring and the number of sires but did not include the information required for effect size calculation (SI1, Table S2). These studies used a variety of tests: t tests, ANOVA, ANCOVA, GLM, GLMM and logistic regression. While it may be possible to convert between some types of effect sizes (Rosenberg et al. 2000), converting the results of statistical models that include multiple variables or tests that control for covarying factors may not be appropriate. Conversions make assumptions about the underlying data, and if these assumptions were violated or if there were substantive differences between the studies, then the validity of the meta-analysis will be impacted (Borenstein et al. 2009). We preferred to take a conservative approach by excluding these studies.
Statistical analyses
We performed statistical analyses in R v. 3.6.1 (R Core Team 2019) using R Studio v. 1.1.453 (RStudio Team 2015) and used the R package metafor (Viechtbauer 2010) for meta-analyses. The multilevel meta-analytical framework in metafor fits random-effects models weighted by the inverse of the marginal variance–covariance matrix estimated for the effect sizes (Viechtbauer 2010). We used the rma.mv() function to build random-effects meta-models with the default restricted maximum likelihood estimation (REML) method and the option ‘test’ set to ‘t’ (see SI1 for R scripts). Random effects were the statistic used for effect size calculation (correlation r or Hedges’ g), common ancestry (phylogeny) and estimates from the same study or from the same species. Funnel plots (the funnel() function), Egger’s tests (the regtest() function) and estimation of ‘fail-safe’ numbers (the fsn() function) enabled the examination of the model results for the presence of heterogeneity and publication bias (Rothstein et al. 2005). We used the influence() function for diagnostic tests of influential outliers. We used the cumul() function to observe how summary effect sizes accumulated with a specific variable (e.g. the year of publication).
To control for phylogenetic relatedness (Lajeunesse 2009), we acquired the correlation matrix of phylogenetic relationships. We used the R package rotl (Michonneau et al. 2016) to interface with the ‘Open Tree of Life’ (opentreeoflife.org) database (Redelings and Holder 2017) and retrieve phylogenetic trees (SI1, Fig. S3). With the R package ape (Paradis and Schliep 2018), we used the compute.brlen() function to estimate branch lengths, and the vcv() function to construct the correlation matrix.
We ran a random-effects model using the rma.mv() function to estimate a pooled (summary) effect size. The random-effects model allows the true effect size to vary from case to case, so we estimated heterogeneity (variation in true effect sizes) from observed effects as the statistic T2, which includes both true heterogeneity and random error (Borenstein et al. 2009). We calculated Cochran’s Q to test for heterogeneity (this tests for differences between the effect sizes of cases and the meta-analytical mean effect size) and the I2 statistic to show the proportion of heterogeneity that is due to a variation among studies (Borenstein et al. 2009).
We addressed significant heterogeneity in two ways. First, we evaluated separate meta-analytic models for taxonomic subsets of the data (subgroup meta-analyses) to identify where significant heterogeneities were occurring (Borenstein et al. 2009). Second, we carried out meta-analyses that tested for ‘moderators’, which are variables that are case-level covariates of effect sizes (Viechtbauer 2010). We used the pairs.panels() function of the R package psych (Revelle 2018) to estimate correlations between variables that were potential moderators and effect sizes (Zr) and to estimate pairwise correlations among these variables. We then conducted meta-analyses incorporating and testing for moderators. These models included random factors (see previous text for details of random effects) and controlled for time-lag bias (mean-centred publication year, which is the year of publication minus the mean of all years of publications) and small study effect (inverse of Zr standard error, which is (1/√N − 3)) (see SI1 for R scripts). The program metafor tested whether variables were significant moderators and tested for remaining residual heterogeneity. We initially fitted a model including all candidate moderators. Next, we fitted models with single candidates to work out the best (according to the test of the moderator) to include in models with two moderators. We further considered whether the test for residual heterogeneity remained significant and used corrected Akaike information criterion values (AICC) to confirm the model best fitting the data. We then tested combinations of two candidates for variables that were not significantly correlated with each other. Finally, we tested between models with different numbers of moderators and ran a sensitivity test (see SI1 for details).
Results
In assessing the full-text articles of 370 studies, we found 154 reporting on the occurrence of multiple paternity among clutches or litters from 184 different amphibian and reptile populations (see also SI1 Fig. S1; SI2 lists all studies and references). The majority, 172 of 184 cases, used microsatellite loci. Of the others, one used single nucleotide polymorphisms (SNPs), two used amplified fragment length polymorphism (AFLP), six used allozymes, two used multilocus DNA fingerprinting (minisatellites or variable number of tandem repeats (VNTRs)) and one used both multilocus DNA fingerprinting and a single-locus DNA probe (see SI2 for all references). We estimated PA at 48% after excluding 36 cases with fewer than five samples and two with unknown sample sizes (SD = 30%, K = 146 cases; N.B., for all 184 cases, PA = 49%, SD = 34%). There were more cases reported for the northern hemisphere than for the southern hemisphere (100 versus 46 for cases with N ≥ 5) (Fig. 1A; see also SI1, Fig. S4).
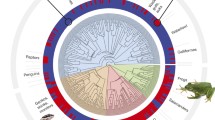
A–H The prevalence of polyandry (PA) in amphibian and reptile breeding populations across taxonomic groups and maps of sampling locations (excludes cases with fewer samples than five). Jittered points in graphs are the PA for individual cases within each taxonomic group, with values summarised above as a box plot (central lines in boxes are median values, numbers are the number of cases, boxes correspond to the first and third quartiles and whiskers extend to 1.5 times of the inter-quartile range). Sampling locations are plotted in maps, with the bubble size reflecting the degree of PA and colour corresponding to the colour used in the associated graph. Single-case taxonomic groups (excluded from summary graphs) are plotted on the map with a black bubble and numbered: (1) Gymnophiona (Boulengerula taitanus), (2) Rhynchocephalia (Sphenodon punctatus), (3) Centrolenidae (Hyalinobatrachium valerioi), (4) Dendrobatidae (Ranitomeya imitator), (5) Leptodactylidae (Thoropa taophora), (6) Megophryidae (Leptobrachium boringii), (7) Phrynosomatidae (Uta stansburiana), (8) Crotaphytidae (Crotaphytus collaris), (9) Chamaeleonidae (Bradypodion pumilum), (10) Diplodactylidae (Oedura reticulata), (11) Pythonidae (Liasis fuscus), (12) Chelidae (Elseya albagula) and (13) Cheloniidae (Natator depressus)
PA varied randomly with respect to absolute latitude (Kendall’s rank correlation tau test: z = 0.84, p = 0.402) and sample size (z = − 1.61, p = 0.107). There were also no significant relationships between PA and the number genotyped per clutch/litter or the proportion genotyped (Kendall’s rank correlation tau tests: z = 1.53, p = 0.126, for the former after excluding two cases with clutch sizes of over a thousand, and z = 0.05, p = 0.959, for the latter). For cases based on microsatellite markers, there were no significant relationships between PA and the number of loci nor the average number of alleles (Kendall’s rank correlation tau tests; z = − 1.65 and 0.45, p = 0.100 and 0.652, respectively). There were no significant relationships with the average observed or expected heterozygosities (Kendall’s rank correlation tau tests: z = 1.58 and 1.88, p = 0.115 and 0.060, respectively) or the year of publication (z = 1.09, p = 0.277).
