Abstract
Phenyllactic acid (PLA) generally recognized as a natural organic acid shows against Vibrio parahaemolyticus activity. In this study, V. parahaemolyticus ATCC17802 (Vp17802) was cultured under the stress of 1/2MIC PLA, and then the antibacterial mechanisms were explored via transcriptomics. The minimum inhibitory concentration (MIC) of PLA against Vp17802 was 3.2 mg/mL, and the time-kill analysis resulted that Vp17802 was inhibited. PLA was able to destroy the bacterial membrane, leading to the leakage of intracellular substances and decline of ATP levels. The RNA-sequencing analysis results indicated that 1616 significantly differentially expressed genes were identified, among which 190 were up-regulated and 1426 were down-regulated. Down-regulation of the icd2 gene in the TCA cycle mediates blockage of tyrosine metabolic, arginine biosynthesis, and oxidative phosphorylation, causing insufficient energy supply of Vp17802. Moreover, PLA could cause amino acids, metal ions, and phosphate transporters to be blocked, affecting the acquisition of nutrients. The treatment by PLA altered the expression of genes encoding functions involved in quorum sensing, flagellar assembly, and cell chemotaxis pathway, which may be interfering with the biofilm formation in Vp17802, reducing cell motility. Overall, 1.6 mg/mL PLA inhibited the growth of Vp17802 by disrupting to uptake of nutrients, cell metabolism, and the formation of biofilms. The results suggested a new direction for exploring the activity of PLA against Vp17802 and provided a theoretical basis for bacterial pathogen control in the food industry.
Key points
•RNA sequencing was carried out to indicate the antibacterial mechanism of Vp17802.
•The icd2 gene in the TCA cycle mediates blockage of metabolic of Vp17802.
•The biofilm formation has interfered with 1.6 mg/mL PLA, which could reduce cell motility and virulence.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Food safety problems are becoming increasingly prominent due to foodborne pathogenic contamination. Vibrio parahaemolyticus is a gram-negative halophilic bacterium, which is mainly distributed in the marine environment, and widely exists in fresh aquatic products such as Penaeus vannamei (Feng et al. 2021), oyster (Mohamad & Li 2022) and raw salmon fillets (Fang et al. 2021). V. parahaemolyticus has a strong pathogenic ability that can cause watery diarrhea, abdominal convulsions, nausea, vomiting, and headaches in humans, leading to sepsis and even death in severe cases (Li et al. 2019). According to the Disease Control and Prevention Center, this bacterium causes 215 foodborne disease cases per year, causing food poisoning or life threats (Elmahdi et al. 2016). Wu et al. (2014) have reposed that V. parahaemolyticus caused 322 foodborne outbreaks from 2003 to 2008 in China, and the number of patients reached 9041. In Japan, V. parahaemolyticus was also associated with 20–30% of human food poisoning (Jahangir et al. 2002).
The pathogenesis mechanism of V. parahaemolyticus is mainly related to its virulence factor, including two types III secretion systems (T3SS1 and T3SS2), adhesion factor, flagellar system, biofilm, and heat-resistant direct hemolysin (TDH). T3SS1 releases essential nutrients to ensure the survival of V. parahaemolyticus, and T3SS2 or TDH can release different virulence factors leading to different pathogenicity strains (Makino et al. 2003). There is a complex regulatory network in virulence factors. Adhesion factors, flagellar system, and biofilm mediate the contact between bacteria and host, and the secretion system mediates the secretion of effector proteins to induce invasion of host cells.
Recently, many natural antimicrobial agents from plant, animal, or microbial fermentation have been developed due to the insecurity of synthetic antibacterial substances and the increasing demand of consumers (Juneja et al. 2012). Phenyllactate acid (PLA) was first discovered from Geotrichum candidum by Dieuleveux in 1998 isolated from a metabolite (Dieuleveux & Guéguen 1998). PLA is a natural organic acid with broad antibacterial activity widely found in fermented goods, metabolites of lactic acid bacteria, and natural plants (Nhung et al. 2017). It exhibits inhibitory effects on a wide range of bacteria, including Listeria monocytogenes (Dieuleveux & Guéguen 1998), Salmonella (Rodríguez et al. 2012), and Escherichia coli O157:H7 (Zheng et al. 2019). The hydrophobic benzene ring and hydrophilic carboxyl structure confer amphiphilicity on PLA, which may facilitate cross-linking with bacterial membrane lipids and membrane proteins. Furthermore, PLA has water solubility and non-toxicity, which can stimulate active immune cells to show anti-inflammatory effects in the human body (Peters et al. 2019). It has been reported that PLA has heat stability whose antimicrobial activity is unaffected even at 121 °C/20 min (Cortes et al. 2014), which can enhance the shelf-life of food. However, the activity of PLA is greatly affected by acidity, and lower pH conditions can enhance the antimicrobial activity of PLA. We previously found that 1/2MIC PLA significantly reduced the CFU of Vp17802 on the salmon during the 8-day storage, and the antibacterial activity of PLA was found to be in a dose-dependent manner (Feng et al. 2021).
