Abstract
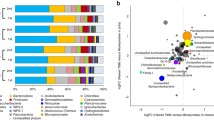
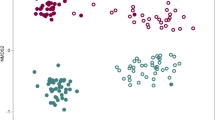
The microbial community within the root system, the rhizosphere closely connected to the root, and their symbiotic relationship with the host are increasingly seen as possible drivers of natural pathogen resistance. Resistant cultivars have the most effective strategy in controlling the Chinese wheat yellow mosaic disease, but the roles of the root and rhizosphere microbial interactions among different taxonomic levels of resistant cultivars are still unknown. Thus, we aimed to investigate whether these microbial community composition and network characteristics are related to disease resistance and to analyze the belowground plant-associated microflora. Relatively high microbial diversity and stable community structure for the resistant cultivars were detected. Comparison analysis showed that some bacterial phyla were significantly enriched in the wheat root or rhizosphere of the resistant wheat cultivar. Furthermore, the root and rhizosphere of the resistant cultivars greatly recruited many known beneficial bacterial and fungal taxa. In contrast, the relative abundance of potential pathogens was higher for the susceptible cultivar than for the resistant cultivar. Network co-occurrence analysis revealed that a much more complex, more mutually beneficial, and a higher number of bacterial keystone taxa in belowground microbial networks were displayed in the resistant cultivar, which may have been responsible for maintaining the stability and ecological balance of the microbial community. Overall, compared with the susceptible cultivar, the resistant cultivar tends to recruit more potential beneficial microbial groups for plant and rhizosphere microbial community interactions. These findings indicate that beneficial rhizosphere microbiomes for cultivars should be targeted and evaluated using community compositional profiles.
Key points
• Different resistance levels in cultivars affect the rhizosphere microbiome..
• Resistant cultivars tend to recruit more potential beneficial microbial groups.
• Bacteria occupy a high proportion and core position in the microflora network.





Similar content being viewed by others
Data availability
All data generated and/or analyzed during this study are included in this published article and its supplementary files.
Code availability
Not applicable.
References
Adams M, Jacquier C (1994) Infection of cereals and grasses by isolates of Polymyxa graminis (Plasmodiophorales). Ann Appl Biol 125(1):53–60. https://doi.org/10.1111/j.1744-7348.1994.tb04946.x
Adams MJ, Antoniw JF, Kreuze J (2009) Virgaviridae: a new family of rod-shaped plant viruses. Arch Virol 154(12):1967–1972. https://doi.org/10.1007/s00705-009-0506-6
Andika IB, Zheng SL, Tan ZL, Sun LY, Kondo H, Zhou XP, Chen JP (2013) Endoplasmic reticulum export and vesicle formation of the movement protein of Chinese wheat mosaic virus are regulated by two transmembrane domains and depend on the secretory pathway. Virology 435(2):493–503. https://doi.org/10.1016/j.virol.2012.10.024
Badri DV, Chaparro JM, Zhang RF, Shen QR, Vivanco JM (2013) Application of natural blends of phytochemicals derived from the root exudates of Arabidopsis to the soil reveal that phenolic-related compounds predominantly modulate the soil microbiome. J Biol Chem 288(7):4502–4512. https://doi.org/10.1074/jbc.M112.433300
Bakker PA, Berendsen RL, Doornbos RF, Wintermans PC, Pieterse CM (2013) The rhizosphere revisited: root microbiomics. Front Plant Sci 4:165. https://doi.org/10.3389/fpls.2013.00165