Patterns across taxonomic groups
PA was widespread and variable across major taxonomic groups (Fig. 1A). There was a large range of PA values within groups, without significant differences among groups (Kruskal–Wallis rank sum test = 8.27, df = 5, p = 0.142). One notable pattern was that Anura had the smallest median PA (18.9%; Fig. 1A) with many cases of relatively low values being of Ranidae (the largest value was no more than 44%). The PA data for Anura were significantly lower than those for Caudata (Wilcoxon rank sum test: W = 41.5, p = 0.031). In contrast, Caudata had the highest median PA (72.4%; Fig. 1A). Thus, cases for Caudata and Anura tended towards opposite ends of the range of values for PA, even though both groups were amphibians. For reptile groups, median PA values were 48.7–56.3% (Fig. 1A).
Within most taxonomic families, PA values ranged widely (Fig. 1B–G), but with some exceptions. Ranidae and Iguanidae were the only families with relatively low PA for all cases (K = 5 cases, range: 0–44%, and K = 2 cases, range: 11–38%, respectively). While values for the five cases of Ranidae only averaged 19%, these were not significantly different from the PA values of the other genera in Anura (Kruskal–Wallis χ2 = 11.44, df = 10, p = 0.324). Iguanidae only had two cases, so it remains to be seen if a greater range of PA values will be reported in future studies. Many taxonomic families were represented by few cases (< 5), so general conclusions would be tentative.
Among the taxonomic groups, Testudines was the most studied (K = 58 cases), with PA values spanning the entire range (0–100%; Fig. 1G). PA for Podocnemididae tended towards high values (median = 95.8%), whereas those of Testudinidae were lower (median = 28.1%), but overall, there were no significant differences among families (Kruskal–Wallis χ2 = 8.22, df = 4, p = 0.084). Among Testudines, Cheloniidae, the taxonomic family with the greatest number of cases (K = 36; 62% of all cases), provided the only opportunity to observe patterns of PA among genera within the family (Fig. 1H). In Cheloniidae, we found a wide range of PA values (0–93%), with one noteworthy pattern. There were significantly lower PA values for Dermochelys (K = 3, range: 10–24%) and Eretmochelys (K = 8 with a range of 0–17%, and one case of 61%) compared to PA values of the other genera (Kruskal–Wallis χ2 = 19.03, df = 5, p = 0.002; with Dermochelys and Eretmochelys excluded: χ2 = 1.35, df = 3, p = 0.716).
Testing the relationship between the number of offspring and the number of sires
Effect size data (36 cases from 35 studies) obtained from all taxonomic groups, except Anura, were checked for influential cases, outliers and publication bias. Egger’s test did not find significant asymmetry in funnel plots (SI1, Fig. S6), so publication bias was not an extensive problem. A forest plot of cumulative meta-analysis conducted according to the year of publication showed initial cumulative effect sizes had high values influenced by a few early cases, but as more were added, the cumulative effect sizes settled close to zero (SI1, Fig. S7). Influential case diagnostics discovered one case for a population of Zootoca vivipara (Eizaguirre et al. 2007), exerting substantial changes in fitted models (see SI1 for Fig. S5 and further details). Funnel plots confirmed this case as an outlier (SI1, Fig. S6). We therefore fitted meta-analysis models with and without this case (Fig. 2). Summary effect sizes estimated by phylogenetically controlled meta-analyses were not significant (Fig. 2), but significant heterogeneity in the data remained even after the influential outlier was excluded (Table 2), and even though the meta-analysis controlled for phylogenetic non-independence and a number of other random factors (see methods and R script in SI1). While heterogeneity may be caused by random error, some may be due to real differences between cases (Borenstein et al. 2009). To clarify the heterogeneity, we used subgroup meta-analyses and tested for moderators.
Forest plot of effect sizes, and summary effect sizes estimated by phylogenetically controlled meta-analysis (random-effects maximum likelihood method) of the relationship between the number of offspring and the number of sires. The star indicates the outlying influential case (see text). Within taxonomic subgroups, cases are organised by the effect size (Zr) and labelled with the reference of the study. Point sizes are proportional to the precision of the estimates, and whiskers, 95% confidence intervals (CI). An effect size is considered significantly different from zero if its 95% confidence interval is either larger than or smaller than zero. Positive values indicate that there are larger numbers of offspring in groups of siblings and half-siblings with more sires, whereas negative values indicate the opposite. ‘PA %’ is the estimated prevalence of polyandry in the population with sample size ‘N’. The black diamond polygon below each subgroup is the estimated summary effect size including the 95% confidence interval, labelled by the taxonomic name of the subgroup. The grey diamond polygon is the summary effect size excluding the outlying influential case. Summary effect sizes for the whole dataset are represented by diamond polygons at the bottom of the graph. The black polygon represents the estimate for all cases (36 cases, labelled ‘Full dataset’), the grey polygon represents the estimate for cases without the outlier (35 cases, labelled ‘Without outlier’) and the white polygon represents the estimate based on a mixed-effects model that included average expected heterozygosity as a moderator (27 cases, labelled ‘With moderator’). The latter excluded the outlying influential case, and cases that were missing data for average expected heterozygosity (see also Fig. 3)
Subgroup meta-analyses
Subgroup meta-analyses revealed significant heterogeneity in just one taxon, Lacertilia (Table 3). Indeed, values of Zr for Lacertilia ranged from − 0.30 to as high as 1.58, the latter being the largest effect size in the dataset (Fig. 2). The meta-analysis included species from diverse families: Scincidae (Tiliqua rugosa and Egernia whitii) (Bull et al. 1998; Chapple and Keogh 2005), Agamidae (Intellagama lesueurii) (Frere et al. 2015), Iguanidae (Cyclura nubila caymanensis) (Moss et al. 2019) and Lacertidae (Z. vivipara) (Eizaguirre et al. 2007). Only two cases had significantly positive effect sizes (Fig. 2): T. rugosa (Bull et al. 1998) and Z. vivipara (Eizaguirre et al. 2007). However, T. rugosa have few offspring (average of two in this case), so the result could be an artefact of multiple paternity being difficult to detect when few offspring are tested. The other case, however, was the outlier identified in influential diagnostics and in funnel plots and likely contributed heterogeneity to the data. The funnel plot for Lacertilia revealed a further case as a possible source of heterogeneity (SI1, Fig. S6): a population of E. whitii studied by Chapple and Keogh (2005). The two outliers had the largest effect sizes in Lacertilia, but in opposite directions (Fig. 2). Maternal size was reported as significantly correlated with the number of mates and the number of offspring in the study on Z. vivipara (Eizaguirre et al. 2007). However, we could not control for maternal size in our meta-analysis because the information was lacking for other cases. Significant heterogeneity remained after leaving out the influential outlier (Table 3). Excluding both outliers reduced heterogeneity to non-significance (I2 = 35.5%, Q = 3.37, p = 0.185), but left only three cases for Lacertilia.
Subgroup meta-analysis showed Caudata as the only group with a significant summary effect size (Fig. 2). Total heterogeneity for Caudata was low (I2 = 2.5%, Q = 3.82, p = 0.431; Table 3), suggesting that the effect sizes were consistent in supporting the summary effect size for the group. All species were salamanders, and of the five, effect sizes for Ambystoma texanum (Gopurenko et al. 2007) and Salamandra salamandra (Caspers et al. 2014) were significant. Individual effect sizes for Ambystoma tigrinum tigrinum (Gopurenko et al. 2006), Salamandrina perspicillata (Rovelli et al. 2015) and Plethodon cinereus (Liebgold et al. 2006) were also positive in value, though not significant (Fig. 2).
