Abstract
Although great advances have been made on large-scale manufacturing of vaccines and antiviral-based drugs, viruses persist as the major cause of human diseases nowadays. The recent pandemic of coronavirus disease-2019 (COVID-19) mounts a lot of stress on the healthcare sector and the scientific society to search continuously for novel components with antiviral possibility. Herein, we narrated the different tactics of using biopeptides as antiviral molecules that could be used as an interesting alternative to treat COVID-19 patients. The number of peptides with antiviral effects is still low, but such peptides already displayed huge potentials to become pharmaceutically obtainable as antiviral medications. Studies showed that animal venoms, mammals, plant, and artificial sources are the main sources of antiviral peptides, when bioinformatics tools are used. This review spotlights bioactive peptides with antiviral activities against human viruses, especially the coronaviruses such as severe acute respiratory syndrome (SARS) virus, Middle East respiratory syndrome (MERS) virus, and severe acute respiratory syndrome coronavirus 2 (SARS-COV-2 or SARS-nCOV19). We also showed the data about well-recognized peptides that are still under investigations, while presenting the most potent ones that may become medications for clinical use.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
• The mechanisms of limitedly available antiviral peptides show that they are potent.
• The peptides could be derived from animal venoms, mammals, plants, or artificial synthesis.
• They are potent against human coronaviruses-related diseases like SARS, MERS, and SARS-nCoV19.
Introduction
Viruses are found in birds, animals, and humans and are responsible for numerous epidemics or even pandemics. Since the last century, major breakthroughs regarding the control of viral replication, infection, and the production of vaccines have led to remarkable advances in human-virus interactions, such as smallpox eradication and the control of measles and poliomyelitis transmission (Vilas Boas et al. 2019). The same last century has been characterized with 5 pandemic respiratory diseases associated with influenza viruses (H1N1 or Spanish flu, H2N2 or Asian flu, H3N2 or Hong Kong flu, H5N1 or Bird flu and H1N1 Swine flu) with pigs serving as the natural hosts, infecting humans, pigs, and birds (Boopathi et al., 2020). The most recent pandemics are from the coronavirus family, which caused severe acute respiratory syndrome (SARS), Middle East respiratory syndrome (MERS), and novel coronavirus (SARS-nCoV19) in this present century.
The present respiratory disease (SARS-nCoV19 or COVID-19) is caused by novel coronaviruses. The disease has taken over the world with symptoms like cold, fever, cough, sore throat, runny nose, and breathing difficulty. For reasons like this, the most utilized treatment for viral control is the use of antiviral drugs (Lou et al. 2014). Due to the side effects of certain antiviral drugs that have developed over the past decades, advances into newer and safer antiviral molecules are being made. Isolation of new compounds and peptides from diverse sources with the aid of molecular, structural, and bioinformatic analyses has contributed to the prospects of antiviral peptides discovery for therapeutic purposes against various human viruses (Wang et al. 2010; Rothan et al. 2014).
Therefore, this present pandemic situation versus available information on antiviral peptides putative of antiviral therapy or drugs are instigators of the present study. The therapeutic potentials of antiviral bioactive peptides against the human viruses, especially the severe respiratory syndrome coronaviruses, were examined. This review comprehensively summarizes the occurrence of antiviral peptides, mechanisms of peptide inhibition and viral action, along with futuristic needs and recommendations. Most importantly, we systematically discussed the antiviral peptides against SARS, MERS, and SARS-nCoV19. We do believe that this kind of ordered knowledge will guide researchers to extend their studies and explore the promising peptides with antiviral effects.
To do that, we used “antiviral peptides,” “bioactive peptides,” “peptides sources,” “coronavirus,” and “COVID-19” as research terms, applied Boolean operators “AND/OR” combination, performed in April 2020 from the following electronic databases: Scopus, PubMed, ScienceDirect, Wiley Online, Springer, Web of Science, ACS pubs, RCS-website, and Google Scholar. We initially checked the content of the English published articles to detect both included or excluded criteria, and therefore, some irrelevant articles were omitted according to the eligibility criteria published by Peterson et al. (2006). Then, we narrated the extractable data in the following sections.
Occurrence of antiviral peptides
Production of bioactive peptides
Bioactive peptides are short fragments of food proteins capable of providing positive influence and health-promoting properties when consumed by humans. They act as nutrients that potentate physiological functions and disease prevention when passed through the absorptive pathways in the digestive system (Ashaolu 2020a). Based on the bioactive peptide sequence, their functions may include satiety and antiobesity (gastrointestinal system), antithrombotic, antihypertensive, antioxidant, hypocholesterolemic (cardiovascular system), opioid (nervous system), cytomodulatory, immunomodulatory, antiviral, and antimicrobial effects (immune system) (Vilas Boas et al. 2019; Mora et al. 2019; Ashaolu 2020a, 2020b and Ashaolu 2020c).
The production of bioactive peptides may involve classical or bioinformatic (in silico) approaches (Daliri et al. 2017). The classical way includes enzymatic hydrolysis or microbial fermentation, both of which involve using enzymes or microorganisms to hydrolyze proteins into a pool of hydrolysates. This will be followed by bioassay purification, peptide size separation via membrane filtration, high-performance liquid chromatography (HPLC), and in vitro and in vivo bioactivity tests (Ashaolu 2020a). There would also be a further peptide purification and sequencing step to isolate the desired bioactive peptide for the final in vitro and in vivo biological tests. The in silico method employs database sets composing of already isolated and identified peptide sequences to match the target protein. Based on the information, the identified sequences in the target protein are hydrolyzed by specific enzymes associated with their cleavage sites. It enhances the identification of known peptides from unknown proteins (Daliri et al. 2017). Proteins from microorganisms, fungi, plants (soy, wheat, barley, rye, oats, rice, corn, sorghum, and millet), and animals (meat, eggs, milk, and fish) have been recognized as main sources of bioactive peptides (Mora et al. 2015; Ashaolu 2020a). Almost half of the past century has been invested into peptides research with numerous angles to their bioactivities. Till now, characterizing the bioactive peptides has more often favored antibacterial or antimicrobial functions, of which the cationic and amphipathic properties of the antimicrobial peptides are not dissimilar to antiviral peptides (Vilas Boas et al. 2019).