The antibacterial mechanism of PLA mainly focuses on the cellular level, including cell membrane integrity destruction and genomic DNA degradation (Fang et al. 2017). Dieuleveux and Guéguen (1998) have revealed that the antibacterial mechanism of PLA is likely associated with destroying the cell wall, and Ning et al. (2020) have reported that PLA can inactivate the cell membrane integrity of Listeria monocytogenes, while for Escherichia coli, it only destroys the outer membrane. The active sites of PLA on gram-positive bacteria and gram-negative bacteria are different. Recent evidence suggests that energy metabolism disorder is a new target of PLA inhibition of Rhizopus oryzae (Fan et al. 2022). PLA has been recognized as a natural, safe, and green antibacterial substance in the food industry, a deeper understanding of the subcellular targets and antibacterial mechanisms of PLA were required for their practical applications.
Therefore, transcriptome sequencing (RNA-seq) further clarifies the antibacterial mechanism of PLA in V. parahaemolyticus, which plays a vital role in cell growth and proliferation, promoting their application in the food industry.
Materials and methods
Bacterial strain and chemical
V. parahaemolyticus ATCC 17802 (Vp17802) was obtained from the Gongbei Customs Technical Center (Zhuhai, Guangzhou), and it was stored in tryptic soy broth (TSB) supplied with 3% NaCl (TSB-s) and 25% glycerol at − 80 °C before use. The bacterium was activated by incubation at 37 °C for 24 h after streaking the stock culture onto tryptone soy agar (TSA) supplied with 3% NaCl (TSA-s, pH 7.3). Then, a single colony from TSA-s was inoculated into TSB-s to reach the stationary phase after incubation at 37 °C for 18 h. Stock solutions of phenyllactate acid (PLA, 98%; Aladdin Reagent Co., Ltd., Shanghai, China) was prepared just before use and briefly stored at 4 °C.
Determination of minimum inhibitory concentration and time-kill analysis
The minimum inhibitory concentration (MIC) of PLA against Vp17802 was determined using broth microdilution method described previously (Fang et al. 2021). The time-kill assay was carried out to measure the bactericidal effect of PLA against Vp17802. The Vp17802 was diluted to about 106 CFU/mL in the TSB-S containing PLA with the final concentrations of 0.8 mg/mL, 1.6 mg/mL, 3.2 mg/mL, and 6.4 mg/mL respectively. The sterile TSB-s without PLA was used as the control group, and all samples were incubated under microaerophilic conditions with treatment for 0 h, 4 h, 8 h, 12 h, 16 h, 20 h, and 24 h. The viable bacteria were calculated by plate counting.
Determination of cytoplasmic leakages
The bacterial suspension was filtered through a membrane of 0.22 μm pore size. The filtrate was collected, and its absorbance at 260 nm and 280 nm was determined using an ND2000C ultra-micro ultraviolet–visible spectrophotometer (Thermo Fisher Scientific (China) Co., Ltd.).
Determination of ATPase activity and ATP level
The ATPase activity was determined using an ultra-micro total ATPase test kit (Nanjing Jiancheng Bioengineering Institute, A070-1), and the ATP concentration was determined using an ATP test kit (Biyuntian Biotechnology, S0026).
Swimming test
The final concentration of 1/4 MIC and 1/2 MIC PLA were added to the TSB-S which contains 0.2% agar as the treated group, and an equal volume of sterile water as the control group. All plates were incubated at 37 ℃ for 12 h to use for swimming tests.
RNA isolation and processing
Two sets of samples were selected for transcriptomic analyses to explore the antibacterial mechanisms of PLA. The bacterial suspensions were treated with (1) CK, sterile TSB-s without antibacterial substances; (2) T, sterile TSB-s with 1/2 MIC PLA, and then incubated at 37 ℃ for 6 h. After incubation, the cell suspensions were centrifuged at 8000 g for 3 min to remove the culture medium. The obtained cell pellets were immediately inactivated with liquid nitrogen and temporarily stored at − 80℃ before the following RNA extraction.
The total RNA of Vp17802 was extracted by TRIzol reagent (Invitrogen, Carlsbad, USA). Briefly, the bacterial cells were fully ground in liquid nitrogen and the intracellular substances were released from bacterial cells by 1 mL TRIzol reagent. Two hundred microliters of chloroform (Guangzhou Chemical Reagent Factory, Guangzhou, China) was added to separate from the DNA and proteins at 12,000 rpm for 10 min. Then, the upper aqueous phase was taken and 1 mL equal volumes of isopropanol (75%, v/v, Guangzhou Chemical Reagent Factory, Guangzhou, China) were added to precipitate total RNA under vacuum. The quality and integrity of total RNA were assessed using an Agilent 2100 Bio-analyzer (Agilent Technologies, Palo Alto, USA) and RNase-free agarose gel electrophoresis. After total RNA extraction, the ribosomal RNA was removed by Ribo-ZeroTM Magnetic Kit (Epicentre, Madison, USA), and the mRNA was fragmented into short fragments with 200–700 nt using a fragmentation buffer. The mRNA fragments were reversed transcription into cDNA with random hexamer primers, and the second strand of cDNA was synthesized with dNTP to promote DNA polymerase I. The synthesized cDNA fragments were purified using a QiaQuick PCR extraction kit (Qiagen, Venlo, the Netherlands) and then subjected to end repair, the ligation of poly (A), and sequencing adapters. The ligated cDNA fragments were size-screened by the agarose gel electrophoresis. The selected cDNA fragments were amplified and sequenced using Illumina HiSeq2500 by Gene Denovo Biotechnology Co. (Guangzhou, China).