For Serpentes, the effect size for one population of Thamnophis sirtalis was significant (McCracken et al. 1999), but those of the other population (Garner et al. 2002) and for other species (Barry et al. 1992; Garner and Larsen 2005; Madsen et al. 2005; Ursenbacher et al. 2009; Clark et al. 2014; Levine et al. 2015) were not (Fig. 2). However, two singly sired litters in the T. sirtalis case only had four offspring each (McCracken et al. 1999), so this may be an artefact of having few offspring to test in paternity analysis.
The summary effect size for three species of Crocodylia (McVay et al. 2008; Lance et al. 2009; Lafferriere et al. 2016; Zajdel et al. 2019) was non-significant, lacked heterogeneity and showed a negative trend (Fig. 2; Table 3).
Subgroup meta-analysis of Testudines included Gopherus polyphemus (Testudinidae) (Moon et al. 2006; Tuberville et al. 2011; White et al. 2018), Podocnemis sextuberculata (Podocnemididae) (Freda et al. 2016) and species of Cheloniidae: Chelonia mydas (Lee and Hays 2004; Wright et al. 2013; Alfaro-Núñez et al. 2015; Turkozan et al. 2019), Eretmochelys imbricata (Phillips et al. 2014), Caretta caretta (Sari et al. 2017), Natator depressus (Theissinger et al. 2009) and Lepidochelys olivacea (Hoekert et al. 2002; Jensen et al. 2006). This resulted in a non-significant summary effect size (Fig. 2) with individual effect sizes not significant except for P. sextuberculata (Fig. 2). Given the lack of significant heterogeneity (Table 3), we did not partition Testudines any further for meta-analysis.
Testing for moderators of the effect size data
In fitting a model with all candidate moderators, we obtained a non-significant result for the omnibus test of moderators and significant residual heterogeneity (see SI1, Table S3). Some candidate moderators were significantly correlated (SI1, Fig. S8). In testing candidate moderators individually (SI1, Table S3B), we found that the model including latitude had the best fit to the data for models based on 35 cases (SI1, white box in Table S3B), but latitude proved not to be a significant moderator of effect size data. Among models with fewer cases (fewer because of missing data), only average expected heterozygosity was found to be a significant moderator (t = 2.213, p = 0.037, K = 27; see SI1, Table S3B). The model with average expected heterozygosity also had the best fit to the data for models that could be compared (see SI1: comparison of AICC values in a light grey box in Table S3B). A cumulative meta-analysis show how predicted summary effect sizes increases with increasing average expected heterozygosity (Fig. 3).
Forest plot of a cumulative meta-analysis for relationships between the number of offspring and the number of sires, cumulated according to the average expected heterozygosity estimated for the population. Points are cumulative effect sizes (Zr) with 95% confidence intervals (CI) and labelled with the reference of the study (see Fig. 2 for species names). The data for each study is cumulated progressing from top to bottom (each addition symbolized with ‘ + ’). The plot excludes cases that lack data for average expected heterozygosity. Diamond polygons at the bottom show summary effect sizes including the 95% confidence interval that are predicted for various values of average expected heterozygosity (minimum, mean and maximum of the dataset), for a mixed-effects model fitted with average expected heterozygosity specified as the moderator. This model also controlled for time-lag bias and small study effect and included random factors. The predicted effect size based on the mean value of average expected heterozygosity is also displayed as a white polygon in Fig. 2. Stars mark cases that, when left out of the dataset, result in non-significant results in tests of average expected heterozygosity as a moderator
We next fitted models including average expected heterozygosity and other candidate moderators (SI1, Table S3C), excluding those that were significantly correlated with average expected heterozygosity (i.e. observed heterozygosity, the number of loci sampled and the average number of alleles per locus; see SI1, Fig. S8). However, fitting models with two candidate moderators did not result in discernibly better models: omnibus results were non-significant, and residual heterogeneity remained significant (SI1, Table S3C). Differences between models with one, two or three moderators were non-significant (SI1, Table S4). We ceased further model fitting.
Curiously, the average expected heterozygosity was not a significant moderator for the reduced dataset of 26 cases (SI1, Table S3B). A sensitivity test (models were fitted repeatedly, leaving out a different case each time) revealed three cases that, when each was excluded, resulted in non-significant tests (the cases are marked by stars in Fig. 3). Furthermore, differences between models with or without average expected heterozygosity as a moderator were not significant (SI1, Table S4). These tests highlighted the tenuous nature of average expected heterozygosity as moderating the effect size data.
Discussion
Our tests showed that Caudata females had significantly more offspring with increasing numbers of sires (Fig. 2), whereas there was no evidence for this relationship among reptiles. Summary effect sizes for Testudines and Serpentes were non-significant (Fig. 2), with little heterogeneity among cases (Table 3). This was also true for a smaller number of Crocodylia cases, but effect sizes tended towards negative values (Fig. 2), which implied that multiple paternity may incur costs for Crocodylia (e.g. Zajdel et al. 2019). Results for Lacertilia were less clear because significant heterogeneity could not be resolved, but individual effect sizes for species and populations were mostly non-significant (Fig. 2). Lacertilia is the largest group of living reptiles, and families are diverse with different morphologies, habitats, reproductive modes (e.g. egg-laying or live bearing), mating systems (monogamy, polygyny, polyandry and polygynandry; see Table 1) and behaviours (e.g. social bonds, community living, territoriality, male dominance hierarchies and male combat). Many are capable of sperm storage, and male competition for females can be intense (Wapstra and Olsson 2014). There are over twenty-five families of Lacertilia, yet studies of polyandry have only been reported for nine (Fig. 1E). Taxonomically limited and biased studies meant patchy coverage of Lacertilia in our meta-analysis, which probably exacerbated the problem with heterogeneity. Overall, we concluded that most female reptiles that produced offspring with different sires did not have more offspring than monogamous females. For cases where polyandrous females did have significantly more offspring, authors offered alternative explanations, proposed other benefits to females or discovered confounding factors such as maternal condition, age or attractiveness (e.g. Table 2).
Among amphibians, Anura are mostly external fertilisers, which allows simultaneous polyandry to result from multiple amplexus (Table 1) (e.g. Sztatecsny et al. 2006) or from the presence of free swimming sperm from multiple males (e.g. Hase and Shimada 2014). However, we found relatively low PA, with some populations exhibiting little or no evidence of polyandry (Fig. 1A). There was a lack of data on whether or not female Anura produced more offspring from polyandrous matings. Counting the number of offspring to estimate female reproductive output for some species is not straightforward since females may produce hundreds to thousands of eggs in a single deposition. Anura capable of sequential polyandry may insure against nest failure by partitioning eggs among multiple males (Byrne and Keogh 2009), but aside from this, there has been little other evidence that female Anura could benefit from polyandry (Roberts and Byrne 2011; Table 2).