Barberán A, Bates ST, Casamayor EO, Fierer N (2012) Using network analysis to explore co-occurrence patterns in soil microbial communities. ISME J 6(2):343–351. https://doi.org/10.1038/ismej.2011.119
Bastian M, Heymann S, Jacomy M (2009) Gephi: an open source software for exploring and manipulating networks. In third international ICWSM conference,2009, San Jose, California, USA (pp. 361–362) https://www.aaai.org/ocs/index.php/ICWSM/09/paper/view/154
Berendsen RL, Pieterse CMJ, Bakker PAHM (2012) The rhizosphere microbiome and plant health. Trends Plant Sci 17(8):478–486. https://doi.org/10.1016/j.tplants.2012.04.001
Berendsen RL, Vismans G, Yu K, Song Y, de Jonge R, Burgman WP, Burmolle M, Herschend J, Bakker P, Pieterse CMJ (2018) Disease-induced assemblage of a plant-beneficial bacterial consortium. ISME J 12(6):1496–1507. https://doi.org/10.1038/s41396-018-0093-1
Bolyen E, Rideout JR, Dillon MR, Bokulich N, Abnet CC, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F, Bai Y, Bisanz JE, Bittinger K, Brejnrod A, Brislawn CJ, Brown CT, Callahan BJ, Caraballo-Rodriguez AM, Chase J, Cope EK, Da Silva R, Diener C, Dorrestein PC, Douglas GM, Durall DM, Duvallet C, Edwardson CF, Ernst M, Estaki M, Fouquier J, Gauglitz JM, Gibbons SM, Gibson DL, Gonzalez A, Gorlick K, Guo JR, Hillmann B, Holmes S, Holste H, Huttenhower C, Huttley GA, Janssen S, Jarmusch AK, Jiang LJ, Kaehler BD, Bin Kang K, Keefe CR, Keim P, Kelley ST, Knights D, Koester I, Kosciolek T, Kreps J, Langille MGI, Lee J, Ley R, Liu YX, Loftfield E, Lozupone C, Maher M, Marotz C, Martin BD, McDonald D, McIver LJ, Melnik AV, Metcalf JL, Morgan SC, Morton JT, Naimey AT, Navas-Molina JA, Nothias LF, Orchanian SB, Pearson T, Peoples SL, Petras D, Preuss ML, Pruesse E, Rasmussen LB, Rivers A, Robeson MS, Rosenthal P, Segata N, Shaffer M, Shiffer A, Sinha R, Song SJ, Spear JR, Swafford AD, Thompson LR, Torres PJ, Trinh P, Tripathi A, Turnbaugh PJ, Ul-Hasan S, vander Hooft JJJ, Vargas F, Vazquez-Baeza Y, Vogtmann E, von Hippel M, Walters W, Wan YH, Wang MX, Warren J, Weber KC, Williamson CHD, Willis AD, Xu ZZ, Zaneveld JR, Zhang YL, Zhu QY, Knight R, Caporaso JG, (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37(8):852–857. https://doi.org/10.1038/s41587-019-0209-9
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13(7):581. https://doi.org/10.1038/nmeth.3869
Chen J, MacFarlane SA, Wilson TM (1994) Detection and sequence analysis of a spontaneous deletion mutant of soil-borne wheat mosaic virus RNA2 associated with increased symptom severity. Virology 202(2):921–929. https://doi.org/10.1006/viro.1994.1414
Deng Y, Jiang Y-H, Yang Y, He Z, Luo F, Zhou J (2012) Molecular ecological network analyses. Bmc. Bioinformatics 13(1):1–20. https://doi.org/10.1186/1471-2105-13-113
Diao AP, Chen JP, Ye R, Zheng T, Yu SQ, Antoniw JF, Adams MJ (1999) Complete sequence and genome properties of Chinese wheat mosaic virus, a new furovirus from China. J Gen Virol 80:1141–1145. https://doi.org/10.1099/0022-1317-80-5-1141
Dick WA, Cheng L, Wang P (2000) Soil acid and alkaline phosphatase activity as pH adjustment indicators. Soil Biol Biochem 32(13):1915–1919. https://doi.org/10.1016/S0038-0717(00)00166-8
Dixon P (2003) VEGAN, a package of R functions for community ecology. J Veg Sci 14(6):927–930. https://doi.org/10.1111/j.1654-1103.2003.tb02228.x
Durán P, Thiergart T, Garrido-Oter R, Agler M, Kemen E, Schulze-Lefert P, Hacquard S (2018) Microbial interkingdom interactions in roots promote Arabidopsis survival. Cell 175(4):973-983.e14. https://doi.org/10.1016/j.cell.2018.10.020
Fargione JE, Tilman D (2005) Diversity decreases invasion via both sampling and complementarity effects. Ecol Lett 8(6):604–611. https://doi.org/10.1111/j.1461-0248.2005.00753.x
