Abstract
Purpose
This study investigated the correlations between parameters of 18F-fluorodeoxyglucose (FDG) uptake on positron emission tomography (PET) scan and indices of genetic properties, heterogeneity index (HI), and tumor mutation burden (TMB), in patients with lung cancer.
Methods
We produced 106 PET indices for each tumor site that underwent genomic analysis in a total of 176 study subjects (age, 62.0 ± 10.0 y; males, 68.2%), comprising 101 adenocarcinoma (ADC), 29 squamous cell carcinoma (SQCC), and 46 small cell lung cancer (SCLC) patients. We then examined the correlations of the PET parameters with genetic properties of HI and TMB, according to pathology and tumor site.
Results
Comparisons between PET parameters and the genetic properties with false discovery rate (FDR) correction revealed that the surface standard uptake value (SUV) entropy of SUV statistics had a significant correlation with HI only in patients with SCLC who underwent a genetic test in lymph nodes (r = 0.592, p = 0.028), whereas PET parameters did not show a significant correlation with HI or TMB in patients with SCLC who underwent a genetic test in lung tissue. In patients with ADC and SQCC, there was no significant correlation between PET parameters and the genetic properties. Although SUVmax showed raw p values less than 0.05 in correlation with HI (r = 0.315, raw p = 0.048) and TMB (r = 0.206, raw p = 0.043) in ADC, and SUVpeak had a raw p value less than 0.05 in correlation with HI (r = 0.394, raw p = 0.046) in SQCC, these parameters were not significant when corrected by FDR.
Conclusions
In this study, surface SUV entropy had a significant correlation with HI in SCLC. Regarding other PET parameters and tumors, no significant correlation with genetic parameters existed.

Similar content being viewed by others
References
Bai HX, Lee AM, Yang L, Zhang P, Davatzikos C, Maris JM, et al. Imaging genomics in cancer research: limitations and promises. Br J Radiol. 2016;89:20151030.
Jaffe CC. Imaging and genomics: is there a synergy? Radiology. 2012;264:329–31.
Mazurowski MA. Radiogenomics: what it is and why it is important. J Am Coll Radiol. 2015;12:862–6.
Andreassen CN, Schack LM, Laursen LV, Alsner J. Radiogenomics - current status, challenges and future directions. Cancer Lett. 2016;382:127–36.
Sala E, Mema E, Himoto Y, Veeraraghavan H, Brenton JD, Snyder A, et al. Unravelling tumour heterogeneity using next-generation imaging: radiomics, radiogenomics, and habitat imaging. Clin Radiol. 2017;72:3–10.
Gerlinger M, Rowan AJ, Horswell S, Math M, Larkin J, Endesfelder D, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med. 2012;366:883–92.
Burrell RA, McGranahan N, Bartek J, Swanton C. The causes and consequences of genetic heterogeneity in cancer evolution. Nature. 2013;501:338–45.
Morris LG, Riaz N, Desrichard A, Senbabaoglu Y, Hakimi AA, Makarov V, et al. Pan-cancer analysis of intratumor heterogeneity as a prognostic determinant of survival. Oncotarget. 2016;7:10051–63.
Liu J, Dang H, Wang XW. The significance of intertumor and intratumor heterogeneity in liver cancer. Exp Mol Med. 2018;50:e416.
Goodman AM, Kato S, Bazhenova L, Patel SP, Frampton GM, Miller V, et al. Tumor mutational burden as an independent predictor of response to immunotherapy in diverse cancers. Mol Cancer Ther. 2017;16:2598–608.
Nair VS, Gevaert O, Davidzon G, Napel S, Graves EE, Hoang CD, et al. Prognostic PET 18F-FDG uptake imaging features are associated with major oncogenomic alterations in patients with resected non-small cell lung cancer. Cancer Res. 2012;72:3725–34.
Nair VS, Gevaert O, Davidzon G, Plevritis SK, West R. NF-kappaB protein expression associates with (18)F-FDG PET tumor uptake in non-small cell lung cancer: a radiogenomics validation study to understand tumor metabolism. Lung Cancer. 2014;83:189–96.
Gevaert O, Xu J, Hoang CD, Leung AN, Xu Y, Quon A, et al. Non-small cell lung cancer: identifying prognostic imaging biomarkers by leveraging public gene expression microarray data--methods and preliminary results. Radiology. 2012;264:387–96.
Hyun SH, Kim HS, Choi SH, Choi DW, Lee JK, Lee KH, et al. Intratumoral heterogeneity of (18)F-FDG uptake predicts survival in patients with pancreatic ductal adenocarcinoma. Eur J Nucl Med Mol Imaging. 2016;43:1461–8.
Fang YH, Lin CY, Shih MJ, Wang HM, Ho TY, Liao CT, et al. Development and evaluation of an open-source software package "CGITA" for quantifying tumor heterogeneity with molecular images. Biomed Res Int. 2014;2014:248505.
Shin HT, Choi YL, Yun JW, Kim NKD, Kim SY, Jeon HJ, et al. Prevalence and detection of low-allele-fraction variants in clinical cancer samples. Nat Commun. 2017;8:1377.
Magurran A. Measuring biological diversity. Oxford: Blackwell; 2004.
Sottoriva A, Kang H, Ma Z, Graham TA, Salomon MP, Zhao J, et al. A big bang model of human colorectal tumor growth. Nat Genet. 2015;47:209–16.