Thus, among amphibians, we only have evidence of female salamanders having increased reproductive success with polyandry (Fig. 2). Many Caudata use spermatophore transfer (Table 1) to internally fertilise eggs, which accords females greater control over sperm acquisition than other amphibians and reptiles, since they can choose to reject or pick up spermatophores. Polyandry for some species is further facilitated by sperm storage (e.g. Adams et al. 2005) or explosive breeding (Table 1) (e.g. Myers and Zamudio 2004). Interestingly, female Caudata also had relatively high levels of polyandry (highest median PA among amphibians and reptiles; Fig. 1). Even socially monogamous species of Caudata were found to be predominantly genetically polyandrous (Liebgold et al. 2006). Female Caudata having virtually complete pre-copulatory choice have little need for post-copulatory choice (Jones et al. 2002; Rafinski and Osikowski 2002; Gopurenko et al. 2006). There is evidence that female choice may be contingent on relatedness—with preferred mates being genetically similar (Caspers et al. 2014) or dissimilar from themselves (Rovelli et al. 2015). For some species, females may require multiple insemination events because eggs are produced over extended periods (Osikowski and Rafiński 2001). Thus, having the pre-copulatory choice to be polyandrous seems a key to enhancing female reproductive success for Caudata. Experimental studies suggest the females of some species ensured fertilisation by mating with the first encountered male, but thereafter choose higher-quality (Gabor and Halliday 1997) or more genetically compatible mates (Garner and Schmidt 2003). Furthermore, mate choice may be condition- (Bos et al. 2009) or context-dependent (Chandler and Zamudio 2008), which may explain contrasting patterns found for some populations (e.g. Caspers et al. 2014 versus Rovelli et al. 2015).
Our study tested one indicator of a benefit for females (reproductive output), but there may be other benefits for female reptiles and amphibians (Table 2). Polyandry facilitated by sperm storage extends the potential for female choice when mating opportunities or choice of mates is limited or unpredictable. Stored sperm can be used in the same season (e.g. Fitzsimmons 1998; Roques et al. 2004, 2006), between years (e.g. Pearse et al. 2002; Roques et al. 2006; Anthonysamy et al. 2014; McGuire et al. 2014; Phillips et al. 2014; Riley et al. 2021) or over decades in some species (Whitaker 2006; Murphy et al. 2007). Female Caudata may have little need for post-copulatory choice, but for other species of amphibians and reptiles, cryptic choice enables the female to diversify offspring characteristics and/or to select optimal genotypes/phenotypes for offspring (e.g. Calsbeek and Bonneaud 2008). For social reptiles, inbreeding avoidance could be achieved through polyandry (While et al. 2014), which may be important for socially monogamous species with limited choice of mates beyond close kin.
One certain outcome of polyandry is that the diversity of paternal alleles will be greater for groups of offspring with multiple sires compared with groups of offspring with only one sire (e.g. Calsbeek et al. 2007; Bouchard et al. 2018; Moss et al. 2019): some argue that the augmented genetic diversity of offspring per se benefits females. For example, greater diversity could boost the chances of offspring surviving variable or unpredictable environments and reduce sibling competition (Calsbeek et al. 2007; Eales et al. 2010; Bouchard et al. 2018). Offspring viability may be improved with increased offspring heterozygosity or genetic diversity (examples in Table 2), potentially because of better male–female genetic compatibility (Madsen et al. 2005). These are intriguing examples, but definitive genetic benefits have been difficult to detect (e.g. Noble et al. 2013; also see reviews listed in SI1, Table S1). Perhaps a more general route for genetic benefits is through “heterozygosity-fitness-correlations” (HFCs), in which genome-wide multilocus heterozygosity is interpreted as a proxy for the degree of inbreeding and, thereby, the potential for inbreeding depression (Chapman et al. 2009). From a female’s perspective under an HFC scenario, multiple sires for her offspring increase mean heterozygosity and reduce risks of loss of viability, survival or fecundity to inbreeding depression. We did detect average expected heterozygosity of the population as a moderator of the effect size data (Fig. 3). The calculation of expected heterozygosity is based on the relative frequencies of alleles at loci surveyed for unrelated adults in a population. Therefore, an increase in expected heterozygosity may simply be a by-product of relatively more individuals (males) mating in the system (Rafajlović et al. 2013), even though the HFC model would argue that this would have fitness benefits. However, because our evidence proved tenuous (leaving out one of three cases led to non-significant results), it remains for future work to re-examine links between heterozygosity, polyandry and reproductive success when more cases become available.
Problems, solutions and future research directions
Paternity analysis will continue to be an invaluable tool underpinning studies on the behaviour, ecology, evolution and conservation of amphibians and reptiles (Lee 2008; Flanagan and Jones 2019). For example, paternity data helps identify the genetic mating system (Gardner et al. 2002; Chapple and Keogh 2005; Clark et al. 2014), and comparisons between different seasons or populations reveal variation in mate encounter rate and mating opportunity for the different sexes (Lodé et al. 2005; Olsson et al. 2011; Lasala et al. 2013, 2018; Lee et al. 2018), while temporal changes potentially track population decline or recovery (Figgener et al. 2016). Paternal alleles determined from offspring provide genetic data about breeding males, which, for some species, cannot be easily obtained in any other way (Madsen et al. 2005; Lee et al. 2007). The current study has additionally demonstrated the value of obtaining data on polyandry across a broad diversity of populations and species. However, in terms of understanding global and trans-taxa patterns of polyandry, there remains outstanding problems with current methods of study and particularly, with sampling.
Sampling across all levels, from within-study, to across geographical locations and within and among taxa, was a critical issue. Sampling was extremely limited in some studies: 18% of cases in our dataset had sample sizes fewer than five. Furthermore, estimates of polyandry for many species were based on a single case. There are still gaps in our knowledge of female reproductive strategies for many amphibian and reptile species, and limited data for many taxonomic groups. For example, Caudata could offer original, powerful, under-tested models for advancing understanding of polyandry, and some species are already used in experiments on sexual selection, sperm competition and mate choice (e.g. Jones et al. 2004; Chandler and Zamudio 2008; Bos et al. 2009). However, we had few cases for Caudata (ten in Fig. 1 and five in Fig. 2). Some of the other taxonomic groups were even more poorly represented—only by a single case (numbered black points in Fig. 1). The geographical distribution of cases was biased towards temperate and the northern hemisphere, and studies of species from larger groups, such as Lacertilia, Serpentes and Anura, concentrated on certain families (Scincidae, Natricidae and Ranidae, respectively), perhaps reflecting interests of researchers rather than species distribution. Improved coverage within and among species will lead to estimates that are more accurate, and accumulation of data for many more populations and species will facilitate hypothesis testing (e.g. Zbinden et al. 2007; Lee et al. 2018).
Heterogeneity in the effect size data across studies was a significant problem. We attempted to include available data from any species of amphibian or reptile in one meta-analysis, but could not resolve the heterogeneity, even by controlling for random factors and candidate moderators. However, most subgroup meta-analyses did not suffer from significant heterogeneity. We recommend that future meta-analyses may be more appropriately conducted at lower taxonomic levels, such as at the family or subfamily level.
Another problem was the lack of information in some studies. For example, out of 154 studies, we only had 36 cases suitable for meta-analysis. Clearly, future meta-analyses would be facilitated if studies reported further data. Aside from measures of reproductive output, we suggest reporting on maternal and offspring characteristics. Most studies reporting the number of offspring provided no further information (e.g. viability, fledging/survival success, sizes, development, relative performance, fecundity). A few reported maternal condition, but we could not test this variable because we lacked the data for most cases. Limited data restricts the scope for meta-analyses and constrains the opportunities for comparing species with contrasting life histories.