Ge AH, Liang ZH, Xiao JL, Zhang Y, Zeng Q, Xiong C, Han LL, Wang JT, Zhang LM (2021) Microbial assembly and association network in watermelon rhizosphere after soil fumigation for Fusarium wilt control. Agric Ecosyst Environ 312:107336. https://doi.org/10.1016/j.agee.2021.107336
Guimera R, Amaral LAN (2005) Functional cartography of complex metabolic networks. Nature 433(7028):895–900. https://doi.org/10.1038/nature03288
Guo LM, He J, Li J, Chen JP, Zhang HM (2019) Chinese wheat mosaic virus: a long-term threat to wheat in China. J Integr Agric 18(4):821–829. https://doi.org/10.1016/S2095-3119(18)62047-7
Guttman DS, McHardy AC, Schulze-Lefert P (2014) Microbial genome-enabled insights into plant–microorganism interactions. Nat Rev Genet 15(12):797–813. https://doi.org/10.1038/nrg3748
Haichar FZ, Marol C, Berge O, Rangel-Castro JI, Prosser JI, Balesdent J, Heulin T, Achouak W (2008) Plant host habitat and root exudates shape soil bacterial community structure. ISME J 2(12):1221–1230. https://doi.org/10.1038/ismej.2008.80
Hamonts K, Trivedi P, Garg A, Janitz C, Grinyer J, Holford P, Botha FC, Anderson IC, Singh BK (2018) Field study reveals core plant microbiota and relative importance of their drivers. Environ Microbiol 20(1):124–140. https://doi.org/10.1111/1462-2920.14031
Hen CG, Li DW, Xing YM, Zhu K, Tian ZF, Cai ZN, Yu JL, Liu Y (2000) Wheat yellow mosaic virus widely occurring in wheat (Triticum aestivum) in China. Plant Dis 84(6):627–630. https://doi.org/10.1094/pdis.2000.84.6.627
Hoitink HAJ, Stone AG, Grebus ME (1996) Suppression of plant diseases by composts. In: de Bertoldi M., Sequi P., Lemmes B., Papi T. (eds) The science of composting. Springer, Dordrecht. pp 373–381. https://doi.org/10.1007/978-94-009-1569-5_35
Hothorn T, Bretz F, Westfall P (2008) Simultaneous inference in general parametric models. Biom J 50(3):346–363. https://doi.org/10.1002/bimj.200810425
Huang XQ, Zhou X, Zhang JB, Cai ZC (2019) Highly connected taxa located in the microbial network are prevalent in the rhizosphere soil of healthy plant. Biol Fertil Soils 55(3):299–312. https://doi.org/10.1007/s00374-019-01350-1
Irikiin Y, Nishiyama M, Otsuka S, Senoo K (2006) Rhizobacterial community-level, sole carbon source utilization pattern affects the delay in the bacterial wilt of tomato grown in rhizobacterial community model system. Appl Soil Ecol 34(1):27–32. https://doi.org/10.1016/j.apsoil.2005.12.003
Jiao S, Chen WM, Wang JL, Du NN, Li QP, Wei GH (2018) Soil microbiomes with distinct assemblies through vertical soil profiles drive the cycling of multiple nutrients in reforested ecosystems. Microbiome 6:13. https://doi.org/10.1186/s40168-018-0526-0
Karasov TL, Almario J, Friedemann C, Ding W, Giolai M, Heavens D, Kersten S, Lundberg DS, Neumann M, Regalado J, Neher RA, Kemen E, Weigel D (2018) Arabidopsis thaliana and Pseudomonas pathogens exhibit stable associations over evolutionary timescales. Cell Host Microbe 24(1):168. https://doi.org/10.1016/j.chom.2018.06.011
Kennedy TA, Naeem S, Howe KM, Knops JMH, Tilman D, Reich P (2002) Biodiversity as a barrier to ecological invasion. Nature 417(6889):636–638. https://doi.org/10.1038/nature00776
Koljalg U, Larsson KH, Abarenkov K, Nilsson RH, Alexander IJ, Eberhardt U, Erland S, Hoiland K, Kjoller R, Larsson E, Pennanen T, Sen R, Taylor AFS, Tedersoo L, Vralstad T, Ursing BM (2005) UNITE: a database providing web-based methods for the molecular identification of ectomycorrhizal fungi. New Phytol 166(3):1063–1068. https://doi.org/10.1111/j.1469-8137.2005.01376.x