Antiviral peptides
Antiviral peptides have several commonalities with antimicrobial peptides targeted at all microorganisms but need special attention since their targets are between living and non-living organisms; they are replicative, non-metabolizing, and lack self-generated energy (Pärn et al. 2015). As there are numerous examples of antiviral peptides, which are predominantly cell-based, in vivo protective studies are lacking (Shartouny and Jacob 2019). The demerits of antiviral peptides usage include their expensiveness, short half-life, limited upper gastrointestinal absorption, and poor immunogenicity, while on the other hand, they can be very effective, less toxic, enzymatically biodegradable, and specific (Galdiero et al. 2013). Peptides with antiviral activity against human viruses are listed in Table 1.
Animal-derived antiviral peptides
The antiviral peptides may be obtained from animal or plant sources. The animal-derived antiviral peptides have been scantily sourced from marine organisms, amphibians, mammals, and arthropods. The commonest peptides from the marines are the clavanins from Styela clava and mirabamides E, F, G, and H obtained from Stelletta clavosa, which demonstrated antiviral activities against rotavirus, adenovirus, and human immune deficiency virus (HIV) (Yasin et al. 2000; Lu et al. 2011; Ireland et al. 2008). An antiviral peptide nomenclated as Pa-MAP 1 and capable of interference with the herpes simplex virus (HSV-1 and HSV-2) glycoprotein surface was derived from Pleuronectes americanus, an arctic fish (Migliolo et al. 2012; Vilas Boas et al. 2017). Didemnim (A and B) have also shown virucidal actions against parainfluenza, dengue, and human papillomavirus (HPV) (Canonico et al. 1982). Didemnins-like depsipeptides derived from tunicates such as Didemnins X, Y, M, Nordidemnin N, and Epididemnin A were also found to possess antiviral properties (Aneiros and Garateix 2004).
The physiological and morphological characteristics of amphibians suggest that they can produce antimicrobial and antiviral peptides. Rana temporaria is a frog species capable of producing temporin B peptide against HSV-1 and was found to rupture the viral envelope (Marcocci et al. 2018). Magainin 1 and 2 peptides from Xenopus laevis demonstrated virucidal action against vaccinia virus (VV), HSV-1 and HSV-2, with clear indications that their lysine-rich and alanine-rich regions stimulated certain unclear mechanisms that associated their cationic charge with an amphipathic structure to cause interactions with the anionic phospholipids of the viral envelope, consequently, causing their disruption and death (Egal et al. 1999; Matanic and Castilla 2004; Dean et al. 2010). Obviously, the dermal glands of frogs have proven to be the source of many amphibian-derived peptides because they can release cationic, amphipathic α-helical secondary structure, with 10–50 amino acids when stressed (Marcocci et al. 2018; Vilas Boas et al. 2019).
Other than HSV, peptides targeting HIV-1 envelope were also derived from the genus Phyllomedusa, the dermaseptins (Belaid et al. 2002; Lorin et al. 2005; Bergaoui et al. 2013). The derivatives (S3, S4) of dermaseptins showed virucidal activity against rabies virus when amino acid exchange (methionine-lysine) was involved, using bioassays and mice models (Mechlia et al. 2019). Hydrophylax bahuvistara frog can produce urumin, which demonstrated inhibitory and destructive actions against influenza (H1N1, H1N2) virus when bioassays and mice models were employed (Holthausen et al. 2017). This apparently indicated that hemagglutinin 1 was switched into a reactive mode with the peptide urumin.
In addition, dengue virus (2 and 3) could be attacked by HS-1 peptide isolated from the skin of Hypsiboas semilineatus. This is based on the studies of Monteiro et al. (2018) who observed HS-1 peptide activity at the early stages of dengue viral infection in vitro, subsequent reduction in viral load, and the disruptive impact of the peptide on the viruses that caused some envelopes to invaginate.
Apart from lactoferrin or protegrin-1, the antiviral peptides that mammals commonly produce are defensins and cathelicidins, which are also antimicrobials with cationic and amphipathic characteristics. Cathelicidins were demonstrated to inhibit or destroy influenza A virus, HSV, HIV, respiratory syncytial virus (RSV), varicella zoster virus (VZV), VV, hepatitis C virus (HCV), Zika virus, adenovirus, rhinovirus, Aichi virus, and certain serovars of dengue virus with no same mechanisms of action, which often involve viral envelope and components damage or host cell protection (Gordon et al. 2005; Barlow et al. 2011; Matsumura et al. 2016; Sousa et al. 2017; Alagarasu et al. 2017; He et al. 2018; Ahmed et al. 2019).
Defensins have shown activity against HIV, influenza A virus, VV, and varicella zoster virus among others (Quiñones-Mateu et al. 2003; Salvatore et al. 2007; Howell et al. 2007; Crack et al. 2012; Zapata et al. 2016), indicative of being used as futuristic antiretroviral peptides. Protegrin-1 originating from pigs is a cyclic peptide shown to inhibit dengue virus replication via the replication enzyme pathway, while its mirror structure demonstrated activity against HSV-1 and HSV-2 (Yasin et al. 2000).