We examined the link between polyandry and female reproductive success, but questions that were beyond this study were whether polyandry or female benefits varied according to the method of reproduction (i.e. egg-laying versus live-birth), sexual dimorphism, lifetime reproductive potential or environmental conditions. How these factors may affect polyandry for amphibians and reptiles remains avenues for future research. To progress our understanding of polyandry, other reviews of mating systems have advocated integration of observational and experimental approaches, and a more holistic view of the entire reproductive process to include consideration of multiple drivers, both male- and female-based, that may vary within and between species (Uller and Olsson 2008; Wapstra and Olsson 2014; Lyons et al. 2021). We would extend on these suggestions to propose meta-analysis as a powerful tool in consolidating information on polyandry across studies, populations and species. We further urge researchers to take a broader taxonomic view and to continue studying polyandry within and among taxa. Recording and reporting more widely to expand the baseline data will undoubtedly improve future attempts at meta-analysis and support experimental and comparative studies. Establishing a taxonomically well-sampled framework will enhance the utility of amphibians and reptiles as model systems and stimulate growth in the evolutionary and ecological study of male and female reproductive strategies.
Supplementary information
SI1 Additional detail further to those presented in the main text and R scripts.
SI2 Data and data references. Includes: all cases of multiple paternity and associated references; references for excluded studies, and reasons for exclusion; data used in meta-analyses.
Data availability
Data and R scripts used in analyses are available as Supplementary Information.
References
Adams EM, Jones AG, Arnold SJ (2005) Multiple paternity in a natural population of a salamander with long-term sperm storage. Mol Ecol 14:1803–1810. https://doi.org/10.1111/j.1365-294X.2005.02539.x
Alfaro-Núñez A, Jensen MP, Abreu-Grobois FA (2015) Does polyandry really pay off? The effects of multiple mating and number of fathers on morphological traits and survival in clutches of nesting green turtles at Tortuguero. PeerJ 3:e880. https://doi.org/10.7717/peerj.880
Anthonysamy WJB, Dreslik MJ, Douglas MR, Marioni NK, Phillips CA (2014) Reproductive ecology of an endangered turtle in a fragmented landscape. Copeia 2014:437–446. https://doi.org/10.1643/cg-13-137
Avise JC, Liu J-X (2011) Multiple mating and its relationship to brood size in pregnant fishes versus pregnant mammals and other viviparous vertebrates. P Natl Acad Sci USA 108:7091–7095. https://doi.org/10.1073/pnas.1103329108
Barry FE, Weatherhead PJ, Philipp DP (1992) Multiple paternity in a wild population of northern water snakes, Nerodia sipedon. Behav Ecol Sociobiol 30:193–199. https://doi.org/10.1007/BF00166703
Bateman A (1948) Intra-sexual selection in Drosophila. Heredity 2:349–368
Birkhead TR (2000) Defining and demonstrating postcopulatory female choice – again. Evolution 54:1057–1060
Blouin-Demers G, Gibbs HL, Weatherhead PJ (2005) Genetic evidence for sexual selection in black ratsnakes, Elaphe obsoleta. Anim Behav 69:225–234. https://doi.org/10.1016/j.anbehav.2004.03.012
Borenstein M, Hedges L, Higgens J, Rothstein H (2009) Introduction to meta-analysis. John Wiley & Sons Ltd, West Sussex, UK
Bos DH, Williams RN, Gopurenko D, Bulut Z, Dewoody JA (2009) Condition-dependent mate choice and a reproductive disadvantage for MHC-divergent male tiger salamanders. Mol Ecol 18:3307–3315. https://doi.org/10.1111/j.1365-294X.2009.04242.x
Bouchard C, Tessier N, Lapointe F-J (2018) Paternity analysis of wood turtles (Glyptemys insculpta) reveals complex mating patterns. J Hered 109:405–415. https://doi.org/10.1093/jhered/esx103
Boulton RA, Zuk M, Shuker DM (2018) An inconvenient truth: the unconsidered benefits of convenience polyandry. Trends Ecol Evol 33:904–915. https://doi.org/10.1016/j.tree.2018.10.002
Bretman A, Tregenza T (2005) Measuring polyandry in wild populations: a case study using promiscuous crickets. Mol Ecol 14:2169–2179
Brommer JE, Gustafsson L, Pietiäinen H, Merilä J (2004) Single-generation estimates of individual fitness as proxies for long-term genetic contribution. Am Nat 163:505–517
Bull CM, Cooper SJB, Baghurst BC (1998) Social monogamy and extra-pair fertilization in an Australian lizard, Tiliqua rugosa. Behav Ecol Sociobiol 44:63–72. https://doi.org/10.1007/s002650050515
Byrne PG, Keogh JS (2009) Extreme sequential polyandry insures against nest failure in a frog. Proc R Soc Lond B 276:115–120. https://doi.org/10.1098/rspb.2008.0794
Calsbeek R, Bonneaud C (2008) Postcopulatory fertilization bias as a form of cryptic sexual selection. Evolution 62:1137–1148. https://doi.org/10.1111/j.1558-5646.2008.00356.x
Calsbeek R, Bonneaud C, Prabhu S, Manoukis N, Smith TB (2007) Multiple paternity and sperm storage lead to increased genetic diversity in Anolis lizards. Evol Ecol Res 9:495–503
Caspers BA, Krause ET, Hendrix R, Kopp M, Rupp O, Rosentreter K, Steinfartz S (2014) The more the better – polyandry and genetic similarity are positively linked to reproductive success in a natural population of terrestrial salamanders (Salamandra salamandra). Mol Ecol 23:239–250. https://doi.org/10.1111/mec.12577
Chandler CH, Zamudio KR (2008) Reproductive success by large, closely related males facilitated by sperm storage in an aggregate breeding amphibian. Mol Ecol 17:1564–1576. https://doi.org/10.1111/j.1365-294X.2007.03614.x
Chapman J, Nakagawa S, Coltman D, Slate J, Sheldon B (2009) A quantitative review of heterozygosity-fitness correlations in animal populations. Mol Ecol 18:2746–2765. https://doi.org/10.1111/j.1365-294X.2009.04247.x
Chapple DG, Keogh JS (2005) Complex mating system and dispersal patterns in a social lizard, Egernia whitii. Mol Ecol 14:1215–1227. https://doi.org/10.1111/j.1365-294X.2005.02486.x
Clark R, Schuett G, Repp R, Amarello M, Smith C, Herrmann H-W (2014) Mating systems, reproductive success, and sexual selection in secretive species: a case study of the western diamond-backed rattlesnake, Crotalus atrox. Plos One 9:e90616. https://doi.org/10.1371/journal.pone.0090616
Correia HE, Abebe A, Dobson FS (2021) Multiple paternity and the number of offspring: a model reveals two major groups of species. BioEssays 43:2000247. https://doi.org/10.1002/bies.202000247
Eales J, Thorpe RS, Malhotra A (2010) Colonization history and genetic diversity: adaptive potential in early stage invasions. Mol Ecol 19:2858–2869. https://doi.org/10.1111/j.1365-294X.2010.04710.x