Kwak MJ, Kong HG, Choi K, Kwon SK, Song JY, Lee J, Lee PA, Choi SY, Seo M, Lee HJ, Jung EJ, Park H, Roy N, Kim H, Lee MM, Rubin EM, Lee SW (2018) Kim JF (2018) Rhizosphere microbiome structure alters to enable wilt resistance in tomato (vol 36, pg 1100. Nat Biotechnol 36(11):1117–1117. https://doi.org/10.1038/nbt1118-1117
Langfelder P, Horvath S (2012) Fast R functions for robust correlations and hierarchical clustering. J Stat Softw 46(11):1–17
Latz E, Eisenhauer N, Rall BC, Allan E, Roscher C, Scheu S, Jousset A (2012) Plant diversity improves protection against soil-borne pathogens by fostering antagonistic bacterial communities. J Ecol 100(3):597–604. https://doi.org/10.1111/j.1365-2745.2011.01940.x
Lavecchia A, Curci M, Jangid K, Whitman WB, Ricciuti P, Pascazio S, Crecchio C (2015) Microbial 16S gene-based composition of a sorghum cropped rhizosphere soil under different fertilization managements. Biol Fertil Soils 51(6):661–672. https://doi.org/10.1007/s00374-015-1017-0
Lazcano C, Boyd E, Holmes G, Hewavitharana S, Pasulka A, Ivors K (2021) The rhizosphere microbiome plays a role in the resistance to soil-borne pathogens and nutrient uptake of strawberry cultivars under field conditions. Sci Rep 11(1):3188. https://doi.org/10.1038/s41598-021-82768-2
Li P, Cui Z, Gao G, Sun M, Zhang F, Li X (2017) Research progress of wheat yellow mosaic. Shandong Agricultural Sciences(In Chinese with English abstract), 2017, 49(08):168–172. doi: https://doi.org/10.14083/j.issn.1001-4942.2017.08.038
Liaw A, Wiener M (2002) Classification and regression by randomForest. R News 2(3):18–22
Liu W, Jiang L, Yang S, Wang Z, Tian R, Peng Z, Chen Y, Zhang X, Kuang J, Ling N, Wang S, Liu L (2020) Critical transition of soil bacterial diversity and composition triggered by nitrogen enrichment. Ecology 101(8):e03053. https://doi.org/10.1002/ecy.3053
Liu HW, Li JY, Carvalhais LC, Percy CD, Verma JP, Schenk PM, Singh BK (2021) Evidence for the plant recruitment of beneficial microbes to suppress soil-borne pathogens. New Phytol 229(5):2873–2885. https://doi.org/10.1111/nph.17057
Luti KJK, Mavituna F (2011) Elicitation of Streptomyces coelicolor with dead cells of Bacillus subtilis and Staphylococcus aureus in a bioreactor increases production of undecylprodigiosin. Appl Microbiol Biotechnol 90(2):461–466. https://doi.org/10.1007/s00253-010-3032-2
Mallon CA, Elsas JDv, Salles JF, (2015) Microbial invasions: the process, patterns, and mechanisms. Trends Microbiol 23(11):719–729. https://doi.org/10.1016/j.tim.2015.07.013
Mangeot-Peter L, Tschaplinski TJ, Engle NL, Veneault-Fourrey C, Martin F, Deveau A (2020) Impacts of soil microbiome variations on root colonization by fungi and bacteria and on the metabolome of Populus tremula × alba. Phytobiomes J 4(2):142–155. https://doi.org/10.1094/pbiomes-08-19-0042-r
McDonald D, Price MN, Goodrich J, Nawrocki EP, DeSantis TZ, Probst A, Andersen GL, Knight R, Hugenholtz P (2012) An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J 6(3):610–618. https://doi.org/10.1038/ismej.2011.139
Mendes R, Kruijt M, de Bruijn I, Dekkers E, van der Voort M, Schneider JHM, Piceno YM, DeSantis TZ, Andersen GL, Bakker PAHM, Raaijmakers JM (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332(6033):1097–1100. https://doi.org/10.1126/science.1203980
Mendes LW, Mendes R, Raaijmakers JM, Tsai SM (2018a) Breeding for soil-borne pathogen resistance impacts active rhizosphere microbiome of common bean. ISME J 12(12):3038–3042. https://doi.org/10.1038/s41396-018-0234-6
Mendes LW, Raaijmakers JM, de Hollander M, Mendes R, Tsai SM (2018b) Influence of resistance breeding in common bean on rhizosphere microbiome composition and function. ISME J 12(1):212–224. https://doi.org/10.1038/ismej.2017.158