Chow D, Chang P, Weinberg BD, Bota DA, Grinband J, Filippi CG. Imaging genetic heterogeneity in glioblastoma and other glial tumors: review of current methods and future directions. AJR Am J Roentgenol. 2018;210:30–8.
Rosenberg JE, Hoffman-Censits J, Powles T, van der Heijden MS, Balar AV, Necchi A, et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet. 2016;387:1909–20.
Hatt M, Tixier F, Pierce L, Kinahan PE, Le Rest CC, Visvikis D. Characterization of PET/CT images using texture analysis: the past, the present... Any future? Eur J Nucl Med Mol Imaging. 2017;44:151–65.
Hatt M, Cheze-le Rest C, van Baardwijk A, Lambin P, Pradier O, Visvikis D. Impact of tumor size and tracer uptake heterogeneity in (18)F-FDG PET and CT non-small cell lung cancer tumor delineation. J Nucl Med. 2011;52:1690–7.
Brooks FJ, Grigsby PW. The effect of small tumor volumes on studies of intratumoral heterogeneity of tracer uptake. J Nucl Med. 2014;55:37–42.
Forgacs A, Pall Jonsson H, Dahlbom M, Daver F, M DD, Opposits G, et al. A study on the basic criteria for selecting heterogeneity parameters of F18-FDG PET images. PLoS One. 2016;11:e0164113.
Bashir U, Siddique MM, McLean E, Goh V, Cook GJ. Imaging heterogeneity in lung cancer: techniques, applications, and challenges. AJR Am J Roentgenol. 2016;207:534–43.
Higashi K, Ueda Y, Ayabe K, Sakurai A, Seki H, Nambu Y, et al. FDG PET in the evaluation of the aggressiveness of pulmonary adenocarcinoma: correlation with histopathological features. Nucl Med Commun. 2000;21:707–14.
Deng SM, Zhang W, Zhang B, Chen YY, Li JH, Wu YW. Correlation between the uptake of 18F-Fluorodeoxyglucose (18F-FDG) and the expression of proliferation-associated antigen Ki-67 in Cancer patients: a meta-analysis. PLoS One. 2015;10:e0129028.
Hatt M, Majdoub M, Vallieres M, Tixier F, Le Rest CC, Groheux D, et al. 18F-FDG PET uptake characterization through texture analysis: investigating the complementary nature of heterogeneity and functional tumor volume in a multi-cancer site patient cohort. J Nucl Med. 2015;56:38–44.
Cook GJ, O'Brien ME, Siddique M, Chicklore S, Loi HY, Sharma B, et al. Non-small cell lung Cancer treated with Erlotinib: heterogeneity of (18)F-FDG uptake at PET-association with treatment response and prognosis. Radiology. 2015;276:883–93.
Hatt M, Lee JA, Schmidtlein CR, Naqa IE, Caldwell C, De Bernardi E, et al. Classification and evaluation strategies of auto-segmentation approaches for PET: report of AAPM task group no. 211. Med Phys. 2017;44:e1–e42.
Galavis PE, Hollensen C, Jallow N, Paliwal B, Jeraj R. Variability of textural features in FDG PET images due to different acquisition modes and reconstruction parameters. Acta Oncol. 2010;49:1012–6.
Desseroit MC, Tixier F, Weber WA, Siegel BA, Cheze Le Rest C, Visvikis D, et al. Reliability of PET/CT shape and heterogeneity features in functional and morphologic components of non-small cell lung Cancer tumors: a repeatability analysis in a prospective multicenter cohort. J Nucl Med. 2017;58:406–11.
Acknowledgements
The authors thank Kyunga Kim and Min-Ji Kim from the Statistics and Data Center, Research Institute for Future Medicine, Samsung Medical Center for their important contributions to our statistical analysis. They also thank Yu-Hua Fang from Department of Biomedical Engineering, National Cheng Kung University and Hongyoon Choi from Department of Nuclear Medicine, Seoul National University Hospital for their important contributions to our imaging analysis.
Funding
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIP) (No.NRF-2016R1C1B2013411).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
The institutional review board approved that the requirement for written informed consents were waived in this study.
Electronic supplementary material
Supplemental Figure 1
The variance of FDG PET features. CV of non-texture features (a) and texture features (b) which were obtained from nine different volumetric tumor segmentations on PET/CT are plotted. CV, Coefficient of variance (PNG 989 kb)
Supplemental Figure 2
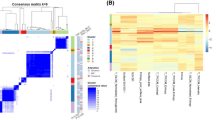
Visualization of the correlation between genetic parameters and image features of PET/CT. Heatmaps of correlation values of 30 PET features with HI in 113 subjects (a) and TMB in 176 subjects (b) according to tumor pathology and site are illustrated. HI, heterogenetic index; TMB, tumor mutation burden (JPEG 178 kb)
Supplemental Table 1
(DOCX 32 kb)
Rights and permissions
About this article
Cite this article
Moon, S.H., Kim, J., Joung, JG. et al. Correlations between metabolic texture features, genetic heterogeneity, and mutation burden in patients with lung cancer. Eur J Nucl Med Mol Imaging 46, 446–454 (2019). https://doi.org/10.1007/s00259-018-4138-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-018-4138-5