Lactoferrin and its derivative, lactoferricin, are peptides obtained from the milk of mammals and have antiviral multiactivity against influenza virus, HPV, HIV, RSV, adenovirus, poliovirus, rotavirus, hepatitis B virus (HBV), HCV, Zika virus, dengue virus cytomegalovirus (CMV), and HSV (Andersen et al. 2001; Van der Strate et al. 2001; Mistry et al. 2007; Marr et al. 2009; Li et al. 2009; Chen et al. 2017; Shestakov et al. 2012; Wang et al. 2016).
Insects, crustaceans, arachnids, and every other member of the arthropodal community are not left out of the supplies of antiviral peptides. Calliphoridae member blowfly peptides and alloferons (1 and 2) have been shown to inhibit influenza virus serovars, coxsackievirus B2 (CBV-2), and HSV-1 (Chernysh et al. 2002; Kuczer et al. 2011, 2013), with less vivid action mechanism. Cecropia moth can produce cecropin A peptide, which has activity against HSV, Junin virus (JV), and HIV (Wachinger et al. 1998; Albiol Matanic and Castilla 2004; Hultmark et al. 2005).
Apart from studies on morphological parts of arthropods, their venoms have widely been investigated by researchers for potential antiviral peptides. Asian spider venom peptide, Lactarcin 1 (from Lachesana tarabaeve) possesses antiviral property against dengue virus-2 through protease activity blockage (Rothan et al. 2014). Melittin, which is derived from bees’ venom, as well as melittin-incorporated nanoparticulates showed activity against HSV, JV, and HIV via lysis of their envelopes (Albiol Matanic and Castilla 2004; Galdiero et al. 2013; Hood et al. 2013). Similarly, wasp venom contains mastoparan capable of rupturing and separating the envelope of vesicular stomatitis virus (VSV) from its capsid (Sample et al. 2013). Mucroporin-M1 peptide from aquatic scorpion (Lychas mucronatus) had inhibitory effects against severe acute respiratory syndrome coronavirus (SARS-CoV), influenza H5N1, measles virus, and HBV (Li et al. 2011; Zhao et al. 2012) while Euscorpiops validus venom-derived peptide, Eva 1418, inhibited the replication of HSV-1 (Zeng et al. 2018). Three peptides derived from scorpion venom, namely Hp1090, Hp1239, and Hp1036, were also demonstrated to act against HSV-1 and HCV replication (Hong et al. 2014; Yan et al. 2011).
Plant-derived antiviral peptides
As a means of self-defense, plants produce multifarious poisonous substances against pathogens and pests. These substances include molecules, proteins, peptides, and toxins that fight against impending viral attacks. This backdrop has led to numerous investigations into these plant defense substances, especially peptides as putative human antiviral peptides (Shartouny and Jacob 2019). Among this group of plant defense peptides, cyclotides have gained vast attention due to their rigid structure and numerous biological activities such as molluscicidal, insecticidal, nematicidal, antimicrobial, and anthelmintic (Weidmann and Craik 2016; Vilas Boas et al. 2019). They have also shown inhibitory actions against dengue virus, HIV, and influenza H1N1 virus (Henriques and Craik 2010; Sencanski et al. 2015). Other related peptides including 2000 Da fractionated peptide, sesquin, and phaseococcin, isolated from Sorghum bicolor seeds, Vigna sesquipedalis, and Phaseolus coccineus, respectively, have demonstrated inhibitory activities against HSV-1 and HIV (Camargo Filho et al. 2008; Jack and Tzi 2005; Ngai and Ng 2005). Moreover, antiviral peptides from pokeweed (PAP-1, PAP-2, and PAP-3) were reported to cause genomic mutations of poliovirus, HSV, HIV-1, influenza virus and lymphocytic choriomeningitis mammarenavirus (LCMV) via depurination (Rajamohan et al. 1999; Uckun et al. 2003; Aron and Irvin 1980; Tomlinson et al. 1974; Uckun et al. 2005), as well as inhibit Japanese encephalitis virus both in vitro and in vivo (Ishag et al. 2013; Domashevskiy and Goss 2015).
Mechanism of inhibition
Peptides with antiviral or virucidal activities integrate into the envelope of the target virus or cell membrane of the host to create an unstable membrane. Some peptides prevent viral spike proteins from binding to host cells (Belaid et al. 2002). The cellular pathways including DNA replication and protein synthesis may be altered in this process, thereby stopping viral infection (Sitaram and Nagaraj 1999; Pärn et al. 2015). Since many antiviral peptides can be obtained from previously demonstrated antimicrobial molecules (Chinchar et al. 2004; Crack et al. 2012), it is logical to relate the idea that peptides with positive charges can have direct interaction with negatively charged host cellular membranes to cause viral inhibition.
From the initial to the final phase of a typical viral cycle, most virucidal peptides inhibit virions by direct contact, or suppress their gene expression, or interfere with the linkages, movements, and adsorption of proteins in the host cell membrane (Galdiero et al. 2013; Zapata et al. 2016). An example is the Tat antiviral peptide, which interacts with CXCR4, a co-receptor (protein) of HIV-1, and inhibits the replication of HIV-1 (Keogan et al. 2012; Pärn et al. 2015). The mechanism of inhibition of antiviral peptides is described within the viral replication cycle in Fig. 1. The antiviral activity of any peptide will be affected by its chemical and structural properties including charge, amino acid sequence, and profile while bioinformatics and in silico designs have paved the edge-cutting way for more modern peptide analysis (Mooney et al. 2012; Sharma et al. 2014).