Messiha NAS, van Bruggen AHC, van Diepeningen AD, de Vos OJ, Termorshuizen AJ, Tjou-Tam-Sin NNA, Janse JD (2007) Potato brown rot incidence and severity under different management and amendment regimes in different soil types. Eur J Plant Pathol 119(4):367–381. https://doi.org/10.1007/s10658-007-9167-z
Mitter B, Pfaffenbichler N, Flavell R, Compant S, Antonielli L, Petric A, Berninger T, Naveed M, Sheibani-Tezerji R, von Maltzahn G, Sessitsch A (2017) A new approach to modify plant microbiomes and traits by introducing beneficial bacteria at flowering into progeny seeds. Front Microbiol 8(11). https://doi.org/10.3389/fmicb.2017.00011
Olesen JM, Bascompte J, Dupont YL, Jordano P (2007) The modularity of pollination networks. P Natl Acad Sci USA 104(50):19891–19896. https://doi.org/10.1073/pnas.0706375104
Parks DH, Tyson GW, Hugenholtz P, Beiko RG (2014) STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30(21):3123–3124. https://doi.org/10.1093/bioinformatics/btu494
Perez-Jaramillo JE, Carrion VJ, Bosse M, Ferrao LFV, de Hollander M, Garcia AAF, Ramirez CA, Mendes R, Raaijmakers JM (2017) Linking rhizosphere microbiome composition of wild and domesticated Phaseolus vulgaris to genotypic and root phenotypic traits. ISME J 11(10):2244–2257. https://doi.org/10.1038/ismej.2017.85
Puigagut J, Salvadó H, García J (2005) Short-term harmful effects of ammonia nitrogen on activated sludge microfauna. Water Res 39(18):4397–4404. https://doi.org/10.1016/j.watres.2005.08.008
Ruiz-Perez CA, Restrepo S, Zambrano MM (2016) Microbial and functional diversity within the phyllosphere of Espeletia species in an andean high-mountain ecosystem. Appl Environ Microbiol 82(6):1807–1817. https://doi.org/10.1128/aem.02781-15
Shi Y, Li Y, Xiang X, Sun R, Yang T, He D, Zhang K, Ni Y, Zhu Y-G, Adams JM (2018) Spatial scale affects the relative role of stochasticity versus determinism in soil bacterial communities in wheat fields across the North China Plain. Microbiome 6(1):1–12. https://doi.org/10.1186/s40168-018-0409-4
Smith KP, Goodman RM (1999) Host variation for interactions with beneficial plant-associated microbes. Annu Rev Phytopathol 37:473–491. https://doi.org/10.1146/annurev.phyto.37.1.473
Sun L, Andika IB, Kondo H, Chen J (2013a) Identification of the amino acid residues and domains in the cysteine-rich protein of Chinese wheat mosaic virus that are important for RNA silencing suppression and subcellular localization. Mol Plant Pathol 14(3):265–278. https://doi.org/10.1111/mpp.12002
Sun L, Andika IS, Shen J, Yang D, Ratti C, Chen J (2013b) The CUG-initiated larger form coat protein of Chinese wheat mosaic virus binds to the cysteine-rich RNA silencing suppressor. Virus Res 177(1):66–74. https://doi.org/10.1016/j.virusres.2013.07.013
Tao J, Meng D, Qin C, Liu X, Liang Y, Xiao Y, Liu Z, Gu Y, Li J, Yin H (2018) Integrated network analysis reveals the importance of microbial interactions for maize growth. Appl Microbiol and Biotechnol 102(8):3805–3818. https://doi.org/10.1007/s00253-018-8837-4
Tonelli ML, Figueredo MS, Rodríguez J, Fabra A, Ibañez F (2020) Induced systemic resistance-like responses elicited by rhizobia. Plant Soil 448(1):1–14. https://doi.org/10.1007/s11104-020-04423-5
Trivedi P, Delgado-Baquerizo M, Trivedi C, Hamonts K, Anderson IC, Singh BK (2017) Keystone microbial taxa regulate the invasion of a fungal pathogen in agro-ecosystems. Soil Biol Biochem 111:10–14. https://doi.org/10.1016/j.soilbio.2017.03.013
van Elsas JD, Chiurazzi M, Mallon CA, Elhottovā D, Krištůfek V, Salles JF (2012) Microbial diversity determines the invasion of soil by a bacterial pathogen. Proc Natl Acad Sci 109(4):1159–1164. https://doi.org/10.1073/pnas.1109326109