Read alignment and normalization of gene expression levels
The high-quality reads were conducted using in-house Perl scripts to remove adapters or low-quality bases. The reads containing adapters, more than 10% of unknown nucleotides, and more than 50% of low quality (Q-value ≤ 20) bases) were removed. Short read alignment tool Bowtie2 was used to exclude the reads mapped to the rRNA database. The remaining clean reads were then mapped to the Vp17802 RIMD 2210633 (NCBI accession: NC_004603.1) using TopHat2. The gene expression levels were normalized using StringTie software. The gene expression abundance was quantified using fragments per kilobase of transcript per million (FPKM) method.
Differentially expressed genes and enrichment analysis
Bioconductor edgeR package (http://www.rproject.org/) was used to identify differentially expressed genes (DEGs) between the samples. The criteria of significant differential expression were FDR ≤ 0.05 and the absolute value of log2 (FC) ≥ 2. Significantly enriched GO terms and KEGG pathways were obtained with the criteria of an adjusted p-value ≤ 0.05. The gene ontology (GO) was applied to define and standardize the function of genes and proteins, including biological process (BP), molecular function (MF), and cellular component (CC). All the expressed genes were mapped to the gene ontology database (http://www.geneontology.org/), and the number of genes mapped to each GO term was calculated. The Kyoto Encyclopedia of Genes and Genomes (KEGG) (http://www.genome.jp/kegg) is the core of the database that was significantly enriched in DEGs compared with the whole genomics defined.
Data analysis
Data are presented as means ± SDs from at least three independent experiments. Statistical significance was determined using a t-test and Waller–Duncan multiple comparison test using GraphPad Prism 5 to analyze the experimental data. Independent sample.
Results
The MIC and time-kill curve
The MIC of Vp17802 treated with different PLA concentrations after 24 h is summarized in Fig. 1A. Compared to the control group, the OD600 of Vp17802 was reduced when exposed to PLA of different concentrations with the increase of incubate time. Besides, 3.2 mg/mL PLA could reduce OD600 of Vp17802 by nearly 0, which can be considered as the MIC. This MIC was consistent with our previous studies (Fang et al. 2021). In addition, the viable bacterial were increased in 1/4MIC and 1/2MIC within 4 h; however, the viable bacterial of MIC and 2MIC groups were completely inactivated (Fig. 1B). These results demonstrated that the antibacterial activity of PLA was found to be in a dose-dependent manner. Moreover, 1/4 MIC and 1/2 MIC of PLA were inactivated gradually after 4 h.
PLA-induced bacterial cytoplasmic material leakages
The cell membrane destroyed will lead to membrane depolarization, membrane destruction, and membrane pore formation, causing the leakage of cytoplasmic materials. Previous studies have proposed that PLA can produce ROS, which increases membrane damage and leads to Vp17802 death (Fang et al. 2021). To further investigate the bacterial membrane damage of PLA in Vp17802, the OD260 and OD280 were analyzed using an ultra-micro ultraviolet–visible spectrophotometer. As shown in Fig. 3C, compared with the control group, the OD value of PLA-treated showed a significant increase, indicating PLA could lead to cytoplasmic leakage in Vp17802.
PLA-induced bacterial ATP level and inhibited swimming ability
The energy conversion of cells and the secretion of various enzymes depend on the cell membrane. We have demonstrated that PLA can cause Vp17802 membrane damage, which may affect the ATP level. As shown in Fig. 2A and B, compared with the control group, the ATPase activity and ATP concentration were significantly reduced by 1/2MIC and MIC of PLA. In addition, our previous studies also showed that Vp17802 membrane damage can reduce the ability of cells to form biofilms (Fang et al. 2021). As shown in Fig. 3, we could find that the swimming ability of 1/2MIC and 1/4MIC significantly declined than the control group. It is obvious that the motility of Vp17802 was significantly reduced by PLA treatment.
DEG analysis
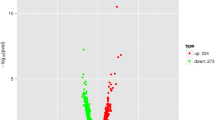
Transcriptomic profiling experiments demonstrated that 1616 DEGs were identified, among which 190 genes were up-regulated and 1426 were down-regulated (Fig. 4c). The Venn diagram shows the number of DEGs of different comparisons (Fig. 4b). Of these, 190 genes were specifically expressed only in the CK group, while 25 genes were differentially expressed only in the T group. The results show that multiple layers of response mechanism may be applied by Vp17802 under PLA stress. Further study of the inhibition mechanism from GO and KEGG results is necessary.
Differentially expressed genes of Vp between different treatments. a Volcano plots of DEGs of CK-vs-T; b Venn diagram of DEGs of CK-vs-T; c the number of up-regulated and down-regulated genes of CK-vs-T. CK: the sterile TSB-s without PLA; T: 1/2 MIC PLA. The criteria of DEGs: | log2 (Fold change) |≥ 2, FDR ≤ 0.05
GO enrichment analysis
The GO enrichment analysis of DEGs was included biological processes, cellular components, and molecular functions. As shown in Fig. 5, BP of CK-T was enriched with a total of 955 differentially expressed genes, mainly involving metabolic process (up, 69; down, 181), cellular process (up, 67; down, 181), organism process (up, 49; down, 165), localization (up, 16; down, 83), and biological regulation (up, 8; down, 64). MF of CK-T was enriched with a total of 490 differentially expressed genes, mainly involving catalytic activity (up, 59; down, 156), binding (up, 49; down, 129), and transporter activity (up, 8; down, 36). CC of CK-T was enriched with a total of 498 differentially expressed genes, main involving cell (up, 38; down, 97), cell part (up, 38; down, 97), membrane (up, 12; down, 86), and membrane part (up, 11; down, 67). Compared with the CK group, the biological process of citric acid and tricarboxylic acid metabolism and the cellular component of cellular ribosomes and ribonucleoprotein complexes were significantly enriched in Vp17802 treated with PLA (Q-value ≤ 0.05). The results demonstrated that PLA exhibited an effect on the transcription of Vp17802 mainly altering its BP and CC.