Eizaguirre C, Laloi D, Massot M, Richard M, Federici P, Clobert J (2007) Condition dependence of reproductive strategy and the benefits of polyandry in a viviparous lizard. Proc R Soc Lond B 274:425–430. https://doi.org/10.1098/rspb.2006.3740
Figgener C, Chacón-Chaverri D, Jensen MP, Feldhaar H (2016) Paternity re-visited in a recovering population of Caribbean leatherback turtles (Dermochelys coriacea). J Exp Mar Biol Ecol 475:114–123. https://doi.org/10.1016/j.jembe.2015.11.014
Firman RC, Gasparini C, Manier MK, Pizzari T (2017) Postmating female control: 20 years of cryptic female choice. Trends Ecol Evol 32:368–382. https://doi.org/10.1016/j.tree.2017.02.010
Fitzsimmons NN (1998) Single paternity of clutches and sperm storage in the promiscuous green turtle (Chelonia mydas). Mol Ecol 7:575–584. https://doi.org/10.1046/j.1365-294x.1998.00355.x
Flanagan SP, Jones AG (2019) The future of parentage analysis: from microsatellites to SNPs and beyond. Mol Ecol 28:544–567. https://doi.org/10.1111/mec.14988
Forstmeier W, Nakagawa S, Griffith SC, Kempenaers B (2014) Female extra-pair mating: adaptation or genetic constraint? Trends Ecol Evol 29:456–464. https://doi.org/10.1016/j.tree.2014.05.005
Freda FP, Bernardes VCD, Eisemberg CC, Fantin C, Vogt RC (2016) Relationship between multiple paternity and reproductive parameters for Podocnemis sextuberculata (Testudines: Podocnemididae) in the Trombetas River Brazil. Genet Mol Res 15:15017335. https://doi.org/10.4238/gmr.15017335
Frere CH, Chandrasoma D, Whiting MJ (2015) Polyandry in dragon lizards: inbred paternal genotypes sire fewer offspring. Ecol Evol 5:1686–1692. https://doi.org/10.1002/ece3.1447
Friesen CR, Kahrl AF, Olsson M (2020) Sperm competition in squamate reptiles. Phil Trans R Soc B 375:20200079. https://doi.org/10.1098/rstb.2020.0079
Gabor CR, Halliday TR (1997) Sequential mate choice by multiply mating smooth newts: females become more choosy. Behav Ecol 8:162–166. https://doi.org/10.1093/beheco/8.2.162
Garcia-Gonzalez F, Yasui Y, Evans JP (2015) Mating portfolios: bet-hedging, sexual selection and female multiple mating. Proc R Soc B 282:20141525. https://doi.org/10.1098/rspb.2014.1525
Gardner MG, Bull CM, Cooper SJB (2002) High levels of genetic monogamy in the group-living Australian lizard Egernia stokesii. Mol Ecol 11:1787–1794. https://doi.org/10.1046/j.1365-294X.2002.01552.x
Garner TWJ, Gregory PT, McCracken GF, Burghardt GM, Koop BF, McLain SE, Nelson RJ (2002) Geographic variation of multiple paternity in the common garter snake (Thamnophis sirtalis). Copeia 2002:15–23. https://doi.org/10.1643/0045-8511(2002)002[0015:gvompi]2.0.co;2
Garner TWJ, Larsen KW (2005) Multiple paternity in the western terrestrial garter snake, Thamnophis elegans. Can J Zool 83:656–663. https://doi.org/10.1139/z05-057
Garner TWJ, Schmidt BR (2003) Relatedness, body size and paternity in the alpine newt, Triturus alpestris. Proc R Soc Lond B 270:619–624. https://doi.org/10.1098/rspb.2002.2284
Gopurenko D, Williams RN, DeWoody JA (2007) Reproductive and mating success in the small-mouthed salamander (Ambystoma texanum) estimated via microsatellite parentage analysis. Evol Biol 34:130–139. https://doi.org/10.1007/s11692-007-9009-0
Gopurenko D, Williams RN, McCormick CR, DeWoody JA (2006) Insights into the mating habits of the tiger salamander (Ambystoma tigrinum tigrinum) as revealed by genetic parentage analyses. Mol Ecol 15:1917–1928. https://doi.org/10.1111/j.1365-294X.2006.02904.x
Gowaty PA (2013) Adaptively flexible polyandry. Anim Behav 86:877–884. https://doi.org/10.1016/j.anbehav.2013.08.015
Griffith SC, Owens IPF, Thuman KA (2002) Extra pair paternity in birds: a review of interspecific variation and adaptive function. Mol Ecol 11:2195–2212. https://doi.org/10.1046/j.1365-294X.2002.01613.x
Hase K, Shimada M (2014) Female polyandry and size-assortative mating in isolated local populations of the Japanese common toad Bufo japonicus. Biol J Linn Soc 113:236–242. https://doi.org/10.1111/bij.12339
Hettyey A, Hegyi G, Puurtinen M, Hoi H, Torok J, Penn DJ (2010) Mate choice for genetic benefits: time to put the pieces together. Ethology 116:1–9
Hoekert WEJ, Neufeglise H, Schouten AD, Menken SBJ (2002) Multiple paternity and female-biased mutation at a microsatellite locus in the olive ridley sea turtle (Lepidochelys olivacea). Heredity 89:107–113. https://doi.org/10.1038/sj.hdy.6800103
Jennions MD, Petrie M (2000) Why do females mate multiply? A review of the genetic benefits. Biol Rev 75:21–64. https://doi.org/10.1111/j.1469-185X.1999.tb00040.x
Jensen MP, Abreu-Grobois FA, Frydenberg J, Loeschcke V (2006) Microsatellites provide insight into contrasting mating patterns in arribada vs. non-arribada olive ridley sea turtle rookeries. Mol Ecol 15:2567–2575. https://doi.org/10.1111/j.1365-294X.2006.02951.x
Jones AG, Adams EM, Arnold SJ (2002) Topping off: a mechanism of first-male sperm precedence in a vertebrate. P Natl Acad Sci USA 99:2078–2081. https://doi.org/10.1073/pnas.042510199
Jones AG, Arguello JR, Arnold SJ (2004) Molecular parentage analysis in experimental newt populations: the response of mating system measures to variation in the operational sex ratio. Am Nat 164:444–456. https://doi.org/10.1086/423826
Kokko H, Mappes J (2013) Multiple mating by females is a natural outcome of a null model of mate encounters. Entomol Exp Appl 146:26–37
Lafferriere NAR, Antelo R, Alda F, Martensson D, Hailer F, Castroviejo-Fisher S, Ayarzaguena J, Ginsberg JR, Castroviejo J, Doadrio I, Vila C, Amato G (2016) Multiple paternity in a reintroduced population of the Orinoco crocodile (Crocodylus intermedius) at the El Frio Biological Station, Venezuela. Plos One 11:e0150245. https://doi.org/10.1371/journal.pone.0150245
Lajeunesse MJ (2009) Meta-analysis and the comparative phylogenetic method. Am Nat 174:369–381
Lance SL, Tuberville TD, Dueck L, Holz-Schietinger C, Trosclair PL III, Elsey RM, Glenn TC (2009) Multiyear multiple paternity and mate fidelity in the American alligator, Alligator mississippiensis. Mol Ecol 18:4508–4520. https://doi.org/10.1111/j.1365-294X.2009.04373.x
Lasala JA, Harrison JS, Williams KL, Rostal DC (2013) Strong male-biased operational sex ratio in a breeding population of loggerhead turtles (Caretta caretta) inferred by paternal genotype reconstruction analysis. Ecol Evol 3:4736–4747. https://doi.org/10.1002/ece3.761
Lasala JA, Hughes CR, Wyneken J (2018) Breeding sex ratio and population size of loggerhead turtles from Southwestern Florida. PLoS ONE 13:e0191615. https://doi.org/10.1371/journal.pone.0191615
Lee PLM (2008) Molecular ecology of marine turtles: new approaches and future directions. J Exp Mar Biol Ecol 356:25–42. https://doi.org/10.1016/j.jembe.2007.12.02