Walters W, Hyde ER, Berg-Lyons D, Ackermann G, Humphrey G, Parada A, Gilbert JA, Jansson JK, Caporaso JG, Fuhrman JA, Apprill A, Knight R (2016) Improved bacterial 16S rRNA gene (V4 and V4–5) and fungal internal transcribed spacer marker gene primers for microbial community surveys. mSystems 1(1):10. https://doi.org/10.1128/mSystems.00009-15
Wang W, Wang H, Zu Y (2014) Temporal changes in SOM, N, P, K, and their stoichiometric ratios during reforestation in China and interactions with soil depths: importance of deep-layer soil and management implications. For Ecol Manage 325:8–17. https://doi.org/10.1016/j.foreco.2014.03.023
Wei Z, Yang T, Friman V-P, Xu Y, Shen Q, Jousset A (2015) Trophic network architecture of root-associated bacterial communities determines pathogen invasion and plant health. Nat Commun 6(1):8413. https://doi.org/10.1038/ncomms9413
Wen T, Yuan J, He X, Lin Y, Huang Q, Shen QJ (2020) Enrichment of beneficial cucumber rhizosphere microbes mediated by organic acid secretion. Hortic Res 7(1):1–13. https://doi.org/10.1038/s41438-020-00380-3
Wickham H (2011) ggplot2. Wires Comp Stat 3(2):180–185. https://doi.org/10.1002/wics.147
Wickham H (2007) Reshaping data with the reshape package. J Stat Softw 21(12):1–20 https://had.co.nz/reshape
Wintermans PCA, Bakker P, Pieterse CMJ (2016) Natural genetic variation in Arabidopsis for responsiveness to plant growth-promoting rhizobacteria. Plant Mol Biol 90(6):623–634. https://doi.org/10.1007/s11103-016-0442-2
Wu B, Jiang S, Zhang M, Wang S, Zhao J, Xin X (2017) Resistance of wheat cultivars to wheat yellow mosaic virus in shandong. Journal of Triticeae Crops 37(3). (in Chinese with English abstract) 2017,37(03):332–336. https://doi.org/10.7606/j.issn.1009-1041.2017.03.10
Xiong C, Zhu YG, Wang JT, Singh B, Han LL, Shen JP, Li PP, Wang GB, Wu CF, Ge AH, Zhang LM, He JZ (2021) Host selection shapes crop microbiome assembly and network complexity. New Phytol 229(2):1091–1104. https://doi.org/10.1111/nph.16890
Xue Y, Chen H, Yang JR, Liu M, Huang B, Yang J (2018) Distinct patterns and processes of abundant and rare eukaryotic plankton communities following a reservoir cyanobacterial bloom. ISME J 12(9):2263–2277. https://doi.org/10.1038/s41396-018-0159-0
Yu P, Wang C, Baldauf JA, Tai H, Gutjahr C, Hochholdinger F, Li C (2018) Root type and soil phosphate determine the taxonomic landscape of colonizing fungi and the transcriptome of field-grown maize roots. New Phytol 217(3):1240–1253. https://doi.org/10.1111/nph.14893
Zhang Z, Yuan Y, Zhao W, He H, Li D, He W, Liu Q, Yin HJ (2017) Seasonal variations in the soil amino acid pool and flux following the conversion of a natural forest to a pine plantation on the eastern Tibetan Plateau, China. Soil Biol Biochem 105:1–11. https://doi.org/10.1016/j.soilbio.2016.11.002
Funding
This work was supported by the China Agriculture Research System from the Ministry of Agriculture of the P.R. China (CARS-03) and sponsored by the K.C. Wong Magna Fund of Ningbo University, China.
Author information
Authors and Affiliations
Contributions
GTD, YJ and CJP conceived and designed the research. WCF and WFF conducted experiments. ZHQ and DYW contributed analytical tools. WCF and CGX analyzed the data. WCF wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wu, C., Wang, F., Zhang, H. et al. Enrichment of beneficial rhizosphere microbes in Chinese wheat yellow mosaic virus-resistant cultivars. Appl Microbiol Biotechnol 105, 9371–9383 (2021). https://doi.org/10.1007/s00253-021-11666-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11666-4