KEGG analysis
The KEGG pathways analysis of DEGs was assisted to further analyze and understand and explore the antibacterial mechanism of PLA in Vp17802. In the results, the metabolism, cellular, genetic information processing, environmental processing, and organismal systems were identified under the PLA treatment in Vp17802 (Fig. 6b). Meanwhile, the top seven significant KEGG pathways were enriched, including carbon fixation pathways in prokaryotes, two-component system, the tricarboxylic acid (TCA) cycle, bacterial chemotaxis, carbon metabolism, methane metabolism, and pyruvate metabolism (Fig. 6a). The carbon fixation pathways, two-component system, citrate cycle, bacterial chemotaxis, and carbon metabolism were significantly enriched in Vp17802 treated with PLA (Q-value ≤ 0.05).
Discussion
Antibacterial effect of PLA against Vp17802
Studies have shown that the MIC of PLA against Vp17802 is 3.2 mg/mL, and other research also revealed the MIC of PLA against bacterial pathogens is 1.5 to 5.0 mg/mL (Ning et al. 2017). These results elucidate that PLA has efficient antibacterial activity on Vp17802. It was previously found that intracellular ROS level in Vp17802 was increased by PLA-treated (Fang et al. 2021), and a large amount of cytoplasmic material leakages further confirmed that PLA caused oxidative damage to cells and affected bacterial membrane integrity (Fig. 2C). Fang et al. (2021) have also reported that damage of the membrane of Vp17802 was treated by PLA. The change in bacterial ATPase activity after PLA treatment also supports these results (Fig. 2A and B). The bacterial membrane was destroyed, which could reduce the ATP synthesis, resulting in a decrease in ATPase activity. In addition, Wadhwa and Berg (2021) have shown that the ATPase activity is related to bacterial motility, which is able to reduce the bacterial swimming ability.
Transcriptomics analysis by RNA-seq
Impairing the energy metabolism
The related pathway of tyrosine metabolism, arginine biosynthesis, and oxidative phosphorylation in the tricarboxylic acid (TCA) cycle has been significantly regulated (Fig. 6). The TCA cycle, known as the citrate cycle, is the most important part of supplying energy for the metabolic hub of a cell. The NADP-dependent isocitrate dehydrogenase (icd2) is a key enzyme involved in cell metabolism and energy generation through the TCA cycle, which can regulate the oxidative respiration and normal operation of cells. It has been demonstrated that icd2 has reducing power to resist oxidative damage, and product virulence factors to adapt to the change of external environment (Chiang et al. 2017; Lv et al. 2016). In mycobacteria, the icd2 is responsible for carbon flux through the oxidative TCA cycle to ensure a balance between energy production and precursor biosynthesis by the glyoxylate shunt enzyme isocitrate lyase (Sharmistha et al. 2005). We found that PLA was able to down-regulate the expression of icd2, which means the energy metabolism of Vp17802 was destroyed (Fig. 7).
Meanwhile, the 2-oxoglutarate dehydrogenase E1 component (sucA) and 2-oxoglutarate dehydrogenase complex dihydrolipoyllysine-residue succinyltransferas (sucB) are important enzymes to convert 2-Oxo-glutarate to Succinyl-CoA in the TCA cycle, which were down-regulated by 2.4 log2 (FC) and 2.51 log2 (FC), respectively. The ketoglutarate dehydrogenase gene cluster sucABCD and the succinyl-CoA synthase gene cluster sdhABCDE were down-regulated by more than 2 log2 (FC) (Table S1). These results further indicate that PLA is able to impact the energy metabolism of Vp17802 and inhibit cell growth. The aconitate hydratase B (acnB) was down-regulated by 1.53 log2 (FC). Several researches have shown that the acnB mutations can lead to decreases in oxidative respiration and increases in antibiotic tolerance, while the changes of the sucC do not affect (Barton et al. 2021).
The pyruvate kinase of pykF and pykA was significantly regulated which affected the degradation of glucose into pyruvate (Table S1). The pykF was up-regulated by 1.35 log2 (FC), while the pykA was down-regulated by 1.06 log2 (FC). The studies have reported that pykF is a key upstream kinase to catalyze phosphoenolpyruvate (PEP) producing ATP and promoting energy metabolism causing the sensitivity of drug-resistant strains (Zuye et al. 2022). Abdelhamid et al. (2019) have demonstrated that pykA plays the dominant role in glycolysis, which is relevant for Pseudomonas aeruginosa virulence, and it is able to influence bacterial pathogenicity. In addition, the glucose-specific IIB component (ptsG) was up-regulated by 2.00 log2 (FC) in the glycolytic pathway; likewise, the gapA encode glyceraldehyde 3-phosphate dehydrogenase was up-regulated by 2.25 log2 (FC). We conclude that the up-regulated gene in the glycolytic pathway may maintain the normal physiology of Vp17802, but the treatment with PLA ultimately causes unbalanced metabolic in the TCA cycle.