Lee PLM, Hays GC (2004) Polyandry in a marine turtle: females make the best of a bad job. P Natl Acad Sci USA 101:6530–6535. https://doi.org/10.1073/pnas.0307982101
Lee PLM, Luschi P, Hays GC (2007) Detecting female precise natal philopatry in green turtles using assignment methods. Mol Ecol 16:61–74. https://doi.org/10.1111/j.1365-294X.2006.03115.x
Lee PLM, Schofield G, Haughey RI, Mazaris AD, Hays GC (2018) A review of patterns of multiple paternity across sea turtle rookeries. Adv Mar Biol 79:1–31. https://doi.org/10.1016/bs.amb.2017.09.004
Levine BA, Smith CF, Schuett GW, Douglas MR, Davis MA, Douglas ME (2015) Bateman-Trivers in the 21st century: sexual selection in a North American pitviper. Biol J Linn Soc 114:436–445. https://doi.org/10.1111/bij.12434
Liebgold EB, Cabe PR, Jaeger RG, Leberg PL (2006) Multiple paternity in a salamander with socially monogamous behaviour. Mol Ecol 15:4153–4160. https://doi.org/10.1111/j.1365-294X.2006.03076.x
Lodé T, Holveck MJ, Lesbarrères D (2005) Asynchronous arrival pattern, operational sex ratio and occurrence of multiple paternities in a territorial breeding anuran, Rana dalmatina. Biol J Linn Soc 86:191–200. https://doi.org/10.1111/j.1095-8312.2005.00521.x
Lyons K, Kacev D, Mull CG (2021) An inconvenient tooth: evaluating female choice in multiple paternity using an evolutionarily and ecologically important vertebrate clade. Mol Ecol 30:1574–1593. https://doi.org/10.1111/mec.15844
Madsen T, Ujvari B, Olsson M, Shine R (2005) Paternal alleles enhance female reproductive success in tropical pythons. Mol Ecol 14:1783–1787. https://doi.org/10.1111/j.1365-294X.2005.02505.x
McCracken GF, Burghardt GM, Houts SE (1999) Microsatellite markers and multiple paternity in the garter snake Thamnophis sirtalis. Mol Ecol 8:1475–1479. https://doi.org/10.1046/j.1365-294x.1999.00720.x
McGuire JM, Congdon JD, Scribner KT, Nagle RD (2014) Female reproductive qualities affect male painted turtle (Chrysemys picta marginata) reproductive success. Behav Ecol Sociobiol 68:1569–1602. https://doi.org/10.1007/s00265-014-1768-x
McGuire JM, Scribner KT, Congdon JD (2013) Spatial aspects of movements, mating patterns, and nest distributions influence gene flow among population subunits of Blanding’s turtles (Emydoidea blandingii). Conserv Genet 14:1029–1042. https://doi.org/10.1007/s10592-013-0493-8
McVay JD, Rodriguez D, Rainwater TR, Dever JA, Platt SG, McMurry ST, Forstner MRJ, Densmore LD III (2008) Evidence of multiple paternity in Morelet’s crocodile (Crocodylus moreletii) in Belize, CA, inferred from microsatellite markers. J Exp Zool A 309A:643–648. https://doi.org/10.1002/jez.500
Michonneau F, Brown JW, Winter DJ (2016) rotl: an R package to interact with the Open Tree of Life data. Methods Ecol Evol 7:1476–1481. https://doi.org/10.1111/2041-210X.12593
Moon JC, McCoy ED, Mushinsky HR, Karl SA (2006) Multiple paternity and breeding system in the gopher tortoise, Gopherus polyphemus. J Hered 97:150–157. https://doi.org/10.1093/jhered/esj017
Moss JB, Gerber GP, Schwirian A, Jackson AC, Welch ME (2019) Evidence for dominant males but not choosy females in an insular rock iguana. Behav Ecol 30:181–193. https://doi.org/10.1093/beheco/ary131
Murphy RW, Berry KH, Edwards T, McLuckie AM (2007) A gentic assessment of the recovery units for the Mojave population of the desert tortoise, Gopherus agassizii. Chelonian Conserv Biol 6:229–251
Myers EM, Zamudio KR (2004) Multiple paternity in an aggregate breeding amphibian: the effect of reproductive skew on estimates of male reproductive success. Mol Ecol 13:1951–1963. https://doi.org/10.1111/j.1365-294X.2004.02208.x
Nguyen T, Moehring A (2015) Accurate alternative measurements of female lifetime reproductive success in Drosophila melanogaster. PLoS ONE 10:e0116679. https://doi.org/10.1371/journal.pone.0116679
Noble DWA, Keogh JS, Whiting MJ (2013) Multiple mating in a lizard increases fecundity but provides no evidence for genetic benefits. Behav Ecol 24:1128–1137. https://doi.org/10.1093/beheco/art040
Olsson M, Madsen T (2001) Promiscuity in sand lizards (Lacerta agilis) and adder snakes (Vipera berus): causes and consequences. J Hered 92:190–197. https://doi.org/10.1093/jhered/92.2.190
Olsson M, Schwartz T, Wapstra E, Uller T, Ujvari B, Madsen T, Shine R (2011) Climate change, multiple paternity and offspring survival in lizards. Evolution 65:3323–3326. https://doi.org/10.1111/j.1558-5646.2011.01387.x
Osikowski A, Rafiński J (2001) Multiple insemination increases reproductive success of female Montandon’s newt (Triturus montandoni, Caudata, Salamandridae). Behav Ecol Sociobiol 49:145–149. https://doi.org/10.1007/s002650000277
Paradis E, Schliep K (2018) ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35:526–528. https://doi.org/10.1093/bioinformatics/bty633
Parker PG, Birkhead TR (2013) Polyandry: the history of a revolution. Phil Trans R Soc B 368:20120335. https://doi.org/10.1098/rstb.2012.0335
Pearse DE, Janzen FJ, Avise JC (2002) Multiple paternity, sperm storage, and reproductive success of female and male painted turtles (Chrysemys picta) in nature. Behav Ecol Sociobiol 51:164–171. https://doi.org/10.1007/s00265-001-0421-7
Phillips KP, Mortimer JA, Jolliffe KG, Jolliffe S-M, Hodgkiss RD, McClelland JHR, Liljevik A (2017) Season-long sperm storage and no multiple paternity in green turtles (Chelonia mydas) nesting on Cousine Island, Seychelles. Mar Turtle Newsl 154:6–11
Phillips KP, Mortimer JA, Jolliffe KG, Jorgensen TH, Richardson DS (2014) Molecular techniques reveal cryptic life history and demographic processes of a critically endangered marine turtle. J Exp Mar Biol Ecol 455:29–37. https://doi.org/10.1016/j.jembe.2014.02.012
Pick JL, Nakagawa SWA, Noble DWA (2019) Reproducible, flexible and high-throughput data extraction from primary literature: the metaDigitise R package. Methods Ecol Evol 10:426–431. https://doi.org/10.1111/2041-210X.13118
Prosser MR, Weatherhead PJ, Gibbs HL, Brown GP (2002) Genetic analysis of the mating system and opportunity for sexual selection in northern water snakes (Nerodia sipedon). Behav Ecol 13:800–807. https://doi.org/10.1093/beheco/13.6.800
R Core Team (2019) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.r-project.org. Accessed 13 Nov 2019
Rafajlović M, Eriksson A, Rimark A, Hintz-Saltin S, Charrier G, Panova M, André C, Johannesson K, Mehlig B (2013) The effect of multiple paternity on genetic diversity of small populations during and after colonisation. PLoS ONE 8:e75587. https://doi.org/10.1371/journal.pone.0075587
Rafinski J, Osikowski A (2002) Sperm mixing in the alpine newt (Triturus alpestris). Can J Zool 80:1293–1298. https://doi.org/10.1139/z02-099