Annotated action mechanism of antiviral peptides based on inhibition sites within a viral replication cycle adapted from different studies. Adapted from Vilas Boas et al. (2019)
Coronavirus family
An overview about coronavirus
Coronaviruses (CoVs) are enveloped, positive-sense, single-stranded RNA viruses with genomic lengths of ≤32 kb and can cause liver, intestinal, respiratory, and nervous system diseases in mammals such as animals and humans (Monajjemi et al. 2020; Liya et al. 2020). Their genomes encode between four and five structural proteins, including hemagglutinin esterase, nucleocapsid, membrane, envelope, and spike proteins (Esper et al. 2005). Spike protein is the most important surface membrane protein of coronavirus, which determines the host range and specificity of the virus (Liya et al. 2020). The coronavirus family belongs to the order Nidovirales, and it can be categorized into 3 chief groups: ɑ-coronavirus (group 1), ß-coronavirus (group 2), and γ-coronavirus (group 3), while the ß-coronavirus is subsequently sub-classified into group 2A and 2B (Chuck et al. 2011). The ɑ- and ß-genera are responsible for human CoVs infections, the HCoVs. ɑ-CoVs include HCoV-229E and HCoVNL63 while ß-CoVs include HCoV-HKU1, HCoV-OC43, SARS-COV, MERS-CoV, and SARS-nCoV19 (Rabaan et al. 2020).
SARS, MERS, and SARS-nCoV19
SARS-CoV, MERS-CoV, and SARS-nCoV19 cause a variety of severe flu-like symptoms and were responsible for recent epidemics (in 2002/3, 2012/5, and 2019/2020, respectively) (Bauchner et al. 2020). Despite of this classification, the ability of CoVs to generate novel strains with high virulence through high frequency of mutations and recombination remains a potential threat to human health.
SARS-CoV is a newly identified member of Coronaviridae family identified in 2002/2003. Coronavirus envelope S-protein is a class I viral fusion protein which is categorized by the presence of two heptad repeat regions (HR1 and HR2), making a multifaceted complex called fusion central. MERS-CoV initially came into the spotlight in 2012, and dispersed worldwide with a high death rate, and now, the world is underneath another severe COVID-19 pandemic instigated by SARS-nCoV19 or novel coronavirus (nCoV-19) (Lai et al. 2020; Prompetchara et al. 2020). So far, more than 120 million people have been infected with close to 3 million deaths recorded across the world (www.worldometers.info/coronavirus). SARS-nCoV19 is vastly spreadable in humans, and so far, no medication has been industrialized to treat the virus infection (Shereen et al. 2020). While drugs such as hydroxychloroquine, remdesivir, and lopinavir (Rosa and Santos 2020) are presently being proposed to hinder SARS-nCoV19 infection, there is no clinical study so far to verify their efficiency in treating these patients. Thus, presently, there is a worldwide search for suitable medication for SARS-nCoV19.
In general, SARS-nCoV19 has exposed 80% genome uniqueness with SARS-CoV. It was found that the viruses could be contracted from their normal hosts (bats) based on their 96.2% genomic similarity (Zhou et al. 2020) through intermediates (civet cats, camels). This is laudable, but information on the number of hosts during transmission process and how humans contract them via animals has not been substantiated. While some reports suggest direct contact with raw meat or milk of intermediate hosts as means of getting HCoV infections, SARS-CoV and MERS-CoV transmission routes were identified as mainly via infectious droplets and close contact (NHFPC 2015). It has also been reported that COVID-19 thrives in densely concentrated areas and can be transmitted via aerosols to the general population (Liya et al. 2020).
Beyond the mechanism of the antiviral drugs, the viral particle is molded by S-glycoprotein, envelope (E), membrane (M), and nucleocapsid (N) (Boopathi et al. 2020). The spike is a type 1 transmembrane protein and is shaped by two subunits (S1 and S2) which are elaborate in the fusion/entry procedure. While S1 has receptor-binding domain (RBD) and is accountable for assembling to the cellular receptor, S2 has fusion-peptide, HR1, HR2, transmembrane-domain, and a cytoplasmic domain-peptide and turns in the viral synthesis before being passed into the cell (De Haan et al. 2006). Both HR1 and HR2 have 3 pieces each that, altogether, form a six-helix bundle synthesis central. Disturbance of this central creation is critical to viral union reserve, thus stopping infection (Steffen and Pohlmann 2010). Meanwhile, HR1 and HR2 of S-protein of SARS-CoV are vastly preserved areas and are attractive boards of passive inhibitors.
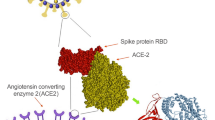
On the other hand, assembling CoVs-S-protein to cellular angiotensin-converting enzyme 2 (ACE2) is the first step in CoV infection. The receptor for SARS-CoV is ACE2, which is a surface molecule located on arteriovenous endothelial cells, arterial smooth muscle cells, small intestinal epithelium, and the respiratory tract (Ren et al. 2008; Liya et al. 2020). Investigation of SARS pathogenesis in different parts and organs of humans has shown that ACE2 is expressed on the surfaces of alveolar epithelial, small intestinal epithelial cells, endothelial cells, and smooth muscle cells (Hamming et al. 2004). Thus, in the bronchial epithelial cells, SARS-nCoV19 may be able to use alveolar type II epithelial cell ACE2 as a receptor for penetration. Speculation arises that there may be similarities in the clinical characteristics of SARS-CoV and SARS-nCoV19 as the amino acid sequence and predicted protein structure between their RBD in the spike protein are highly similar. In fact, the affinity of SARS-nCoV19 to ACE2 binding is higher than SARS-CoV (Wrapp et al. 2020).