Impeding the prokaryotic-type ABC transporters
ABC transporters have been identified as membrane-bound proteins, which use ATP hydrolysis-generated energy to complete transmembrane transport. The substances were transported by the ABC transport system, including minerals, organic ions, phosphates, amino acids, metallic cations, iron-siderophores, vitamins, carbohydrates, and proteins. It is important to mediate the uptake of bacterial nutrients, and the discharge of metabolites and toxic substances. This system functions significantly altered 14 genes that were down-regulated mainly related to mineral and organic ion transporters (modABCD, fbpC, proXW), phosphate and amino acid transporters (pstSAB, aapJQMP), and metallic cation, iron-siderophore, and vitamin B12 transporters (znuA) (Fig. 7).
The ions of iron and zinc are a class of mineral and organic ions, which can be used as important prosthetic groups to inhibit oxidative stress, prevent the production of reactive oxygen species, and reduce cytotoxicity in bacteria. Among them, iron ions are also involved in the tricarboxylic acid cycle and DNA synthesis. On the contrary, zinc ions play a role in cellular metabolism, defense, repair, and other processes by combining with proteins. According to Fig. 2, PLA is able to reduce ATP levels in Vp17802. Based on the result of RNA-seq, the fbpC was notably down-regulated by 1.34 log2 (FC), and the encoding superoxide dismutase of sodAC was also down-regulated by 3.45 log2 (FC) and 3.12 log2 (FC), respectively. The functional ATPase fbpC transported extracellular iron ions into the cells via combing with fbpB to form a ferroportin. Moreover, the pstS was notably down-regulated by 1.25 log2 (FC), which hindered the expression of pstA and further caused the down-regulation of pstB. The bacterial resistance was reduced when there were low expression levels of pstB. The metabolism and balance of phosphorus were controlled by phosphate and amino acid transports, in which pstS is a peripheral binding protein and pstB is an ATP binding protein. Low expression levels of pstB would lead to decreased ATP binding sites and ATP hydrolysis, which cannot provide power for phosphate and amino acid transports. In addition, the genes of aapJQMP mediated the transport of general L-amino acids, which plays a vital role in the metabolic activities of bacteria. In our study, the aapJQMP were significantly inhibited which were down-regulated by 3.10 log2 (FC), 3.14 log2 (FC), 2.37 log2 (FC), and 2.94 log2 (FC), respectively. These phenomena suggested that PLA could cause cell membrane damage, affecting the ATPase activity, further blocking organic iron, phosphate, and amino acid transport, causing damage to Vp17802.
Inhibiting biofilm formation
Biofilm formation is related to quorum sensing (QS), flagellar assembly, cell chemotaxis, and bacterial motility. Previous studies have shown that the biofilm formation was reduced by PLA in Vp17802 (Fang et al. 2021). According to Table S1, it can be found that genes related to quorum sensing pathway (luxO, luxR, luxS, luxQ, and cqsS) were down-regulated by 1.04 ~ 2.57 log2 (FC) and genes related to c-di-GMP signaling pathway (cdgJ, dgcH, dgcC, and mbaA) down-regulated expression by 1.78 ~ 3.32 log2 (FC). In the QS system, bacteria communicate with other bacteria to adjust the expression of genes regulating their growth, adhesion, migration, and biofilm formation. The QS of Vp17802 can produce three kinds of signaling molecules, namely harveyi autoinducer-1 (HAI-1), autoinducer-2 (AI-2), and cholera autoinducer-1 (CAI-1), and the luxN, luxP, luxQ, and cqsS are receptor proteins (Lu et al. 2019). When the external environment changes, the expression of quorum sensing signaling molecules is activated to form polysaccharide-protein complexes further forming a biofilm. In addition, the c-di-GMP signaling pathway is also closely related with biofilm formation. c-di-GMP can inhibit the activity of ATPase to adjust the function of fleQ, which is the main regulator of flagellar synthesis, thereby inhibiting the flagellar assembly and reducing bacterial motility. c-di-GMP can also regulate the production of exopolysaccharides, affecting the maturation of biofilms (GodePotratz et al. 2011). Wang et al. (2021) found that carvacrol can bind to the regulator gene of luxR to block the interaction with signaling molecule C4-HSL and interfere regulatory function of the QS. The results of the transcriptomic analysis also showed that carvacrol reduced the expression of genes related to flagella assembly, chemotaxis, luxR-mediated QS, and c-di-GMP. Therefore, treatment with PLA in Vp17802 may inhibit the QS signal and c-di-GMP signal transduction, reducing the ability to form biofilm.
Type II secretory system (T2SS) related genes (epsDEFGIJKLM) were down-regulated by 1.04–1.86 log2 (FC). T2SS is one of the important virulence factors of gram-negative bacteria, responsible for transporting virulence proteins from the periplasmic space to the outer membrane. The T2SS of bacteria have been studied about Vibrio cholerae, Escherichia coli, Pseudomonas aeruginosa, Klebsiella, Legionella pneumophila, and Yersinia enterocolitica; among them, Vibrio cholerae is highly homology with Vp17802 (Abendroth et al. 2009). The pseudopilus, outer membrane complex, inner membrane platform, and ATPase consisted of T2SS, which contains 12 to 15 different types of proteins. The pseudopilus includes a large pseudopilus epsG, and four small pseudopilus epsH, epsI, epsJ, and epsK. A core component of the outer membrane complex is epsD, which controls the closure and opening of outer membrane proteins during secretion. The inner membrane is composed of three proteins, epsL, epsM, and epsF. The secretion mechanism of T2SS is that the precursor substance of the effector excises the signal peptide through the Sec system and is further folded and modified to be secreted out of the cell. The proteins secreted by T2SS mainly include protease, pectinase, cellulase, and toxin. Both thermostable direct hemolysin (TDH) and thermostable related hemolysin (TRH) of Vp17802 mainly depend on T2SS secretion. In this study, down-regulated expression of T2SS-related genes suggests that PLA affects the secretion of Vp17802-associated toxins, thereby reducing virulence (Fig. 7).