Redelings BD, Holder MT (2017) A supertree pipeline for summarizing phylogenetic and taxonomic information for millions of species. PeerJ 5:e3058. https://doi.org/10.7717/peerj.3058
Revelle W (2018) psych: procedures for psychological, psychometric, and personality research. R package version 1.8.12. https://CRAN.R-project.org/package=psych. Accessed 13 Dec 2019
Riley JL, Stow AJ, Bolton PE, Dennison S, Byrne RW, Whiting MJ (2021) Sperm storage in a family-living lizard, the tree skink (Egernia striolata). J Hered 112:526–534. https://doi.org/10.1093/jhered/esab048
Roberts JD, Byrne PG (2011) Polyandry, sperm competition, and the evolution of anuran amphibians. Adv Stud Behav 43:1–53
Roques S, Diaz-Paniagua C, Andreu AC (2004) Microsatellite markers reveal multiple paternity and sperm storage in the Mediterranean spurthighed tortoise, Testudo graeca. Can J Zool 82:153–159. https://doi.org/10.1139/z03-228
Roques S, Diaz-Paniagua C, Portheault A, Perez-Santigosa N, Hidalgo-Vila J (2006) Sperm storage and low incidence of multiple paternity in the European pond turtle, Emys orbicularis: a secure but costly strategy? Biol Conserv 129:236–243. https://doi.org/10.1016/j.biocon.2005.10.039
Rosenberg MS, Adams DC, Gurevitch J (2000) MetaWin: statistical software for meta-analysis. Version 2.0. Sinauer Associates, Sunderland, MA. https://www.rosenberglab.net/Rosenberg2000MetaWin2.html. Accessed 17 Mar 2012
Rothstein H, Sutton A, Borenstein M (eds) (2005) Publication bias in meta-analysis: prevention, assessment and adjustments. Wiley, Chichester, England
Rovelli V, Randi E, Davoli F, Macale D, Bologna MA, Vignoli L (2015) She gets many and she chooses the best: polygynandry in Salamandrina perspicillata (Amphibia: Salamandridae). Biol J Linn Soc 116:671–683. https://doi.org/10.1111/bij.12613
RStudio Team (2015) RStudio: intergrated development for R. RStudio, Inc., Boston, MA. https://www.rstudio.com. Accessed 13 Nov 2019
Sandrin L, Meunier J, Raveh S, Walser J-C, Koelliker M (2015) Multiple paternity and mating group size in the European earwig, Forficula auricularia. Ecol Entomol 40:159–166. https://doi.org/10.1111/een.12171
Sari F, Koseler A, Kaska Y (2017) First observation of multiple paternity in loggerhead sea turtles, Caretta caretta, nesting on Dalyan Beach, Turkey. J Exp Mar Biol Ecol 488:60–71. https://doi.org/10.1016/j.jembe.2016.11.018
Slatyer RA, Mautz BS, Backwell PR, Jennions MD (2012) Estimating genetic benefits of polyandry from experimental studies: a meta-analysis. Biol Rev 87:1–33
Sztatecsny M, Jehle R, Burke T, Hoedl W (2006) Female polyandry under male harassment: the case of the common toad (Bufo bufo). J Zool 270:517–522. https://doi.org/10.1111/j.1469-7998.2006.00120.x
Taylor ML, Price TAR, Wedell N (2014) Polyandry in nature: a global analysis. Trends Ecol Evol 29:376–383. https://doi.org/10.1016/j.tree.2014.04.005
Theissinger K, FitzSimmons NN, Limpus CJ, Parmenter CJ, Phillott AD (2009) Mating system, multiple paternity and effective population size in the endemic flatback turtle (Natator depressus) in Australia. Conserv Genet 10:329–346. https://doi.org/10.1007/s10592-008-9583-4
Trivers RL (1972) Parental investment and sexual selection. In: Campbell B (ed) Sexual selection and the descent of man, 1871–1971. Aldine-Altherton, Chicago, pp 136–179
Tuberville TD, Norton TM, Waffa BJ, Hagen C, Glenn TC (2011) Mating system in a gopher tortoise population established through multiple translocations: apparent advantage of prior residence. Biol Conserv 144:175–183. https://doi.org/10.1016/j.biocon.2010.08.012
Tummers B (2006) Data Thief III. https://datathief.org/. Accessed 17 Mar 2012
Turkozan O, Karaman S, Yılmaz C, Beşer N (2019) Multiple paternity at the largest green turtle (Chelonia mydas) rookery in the Mediterranean. Reg Stud Mar Sci 31:100777. https://doi.org/10.1016/j.rsma.2019.100777
Uller T, Olsson M (2008) Multiple paternity in reptiles: patterns and processes. Mol Ecol 17:2566–2580. https://doi.org/10.1111/j.1365-294X.2008.03772.x
Ursenbacher S, Erny C, Fumagalli L (2009) Male reproductive success and multiple paternity in wild, low-density populations of the adder (Vipera berus). J Hered 100:365–370. https://doi.org/10.1093/jhered/esn104
Viechtbauer W (2010) Conducting meta-analyses in R with the metafor package. J Stat Softw 36:1–48. https://doi.org/10.18637/jss.v036.i03
Wapstra E, Olsson M (2014) The evolution of polyandry and patterns of multiple paternity in lizards. In: Rheubert J, Siegel D, Trauth S (eds) Reproductive biology and phylogeny of lizards and tuatara. CRC Press, Boca Raton, FL, USA, pp 576–601
While GM, Uller T, Bordogna G, Wapstra E (2014) Promiscuity resolves constraints on social mate choice imposed by population viscosity. Mol Ecol 23:721–732. https://doi.org/10.1111/mec.12618
Whitaker N (2006) Immaculate conception, incubation protocols, and egg characteristics of the Ganges softshell turtle (Aspideretes gangeticus). Contemp Herpetol 2006:1–6
White KN, Rothermel BB, Zamudio KR, Tuberville TD (2018) Male body size predicts reproductive success but not within-clutch paternity patterns in gopher tortoises (Gopherus polyphemus). J Hered 109:791–801. https://doi.org/10.1093/jhered/esy036
Wright LI, Fuller WJ, Godley BJ, McGowan A, Tregenza T, Broderick AC (2013) No benefits of polyandry to female green turtles. Behav Ecol 24:1022–1029. https://doi.org/10.1093/beheco/art003
Zajdel J, Lance SL, Rainwater TR, Wilkinson PM, Hale MD, Parrott BB (2019) Mating dynamics and multiple paternity in a long-lived vertebrate. Ecol Evol 9:10109–10121. https://doi.org/10.1002/ece3.5438
Zbinden JA, Largiadèr CR, Leippert F, Margaritoulis D, Arlettaz R (2007) High frequency of multiple paternity in the largest rookery of Mediterranean loggerhead sea turtles. Mol Ecol 16:3703–3711. https://doi.org/10.1111/j.1365-294X.2007.03426.x
Acknowledgements
We thank Marco Festa-Bianchet, Birgit Szabo, Melissa Peignier and the two anonymous referees for constructive comments and invaluable advice in improving the manuscript.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions KPP was funded by Science Foundation Ireland.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Communicated by M. Festa-Bianchet
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is a contribution to the Topical Collection Measuring individual reproductive success in the wild. Guest Editors: Janet Mann and Marco Festa-Bianchet
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, P.L.M., Sherman, C.D.H., Rollins, L.A. et al. Do female amphibians and reptiles have greater reproductive output if they have more mates?. Behav Ecol Sociobiol 76, 96 (2022). https://doi.org/10.1007/s00265-022-03194-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00265-022-03194-6