Two possible mechanistic pathways have been postulated at this stage. First, the virus particle uses its S1 subunit domain to bind to ACE2 receptor on the human host cell membranes such as alveolar epithelial cells, followed by fusion with S2 subunit at a low pH (Li 2016). Otherwise, the virions undergo membrane fusion after fusion peptide is released from the S2 subunit site and then eat the virus via endocytosis. Here, the virus may exercise receptor-binding reorientation in S-protein, cathepsin L cleavage by host enzymes, and completion of other endosomic fusion mechanisms (Simmons et al. 2005). Then comes the viral release and uncoating of its genetic materials through the proteasomes into the cytoplasm. This process is mediated by the nsp replication-transcription complex (RTC), encoded in the genome and important for double-membrane structural inducement in susceptible cell cytoplasms. The synthesis of M, S, and E proteins ensues and is inserted into the endoplasmic reticulum (ER), which become transferred to endoplasmic reticulum-Golgi intermediate compartment (ERGIC) (Boopathi et al. 2020). After the formation of nucleocapsids by N-protein, new viruses migrate from the cytoplasm toward the cell membrane with vesicular walls to be released in the exocytotic process that leads to host cell death, hence continuing the replication cycle in other susceptible living cells.
The single-stranded RNA genome of CoVs encrypts two polyproteins, which are treated into at least 15 mature proteins by papain-like protease and 3C-like protease (3CLpro) (Lu et al. 1998). The 3CLpro of CoVs is therefore a promising target for development of drugs against CoVs infection due to the inhibition of the proteolytic dispensation, which can diminish viral repetition and viral-made cytotoxic effect (Yao et al. 2018). Antiviral peptides are preferred to conservative drugs and may also be effectual against SARS-nCoV19, due to the continual outbreaks of COVID-19. Thus, it is necessary to enlarge a wide-spectrum drug that can battle against all types of CoVs infection.
Possible antiviral peptides against SARS, MERS, and SARS-nCoV19
The emergence of COVID-19 outbreak needs imperative solutions for operative antiviral tactics. Using the computational chemistry could help in this regard (Zhu et al. 2018). One of these strategies is to inhibit the viral infections using HR-based peptides, which could compete with endogenous HR counterparts and thereby prevent the virus replication and infection. This strategy was successfully used to mitigate the replication of human immunodeficiency (Kliger et al. 2001; Wild et al. 1993), Ebola (Watanabe et al. 2000), and SARS-CoV viruses (Bosch et al. 2004; Zhu et al. 2004). Generally, it is supposed that a sequence of conformational fluctuations happens in the coronavirus HR regions during the viral fusion progression.
Concerning SARS-CoV, Bosch et al. (2004) found that membrane-proximal HR2-peptide of S-protein inhabited SARS-CoV in Vero cells, where the membrane-distal HR1-peptide showed no inhibition activity. Its effectiveness was, however, significantly (P˂0.05) inferior to those of corresponding HR2-peptides of the murine coronavirus and hepatitis virus (MHV) during the inhibition of MHV infection. Interestingly, SARS-CoV HR1- and HR2-peptide mixtures showed a better stability than the individual peptide that was validated using heat-accelerated, biochemical, and electron microscopical analyses, thus providing an appealing basis for developing a therapeutic medication against SARS-CoV. The octapeptide AVLQSGFR was manufactured and tested toward SARS-CoV (BJ-01) (Gan et al. 2006). The results implied that, compared with other components testified, AVLQSGFR is the most potent peptide capable of inhibiting SARS-replication, and that no measurable toxicity is detected in Vero cells. In another study, Ho et al. (2006) evaluated the inhibitory effects of small peptides resultant from S-protein on the interaction of S-protein to ACE2 and on S-protein-pseudotyped retrovirus infection. SP-4 (residues of 192-203), SP-8 (residues of 483-494), and SP-10 (residues of 668-679) pointedly jammed the binding between S-protein and ACE2, with IC50 values of 4.30, 6.99, and 1.88 nM, separately. It was also suggested that SP-10 may be industrialized as an anti-SARS-CoV drug, due to its blocking effects on S-protein which was tested in Vero E6 cells. The HR2 peptide, especially P6 of 23-mer, was found to block the SARS-CoV-fusion with an IC50-value of 1.04 μM (Liu et al. 2009). Most importantly, combination of HR, N46, and its mutated type like N46eg, display synergistic effect with an IC50-value of 1.39 μM and amalgamation index of 0.75, signifying a communal tactic to accomplish hopeful inhibition by HR1-peptide for SARS-CoV. The HR2, either as artificial peptides or as GST-fusion polypeptides, is a powerful inhibitor of SARS-CoV and has been reported to bind with HR1 to create a normal 6-helix bundle (Zhu et al. 2004). Alternatively, Chuck et al. (2013) created 4 nitrile-based peptidomimetic inhibitors with diverse N-terminal defensive clusters and diverse peptide span and scrutinized there in vitro antienzymatic activity of 3CLpro of SARS-CoV. They found out that the nitrile-based peptidomimetic inhibitors are applicable by blocking 3CLpro from a wide series of SARS-CoV with an IC50 value of 4.6-49 μM. This activity was ensured by making a covalent bond with the catalytic Cys145, where AVLQ-peptide creates a sum of auspicious bonds with the S1-S4 substrate-binding pouches. Cbz-AVLQ-CN, which is a peptidomimetic inhibitor, has a wide-spectrum activity on 3CLpro of human-COV strains 229E, NL63, OC43, HKU1, and transferable bronchitis virus, but no noticeable embarrassment on caspase-3. In general, numerous peptides, as anti-SARS-CoV, resulting from HR2 have been recognized, such as P1 (Lu et al. 2013), HR2P (Chambers and Easton 1990), and CP-1 (Otwinowski 1997), which avert fusion core formation by reasonably binding with HR1 and blocking the innate bond betwixt HR1 and HR2.