A total of 18 DEGs related to flagellar assembly were down-regulated by 1.04 log2 (FC) ~ 3.12 log2 (FC), including fliARQPONMGFELIGP and motXAB (Fig. 7). The polar and lateral flagellar are two types of Vp17802, driven by sodium and hydrogen ions, respectively. The flagella of polar and lateral have similar basic elements, mainly composed of basal body, hook, and flagellum. The basal body can be divided into the rotor and the stator. The rotor comprises the C ring of fliG, fliM, and fliN in the cytoplasm and the MS ring of multiple fliFs in the plasma membrane. The basal body also contains a P ring (flgI) in the peptidoglycan layer and an L ring (flgH) in the outer membrane. In addition, the polar flagellar has an H ring (flgT) and a T ring (motX and motY). The stator complex is composed of two proteins, the stator of the polar flagellar is pmoA and pmoB, and the stator of the lateral flagellar is composed of motA and motB. In this study, pmoA, motB, motY, and motX were down-regulated by 1.26 log2 (FC), 1.22 log2 (FC), 2.11 log2 (FC), and 1.59 log2 (FC), respectively, indicating that the polar flagellar and lateral flagellar of Vp17802 have been affected. According to Fig. 3, PLA is able to reduce cell motility in Vp17802. The rapid rotation of the flagellar endowed the bacteria with motility. However, the genes related to flagellar assembly were significantly down-regulated under the stress of PLA, indicating the dynamic system of the flagellar of Vp17802 was inhibited resulting in decreased motility. Studies have shown that the genes related to flagellar assembly in Vp17802, such as the gene fliG and mcps, were also significantly down-regulated by about 0.8 log2 (FC) under the treatment of sub-inhibitory concentrations of benzyl isothiocyanate (Song et al. 2019). Gao et al. (2018) confirmed that the genes flgA, flgC, and flgD were down-regulated in Vibrio anguillarum under starvation stress. Qiao et al. (2022) showed that genes related to flagellar assembly (fliC, fliK, fliG, fliN, fliH, fliI, fliJ, and fliA) were significantly enriched when Vp17802 was co-cultured with Ulva fasciata, showing a trend of down-regulation. Therefore, the flagellar system is an important target for antibacterial substances to exert antibacterial effects.
When bacteria perceive external stimuli, bacterial chemotaxis guides their trend the beneficial movement. Chemotaxis-related genes such as mcpU, mcpP, cheV, cheR, cheW, cheZ, cheY, and fliN were significantly down-regulated by 1.01–3.14 log2 (FC), indicating PLA treatment had a significant effect on the chemotaxis ability of Vp17802 (Fig. 7). The methyl-accepting chemotaxis proteins (MCPs) is able to sense changes of the external environment and stimulate signal transduction through methylation and demethylation processes. The encoding MCP genes were down-regulated by 1 ~ 3 log2 (FC) in PLA treatment, indicating the perceived ability of cells was decreased. In the bacterial chemotaxis system, the methylation level of MCPs was controlled by the methylation function of the methyltransferase cheR and the response regulator cheB, affecting the signaling activities of MCPs. cheW connects MCPs to a histidine kinase cheA, and cheA undergoes autophosphorylation reactions to transfer to cheB, cheY, and cheV. As the only phosphatase, cheZ accelerates the hydrolysis of phosphate on cheY, thereby inhibiting the function of cheY. In addition, the C-terminus of cheZ is responsible for binding to cheY and catalyzing the dephosphorylation of unbound cheY-P, when cheY-P binds to fliM, the cheZ cannot catalyze its dephosphorylation. cheY and cheY-P are able to control the direction of flagellar rotation by binding to fliM. Studies have shown that benzyl isothiocyanate also has the ability to reduce the chemotaxis of bacteria in Pseudomonas aeruginosa (Kafantaris et al. 2021). Thus, PLA inhibits the expression of the chemotactic system genes reducing the survival viability of Vp17802.
In conclusion, this study revealed the antibacterial mechanism of sublethal PLA against Vp17802 using high-throughput transcriptome sequencing technology. The results showed that PLA via multiple pathways inhibited Vp17802, including cell transportation, energy metabolism, quorum sensing, flagellar assembly, cell chemotaxis, and biofilm formation. Among them, the expression of amino acids, metal ions, and phosphate-related transporters was reduced by PLA treatment, impacting on transport capacity. The energy metabolism capacity was inhibited, which could cause decreased antioxidant capacity and toxicity. In addition, PLA effectively reduced the chemotaxis, motility, and virulence related to biofilm formation. This study provides new evidence and ideas to reveal the antibacterial mechanical of PLA on Vp17802.
Data availability
The authors declare that (the/all other) data supporting the findings of this study are available within the article. The transcriptome sequencing raw data have been uploaded to the National Microbiology Data Center (Bio Project: NMDC10018011).
