Abstract
Objectives
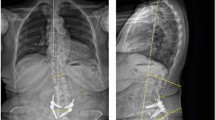
The purpose of this study was to determine whether machine learning algorithms can be utilized to optimize the hanging protocol of lumbar spine radiographs. Specifically, we explored whether machine learning models can accurately label lumbar spine views/positions, detect hardware, and rotate the lateral views to straighten the image.
Methods
We identified 1727 patients with 6988 lumbar spine radiographs. The view (anterior-posterior, right oblique, left oblique, left lateral, right lateral, left lumbosacral or right lumbosacral), hardware (present or not present), dynamic position (neutral, flexion, or extension), and correctional rotation of each radiograph were manually documented by a board-certified radiologist. Various output metrics were calculated, including area under the curve (AUC) for the categorical output models (view, hardware, and dynamic position). For non-binary categories, an all-versus-other technique was utilized designating one category as true and all others as false, allowing for a binary evaluation (e.g., AP vs. non-AP or extension vs. non-extension). For correctional rotation, the degree of rotation required to straighten the lateral spine radiograph was documented. The mean absolute difference was calculated between the ground truth and model-predicted value reported in degrees of rotation. Ensembles of the rotation models were created. We evaluated the rotation models on 3 test dataset splits: only 0 rotation, only non-0 rotation, and all cases.
Results
The AUC values for the categorical models ranged from 0.985 to 1.000. For the only 0 rotation data, the ensemble combining the absolute minimum value between the 20- and 60-degree models performed best (mean absolute difference of 0.610). For the non-0 rotation data, the ensemble merging the absolute maximum value between the 40- and 160-degree models performed best (mean absolute difference of 4.801). For the all cases split, the ensemble combining the minimum value of the 20- and 40-degree models performed best (mean absolute difference of 3.083).
Conclusion
Machine learning techniques can be successfully implemented to optimize lumbar spine x-ray hanging protocols by accounting for views, hardware, dynamic position, and rotation correction.








Similar content being viewed by others
References
Krogue JD, Cheng KV, Hwang KM, Toogood P, Meinberg EG, Geiger EJ, et al. automatic hip fracture identification and functional subclassification with deep learning. arXiv [q-bio.QM]. 2019. Available from: http://arxiv.org/abs/1909.06326
Rayan JC, Reddy N, Kan JH, Zhang W, Annapragada A. Binomial classification of pediatric elbow fractures using a deep learning multiview approach emulating radiologist decision making. Radiology: Artificial Intelligence. Radiol Soc N Am. 2019:1, e180015.
Thian YL, Li Y, Jagmohan P, Sia D, Chan VEY, Tan RT. Convolutional neural networks for automated fracture detection and localization on wrist radiographs. Radiology: Artificial Intelligence. Radiol Soc N Am. 2019:1, e180001.
Olczak J, Fahlberg N, Maki A, Razavian AS, Jilert A, Stark A, et al. Artificial intelligence for analyzing orthopedic trauma radiographs. Acta Orthop. 2017;88:581–6.
Gale W, Oakden-Rayner L, Carneiro G, Bradley AP, Palmer LJ. Detecting hip fractures with radiologist-level performance using deep neural networks. arXiv [cs.CV]. 2017. Available from: http://arxiv.org/abs/1711.06504
Kitamura G. Deep learning evaluation of pelvic radiographs for position, hardware presence, and fracture detection. Eur J Radiol. 2020;130:109139.
Kitamura G, Chung CY, Moore BE 2nd. Ankle Fracture detection utilizing a convolutional neural network ensemble implemented with a small sample, de novo training, and multiview incorporation. J Digit Imaging. 2019;32:672–7.
Kim JR, Shim WH, Yoon HM, Hong SH, Lee JS, Cho YA, et al. Computerized bone age estimation using deep learning based program: evaluation of the accuracy and efficiency. AJR Am J Roentgenol. 2017;1–7.
Tiulpin A, Klein S, Bierma-Zeinstra SMA, Thevenot J, Rahtu E, van Meurs J, et al. Multimodal machine learning-based knee osteoarthritis progression prediction from plain radiographs and clinical data. Sci Rep. 2019;9:20038.
Zhang B, Yu K, Ning Z, Wang K, Dong Y, Liu X, et al. Deep learning of lumbar spine X-ray for osteopenia and osteoporosis screening: a multicenter retrospective cohort study. Bone. 2020;140:115561.
Lee S, Choe EK, Kang HY, Yoon JW, Kim HS. The exploration of feature extraction and machine learning for predicting bone density from simple spine X-ray images in a Korean population. Skeletal Radiology. 2020;613–8. https://doi.org/10.1007/s00256-019-03342-6
Watanabe K, Aoki Y, Matsumoto M. An Application of Artificial Intelligence to Diagnostic Imaging of Spine Disease: Estimating Spinal Alignment From Moiré Images. Neurospine. 2019;16:697–702.
Kim KC, Cho HC, Jang TJ, Choi JM, Seo JK. Automatic detection and segmentation of lumbar vertebrae from X-ray images for compression fracture evaluation. Comput Methods Prog Biomed. 2020;105833. https://doi.org/10.1016/j.cmpb.2020.105833
Mason-Medical Physics D, 2011. SU-E-T-33: Pydicom: An Open Source DICOM Library. Wiley Online Library. 2011. Available from: http://onlinelibrary.wiley.com/doi/10.1118/1.3611983/full.
Abadi M, Agarwal A, Barham P, Brevdo E, Chen Z, Citro C, et al. TensorFlow: large-scale machine learning on heterogeneous distributed systems. arXiv [cs.DC]. 2016. Available from: http://arxiv.org/abs/1603.04467.
Huang G, Liu Z, van der Maaten L, Weinberger KQ. Densely Connected Convolutional Networks. arXiv [cs.CV]. 2016. Available from: http://arxiv.org/abs/1608.06993.
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, et al. Scikit-learn: Machine Learning in Python. J Mach Learn Res. 2011;12:2825–30.
Hunter JD. Matplotlib: A 2D Graphics Environment. Comput Sci Eng IEEE Comput Soc. 2007;9:90–5.
Selvaraju, R. R., Cogswell, M., Das, A., Vedantam, R., Parikh, D., & Batra, D. Grad-cam: Visual explanations from deep networks via gradient-based localization. In: Proceedings of the IEEE international conference on computer vision. 2017. p. 618–626.
Acknowledgments
This research was supported in part by the University of Pittsburgh Center for Research Computing through the resources provided.
Funding
The project described was supported by the National Institutes of Health through Grant Number UL1 TR001857.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kitamura, G. Hanging protocol optimization of lumbar spine radiographs with machine learning. Skeletal Radiol 50, 1809–1819 (2021). https://doi.org/10.1007/s00256-021-03733-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00256-021-03733-8