Regarding MERS-CoV, which is another high-mortality risk kind of coronavirus family, precise, extremely effective therapeutics and vaccines are instantly needed to protect human lives and control the pandemic fears. The antiviral peptides pointing to the membrane fusion architecture signify a favorable type, which had activities towards some viruses, including HIV, SARS-CoV, and MERS-CoV. The HR-based peptide-inhibitor T20 (enfuvirtide) had been used to treat HIV-patients (Cuzin and Alvarez 2003; Imai et al. 2000; Kilby et al. 1998). In a comparison study, the peptide corresponding to the full length of HR2-sequence potently inhibited pseudotyped MERS-CoV with an EC50-value of ~ 3.013 μM, where a similar peptide (HR1) showed no effect even at the supreme concentration (Gao et al. 2013). The changes among similar variance of different peptides were also found in other HR2-peptides of class-I enveloped viruses, but the detailed mechanism explaining these differences has not yet been explored. In a related assay, Gao et al. (2013) characterized the six-helix bundle fusion central construction of MERS-CoV S2-subunit protein and studied the binding of two peptides with each other to form 6-HB-core of MERS-CoV via X-ray crystallography and biophysical investigations. They found that the spanning residues of HR1 (998-1039) and HR2 (1251-1286) peptides formed a stable six-helix bundle fusion central structure, signifying that MERS-CoV goes into the host cell mostly via membrane fusion mechanism. The hydrophilic residues of HR2 (1251-1286) peptide significantly (P˂0.05) improved its stability, solubility, and most importantly enhanced its antiviral activity against MERS-CoV duplication and its S-protein mediated cell-cell fusion, hindering viral fusion central formation, as well as interacting with the viral HR1-domain in a dose-dependent means. Lastly, the authors recommended that HR2 (1251-1286) peptide should be used as a template for design of analogs with enhanced activity against MERS-CoV infection and possibly used in clinical patients. Another study of Sun et al. (2017) inspired by the same fusion core proteins, designed a peptide named MERS-5HB, which contains 3 replicas of HR1 and two copies of HR2. He found that MERS-5HB could bind with HR2P, a peptide resulting from HR2, with a sturdy affinity value of up to 0.24 nM, thereby inhibiting pseudotyped MERS-CoV, pass efficiently with IC50 of ~ 1 μM. In addition, MERS-5HB significantly inhibited S-glycoprotein-mediated syncytial formation in a dose-dependent means (P˂0.05). Further biophysical description displayed that MERS-5HB was a thermo-steady α-helical secondary structure. Finally, the author recommended that MERS-5HB-peptide may offer a decent basis for empathy of an original inhibitor on MERS-CoV. Newly, the same group designed, synthesized, and characterized peptides for HR1 and HR2 regions from the HCoV-229E and tested them against HCoV-229E S-protein-mediated cell-cell fusion, and both pseudotyped and live HCoV-229E. Both 229E-HR1P and 229E-HR2P peptides showed inhibition of cell-cell spread with IC50 of 5.7 and 0.3 μM, respectively, and inhibition of the pseudovirus poison with IC50 of 0.5 and 1.7 μM, respectively. Besides, in mouse-model assays, 229E-HR2P retained its antiviral effects in both higher and inferior breathing areas when intranasally managed. Eventually, the authors proposed that 229E-HR2P could be utilized as an antiviral medication to be utilized lengthwise with diverse antiviral fragments with a diverse underlying mechanism, probably employing synergistic effects (Xia et al. 2018). On the other hand, Ying et al. (2014) identified m336, m337, and m338, as human monoclonal antibodies, directing the receptor (CD26/DPP4), and assembling domain (RBD) of MERS-CoV S-glycoprotein. The extraordinarily utmost nullification effect of such antibodies, especially m336, proposes that they have an inordinate potential for prophylaxis and therapy of MERS-CoV contagion in humans and as a tool for developing vaccine immunogens.
Newly, no known medicines or vaccines to control the COVID-19 pandemic caused by SARS-nCoV19 were found. Interestingly, Barh et al. (2020) disclosed that 3 amino acid in hACE2 bind with nCoV-RBD. AC20 and AC23 derived from hACE2 blocked two key critical positions, where DBP6, which was recognized from databases, blocked 3 sites of nCoV-RBD and assembled with one critical position Gln498. Likewise, it was found that 7 chimeric peptides were measured as promising among which cnCoVP-3, cnCoVP-4, and cnCoVP-7 are the top three, and cnCoVP-4 fits all the criteria as anti-COVID-19 peptide. Generally, the effective peptides ought to bind with 3 key sites of nCoV-RBD, namely Gly485/Phe486/Asn487, Gln493, or Gln498/Thr500/Asn501, or Phe486, Gln493, and Asn501. These are motivating issues that are worth searching in further studies. Small molecules to block 6-helix bundle formation ought to be followed in further studies on the current fusion central structure. As the EC50 is in the micromolar range, for any ideal peptide inhibitors, a thorough screening for peptides of better efficacy should be pursued in the future.