References
Abdelhamid Y, Brear P, Greenhalgh J, Chee X, Rahman T, Welch M (2019) Evolutionary plasticity in the allosteric regulator-binding site of pyruvate kinase isoform PykA from Pseudomonas aeruginosa. J Biol Chem 294:15505–15516. https://doi.org/10.1074/jbc.RA119.009156
Abendroth J, Kreger AC, Wim GJ, Hol WGJ (2009) The dimer formed by the periplasmic domain of EpsL from the type 2 secretion system of Vibrio parahaemolyticus. J Struct Biol 168:313–322. https://doi.org/10.1016/j.jsb.2009.07.022
Barton B, Grinnell A, Morgenstein RM (2021) Disruption of the MreB elongasome is overcome by mutations in the tricarboxylic acid cycle. Front Microbiol 12:664281. https://doi.org/10.3389/fmicb.2021.664281
Chiang YC, Liao CT, Du SC, Hsiao YM (2017) Functional characterization and transcriptional analysis of icd2 gene encoding an isocitrate dehydrogenase of Xanthomonas campestris pv. campestris. Arch Microbiol 199:1–13. https://doi.org/10.1007/s00203-017-1370-5
Cortes ZO, Lopez MA, Hernandez MA, García HS (2014) Antifungal activity of lactobacilli and its relationship with 3-phenyllactic acid production. Int J Food Microbiol 173:30–35. https://doi.org/10.1016/j.foodcont.2021.108184
Dieuleveux V, Guéguen M (1998) Antimicrobial effects of D-3-phenyllactic acid on Listeria monocytogenes in TSB-YE medium, milk, and cheese. J Food Protect 61:1281. https://doi.org/10.4315/0362-028X-61.10.1281
Elmahdi S, DaSilva LV, Parveen S (2016) Antibiotic resistance of Vibrio parahaemolyticus and Vibrio vulnificus in various countries: a review. Food Microbiol 57:128–134. https://doi.org/10.1016/j.fm.2016.02.008
Fan W, Li B, Du N, Hui T, Cao Y, Li X, Ren H (2022) Energy metabolism as the target of 3-phenyllactic acid against Rhizopus oryzae. Int J Food Microbiol 369:109606. https://doi.org/10.1016/j.ijfoodmicro.2022.109606
Fang L, Du L, Tong Z, Ping Z, Doyle MP (2017) Effects of phenyllactic acid as sanitizing agent for inactivation of Listeria monocytogenes biofilms. Food Control 78:72–78. https://doi.org/10.1016/j.foodcont.2017.02.050
Fang M, Wang R, Agyekumwaa AK, Yu Y, Xiao X (2021) Antibacterial effect of phenyllactic acid against Vibrio parahaemolyticus and its application on raw salmon fillets. LWT-Food Sci Technol 154:112586. https://doi.org/10.1016/j.lwt.2021.112586
Feng Y, Lx A, Wh A, Fang L (2021) Highly lethal Vibrio parahaemolyticus strains cause acute mortality in Penaeus vannamei post-larvae. Aquaculture 548:737605. https://doi.org/10.1016/j.aquaculture.2021.737605
Gao X, Pi D, Chen N, Li X, Liu X, Yang H, Wei W, Zhang X (2018) Survival, virulent characteristics, and transcriptomic analyses of the pathogenic Vibrio anguillarum under starvation stress. Front Cell Infect Mi 8:389. https://doi.org/10.3389/fcimb.2018.00389
GodePotratz CJ, Kustusch RJ, Breheny PJ, Weiss DS, McCarter LL (2011) Surface sensing in Vibrio parahaemolyticus triggers a program of gene expression that promotes colonization and virulence. Mol Microbiol 79:240–263. https://doi.org/10.1111/j.1365-2958.2010.07445.x
Jahangir AM, Tomochika KI, Miyoshi SI, Shinoda S (2002) Environmental investigation of potentially pathogenic Vibrio parahaemolyticus in the Seto-Inland Sea, Japan. FEMS Microbiol Lett 208:83–87. https://doi.org/10.1111/j.1574-6968.2002.tb11064.x
Juneja VK, Hari PD, Yan X (2012) Novel natural food antimicrobials. Annu Rev Food Sci T 3:381–403. https://doi.org/10.1146/annurev-food-022811-101241
Kafantaris I, Tsadila C, Nikolaidis M, Tsavea E, Dimitriou TG, Iliopoulos I, Amoutzias GD, Mossialos D (2021) Transcriptomic analysis of Pseudomonas aeruginosa response to pine honey via RNA sequencing indicates multiple mechanisms of antibacterial activity. Foods 10:936. https://doi.org/10.3390/foods10050936
Li L, Meng H, Gu D, Li Y, Jia M (2019) Molecular mechanisms of Vibrio parahaemolyticus pathogenesis. Microbiol Res 222:43–51. https://doi.org/10.1016/j.micres.2019.03.003
Lu R, Tang H, Qiu Y, Yang W, Yang H, Zhou D, Huang X, Hu L, Zhang Y (2019) Quorum sensing regulates the transcription of lateral flagellar genes in Vibrio parahaemolyticus. Future Microbiol 14:1043–1053. https://doi.org/10.2217/fmb-2019-0048
Lv CQ, Wang P, Wang WC, Su RR, Ge YD, Zhu GM (2016) Two isocitrate dehydrogenases from a plant pathogen Xanthomonas campestris pv. campestris 8004. bioinformatic analysis, enzymatic characterization, and implication in virulence. J Basic Microb 56:975–985. https://doi.org/10.1002/jobm.201500648