On the other hand, phytochemicals such as anthocyanins and tannins were previously reported to have possible inhibition against SARS-CoV-2. For example, Khalifa et al. (2020a) found that pedunculagin, tercatain, and castalin which are hydrolysable tannins interacted with Cys145 and His41 of SARS-CoV-2-3CLpro. Likewise, Khalifa et al. (2020b) revealed that the polyacylated anthocyanins, including phacelianin, gentiodelphin, cyanodelphin, and tecophilin, were found to authentically bind with the receptor binding site and catalytic dyad (Cys145 and His41) of 2019-nCoV-3CLpro and could be used as effective anti-COVID-19 natural components. It was also shown that other polyphenols could be believed as promising biologically active substances for the inhibition of COVID-19, mostly due to their antiviral activities and the immune-regulation functions. It was also found that triterpenoid, anthraquinone, flavonoids, and other polyphenols are potential keys to cunning antiviral therapies for inhibiting SARS-CoV-2-pro (Mehany et al. 2021). Additionally, vitamin A, vitamin D, and/or minerals, especially Zn, with their derivatives could provide a promising alternative for CoV therapy (Alpert 2017). These functional compounds are essential substances at the molecular and cellular levels that play critical functions (Khalifa et al. 2020c; Khalifa et al. 2019). The advantage of using mixtures containing polyphenols, bioactive peptides, vitamins, and minerals could be furtherly considered against the SARS-CoV-2 infections.
Further needs
General safety issues associated with all bioactive peptides are not strange to antiviral peptides. It is pertinent to investigate the immunogenicity and toxicities of peptides due to intestinal wall disruption, erythrocytes and lymphocytes toxicity, free radical production, enzymopathic, and immunopathic tissue damage and cytotoxicity (Bhandari et al. 2020). The quality of delivery and treatment processes should be critically guarded, be it by conjugating with nanocarriers, antibodies, carbohydrates, or lipids. Other suggestions on the use of antiviral peptides as adjuvants or combined therapeutic agents (Vilas Boas et al. 2019) cannot be oversighted.
Other needs to be met include reproducibility of antiviral peptides and the design of kinetic model peptide synthesis to keep their bioactivity intact (Daliri et al. 2017). Moreover, several peptides have shown antiviral properties, but their sequencing information are still lacking. Kaspar and Reichert (2013) previously suggested stabilizing peptides in vivo with a probe tail structure or peptide sequence clipping for optimal bioavailability. This is due to the ever-challenging issue of poor oral absorption and short half-life of biopeptides inspired by gut digestive enzymes. Post-translational modifications like acetylation and amidation, membrane permeability improvement by adding fatty acid chains, or D-enantiomers usage for decreased target sensor and binding affinity to proteases (Shartouny and Jacob 2019) can help to solve this need for good oral absorption and peptide stability.
Using the available new technologies, antiviral peptides should be enriched for a complete set of in vitro, in vivo, and human-based studies. Proteases of genetically improved microbial strains could be useful in the industrial production of peptides meant for antiviral therapy. It is also important that validated all-round characterization of antiviral peptides is essential to commence therapeutic claims against HCoVs and all other human viral diseases.
References
Abdul F, Ndeboko B, Buronfosse T, Zoulim F, Kann M, Nielsen PE, Cova L (2012) Potent inhibition of late stages of hepadnavirus replication by a modified cell penetrating peptide. PLoS One 7(11):e48721
Ahmed A, Siman-Tov G, Keck F, Kortchak S, Bakovic A, Risner K, Lu TK, Bhalla N, de la Fuente-Nunez C, Narayanan A (2019) Human cathelicidin peptide LL-37 as a therapeutic antiviral targeting Venezuelan equine encephalitis virus infections. Antivir Res 164:61–69
Alagarasu K, Patil PS, Shil P, Seervi M, Kakade MB, Tillu H, Salunke A (2017) In-vitro effect of human cathelicidin antimicrobial peptide LL-37 on dengue virus type 2. Peptides 92:23–30
Albiol Matanic VC, Castilla V (2004) Antiviral activity of antimicrobial cationic peptides against Junin virus and herpes simplex virus. Int J Antimicrob Agents 23:382–389
Alpert PT (2017) The role of vitamins and minerals on the immune system. Home Health Care Manag Pract 29(3):199–202
Andersen JH, Osbakk SA, Vorland LH, Traavik T, Gutteberg TJ (2001) Lactoferrin and cyclic lactoferricin inhibit the entry of human cytomegalovirus into human fibroblasts. Antivir Res 51(2):141–149
Aneiros A, Garateix A (2004) Diss. Abst. Int. Pt. B: Sci. Eng. Diss. Abst. Int. Pt. B: Sci. Eng. 59, 2200, 1998. J Chrom B: Analyt Technol Biomed Life Sci 803(1):41–53
Aron GM, Irvin JD (1980) Inhibition of herpes simplex virus multiplication by the pokeweed antiviral protein. Antimicrob Agents Chemother 17:1032–1033
Ashaolu TJ (2020a) Antioxidative peptides derived from plants for human nutrition: their production, mechanisms and applications. Eur Food Res Technol 246:853–865
Ashaolu TJ (2020b) Applications of soy protein hydrolysates in the emerging functional foods: a review. Int J Food Sci Technol 55(2):421–428
Ashaolu TJ (2020c) Health applications of soy protein hydrolysates. Int J Pept Res Therapeutics 26:2333–2343