Makino K, Oshima K, Kurokawa K, Yokoyama K, Uda T, Tagomori K, Lijima Y, Najima M, Nakano M, Yamashita A, Kubota Y, Kimura S, Yasunaga T, Honda T, Shinagawa H, Hattori M, Lida T (2003) Genome sequence of Vibrio parahaemolyticus: a pathogenic mechanism distinct from that of V cholerae. The Lancet 361:743–749. https://doi.org/10.1016/S0140-6736(03)12659-1
Mohamad E G & Li D (2022) Photoinactivation of bacteriophage MS2, tulane virus and Vibrio parahaemolyticus in oysters by microencapsulated rose bengal. Food Qual Saf 6:fyac017 https://doi.org/10.1093/fqsafe/fyac017
Nhung NT, Chansiripornchai N, Carrique-Mas JJ (2017) Antimicrobial resistance in bacterial poultry pathogens: a review. Front Vet Sci 4:126. https://doi.org/10.3389/fvets.2017.00126
Ning Y, Yan A, Yang K, Wang Z, Li X, Jia Y (2017) Antibacterial activity of phenyllactic acid against Listeria monocytogenes and Escherichia coli by dual mechanisms. Food Chem 228:533–540. https://doi.org/10.1016/j.foodchem.2017.01.112
Ning YW, Fu YN, Hou LL, Ma MG, Wang ZX, Li XF, Jia YM (2020) iTRAQ-based quantitative proteomic analysis of synergistic antibacterial mechanism of phenyllactic acid and lactic acid against Bacillus cereus. Food Res Int 139:109562. https://doi.org/10.1016/j.foodres.2020.109562
Peters A, Krumbholz P, Jäger E, Heintz-Buschart A, Çakir MV, Rothemund S, Gaudl A, Ceglarek U, Schöneberg T, Stäubert C (2019) Metabolites of lactic acid bacteria present in fermented foods are highly potent agonists of human hydroxycarboxylic acid receptor 3. PLoS Genet 15:e1008145. https://doi.org/10.1371/journal.pgen.1008145
Qiao Y, Feng L, Jia R, Luo Y, Yang Q (2022) Motility, biofilm formation and associated gene expression in Vibrio parahaemolyticus impaired by co-culture with live Ulva fasciata. J Appl Microbiol 132:101–112. https://doi.org/10.1111/jam.15175
Rodríguez N, Salgado JM, Cortés S, Domínguez J (2012) Antimicrobial activity of d-3-phenyllactic acid produced by fed-batch process against Salmonella enterica. Food Control 25:274. https://doi.org/10.1016/j.foodcont.2011.10.042
Sharmistha B, Ashok N, RaviPrasad P, Vishwa MK, Seyed EH (2005) Comparison of mycobacterium tuberculosis isocitrate dehydrogenases (ICD-1 and ICD-2) reveals differences in coenzyme affinity, oligomeric state, pH tolerance and phylogenetic affiliation. BMC Biochem 6:20. https://doi.org/10.1186/1471-2091-6-20
Song J, Hou HM, Wu HY, Li KX, Wang Y, Zhou QQ, Zhang GL (2019) Transcriptomic analysis of Vibrio parahaemolyticus reveals different virulence gene expression in response to benzyl isothiocyanate. Molecules 24:761. https://doi.org/10.3390/molecules24040761
Wadhwa N, Berg HC (2021) Bacterial motility: machinery and mechanisms. Nat Rev Microbiol 20:161–173. https://doi.org/10.1038/s41579-021-00626-4
Wang Y, Feng L, Lu H, Zhu J, Kumar V, Liu X (2021) Transcriptomic analysis of the food spoilers Pseudomonas fluorescens reveals the antibiofilm of carvacrol by interference with intracellular signaling processes. Food Control 127:108115. https://doi.org/10.1016/j.foodcont.2021.108115
Wu Y, Wen J, Ma Y, Ma X, Chen Y (2014) Epidemiology of foodborne disease outbreaks caused by Vibrio parahaemolyticus, China, 2003–2008. Food Control 46:197–202. https://doi.org/10.1016/j.foodcont.2014.05.023
Zheng R, Zhao T, Hung YC, Adhikari K (2019) Evaluation of bactericidal effects of phenyllactic acid on Escherichia coli O157:H7 and Salmonella Typhimurium on beef Meat. J Food Protect 82:2016–2022. https://doi.org/10.4315/0362-028X.JFP-19-217
Zuye F, Fubin L, Kun C, Ziyuan Z, Linlin C, Shiqin L, Yufeng D, Xingfeng Y, Ruiguang G, Qing YH, Xuesong S (2022) Potential role of lysine acetylation in antibiotic resistance of Escherichia coli. Msystems 20:e0064922. https://doi.org/10.1128/msystems.00649-22
Funding
This work was supported by R&D Projects in Key Areas of Guangdong Province [Nos. 2019B020212003 and 2021B0202060001], Collaborative Innovation Center of GDAAS [No. XT202303], Jieyang Science and Technology Project [No. 2022DZX018], and Science and Technology Program of Guangzhou, China [No. 202206010177].
Author information
Authors and Affiliations
Contributions
YLL: investigation; data analysis; writing—original draft. MMF: methodology; writing—original draft. JL: methodology; writing—review and editing. YHZ: supervision, funding acquisition. YGY: supervision; writing—review and editing; funding acquisition.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lin, Y., Fang, M., Liu, J. et al. Transcriptomic analyses of Vibrio parahaemolyticus under the phenyllactic acid stress. Appl Microbiol Biotechnol 108, 180 (2024). https://doi.org/10.1007/s00253-024-13024-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13024-6