Barh D, Tiwari S, Andrade BS, Giovanetti M, Kumavath R, Ghosh P, Góes-Neto A, Alcantara LC, Azevedo V (2020) Potential chimeric peptides to block the SARS-CoV-2 Spike RBD. Harvard Dataverse. Preprints. https://doi.org/10.20944/preprints202004.0347.v1
Barlow PG, Svoboda P, Mackellar A, Nash AA, York IA, Pohl J, Davidson DJ, Donis RO (2011) Antiviral activity and increased host defense against influenza infection elicited by the human cathelicidin LL-37. PLoS One 6(10):e25333
Bauchner H, Fontanarosa PB, Livingston EH (2020) Conserving supply of personal protective equipment—a call for ideas. Jama 323:1911. https://doi.org/10.1001/jama.2020.4770
Belaid A, Aouni M, Khelifa R, Trabelsi A, Jemmali M, Hani K (2002) In vitro antiviral activity of dermaseptins against herpes simplex virus type 1. J Med Virol 66(2):229–234
Bergaoui I, Zairi A, Tangy F, Aouni M, Selmi B, Hani K (2013) In vitro antiviral activity of dermaseptin S4 and derivatives from amphibian skin against herpes simplex virus type 2. J Med Virol 85(2):272–281
Bhandari D, Rafiq S, Gat Y, Gat P, Waghmare R, Kumar V (2020) A review on bioactive peptides: physiological functions, bioavailability and safety. International J Peptide Res Therapeut 1:139–150
Boopathi S, Poma AB, Kolandaivel P (2020) Novel 2019 coronavirus structure, mechanism of action, antiviral drug promises and rule out against its treatment. J Biomol Str Dynamics:1–14
Borst EM, Ständker L, Wagner K, Schulz TF, Forssmann WG, Messerle M (2013) A peptide inhibitor of cytomegalovirus infection from human hemofiltrate. Antimicrob Agents Chemother 57(10):4751–4760
Bosch BJ, Martina BE, Van Der Zee R, Lepault J, Haijema BJ, Versluis C, Heck AJ, De Groot R, Osterhaus AD, Rottier PJ (2004) Severe acute respiratory syndrome coronavirus (SARS-CoV) infection inhibition using spike protein heptad repeat-derived peptides. Proceed Nat Acad Sci 101(22):8455–8460
Camargo Filho I, Cortez DA, Ueda-Nakamura T, Nakamura CV, Dias Filho BP (2008) Antiviral activity and mode of action of a peptide isolated from Sorghum bicolor. Phytomed 15(3):202–208
Canonico PG, Pannier WL, Huggins JW, Rienehart KL (1982) Inhibition of RNA viruses in vitro and in Rift Valley fever-infected mice by didemnins A and B. Antimicrob Agents Chemother 22(4):696–697
Carriel-Gomes MC, Kratz JM, Barracco MA, Bachére E, Barardi CR, Simões CM (2007) In vitro antiviral activity of antimicrobial peptides against herpes simplex virus 1, adenovirus, and rotavirus. Mem Inst Oswaldo Cruz 102(4):469–472
Carvalho CA, Casseb SM, Gonçalves RB, Silva EV, Gomes AM, Vasconcelos PF (2017) Bovine lactoferrin activity against Chikungunya and Zika viruses. J Gen Virol 98(7):1749–1754
Chambers PCR, Easton AJ (1990) Heptad repeat sequences are located adjacent to hydrophobic regions in several types of virus fusion glycoproteins. J Gen Virol 71(12):3075–3080
Chattopadhyay S, Sinha NK, Banerjee S, Roy D, Chattopadhyay D, Roy S (2006) Small cationic protein from a marine turtle has β-defensin-like fold and antibacterial and antiviral activity. Proteins: Struct Funct Bioinformatics 64(2):524–531
Chen Y, Cao L, Zhong M, Zhang Y, Han C, Li Q, Yang J, Zhou D, Shi W, He B, Liu F (2012) Anti-HIV-1 activity of a new scorpion venom peptide derivative Kn2-7. PLoS One 7(4):e34947
Chen JM, Fan YC, Lin JW, Chen YY, Hsu WL, Chiou SS (2017) Bovine lactoferrin inhibits dengue virus infectivity by interacting with heparan sulfate, low-density lipoprotein receptor, and DC-SIGN. Int J Mol Sci 18(9):1957
Chernysh S, Kim SI, Bekker G, Pleskach VA, Filatova NA, Anikin VB, Platonov VG, Bulet P (2002) Antiviral and antitumor peptides from insects. Proc Natl Acad Sci 99(20):12628–12632
Chinchar VG, Bryan L, Silphadaung U, Noga E, Wade D, Rollins-Smith L (2004) Inactivation of viruses infecting ectothermic animals by amphibian and piscine antimicrobial peptides. Virol 323(2):268–275
Chuck CP, Chow HF, Wan DCC, Wong KB (2011) Profiling of substrate specificities of 3C-like proteases from group 1, 2a, 2b, and 3 coronaviruses. PLoS One 6(11):e27228
Chuck CP, Chen C, Ke Z, Wan DC, Chow HF, Wong KB (2013) Design, synthesis and crystallographic analysis of nitrile-based broad-spectrum peptidomimetic inhibitors for coronavirus 3C-like proteases. Eur J Med Chem 59:1–6
Crack LR, Jones L, Malavige GN, Patel V, Ogg GS (2012) Human antimicrobial peptides LL-37 and human β-defensin-2 reduce viral replication in keratinocytes infected with varicella zoster virus. Clin Exper Dermatol: Exper Dermatol 37(5):534–543
Craik DJ, Du J (2017) Cyclotides as drug design scaffolds. Curr Opin Chem Biol 38:8–16
Cuzin L, Alvarez M (2003) Enfuvirtide for prophylaxis against HIV infection. The New Eng J Med 349(22):2169–2170
Daliri EBM, Oh DH, Lee BH (2017) Bioactive peptides. Foods 6(5):32
Daly NL, Gustafson KR, Craik DJ (2004) The role of the cyclic peptide backbone in the anti-HIV activity of the cyclotide kalata B1. FEBS Lett 574(1-3):69–72
De Haan CA, Te Lintelo E, Li Z, Raaben M, Wurdinger T, Bosch BJ, Rottier PJ (2006) Cooperative involvement of the S1 and S2 subunits of the murine coronavirus spike protein in receptor binding and extended host range. J Virol 80(22):10909–10918
Dean RE, O’Brien LM, Thwaite JE, Fox MA, Atkins H, Ulaeto DO (2010) A carpet-based mechanism for direct antimicrobial peptide activity against vaccinia virus membranes. Peptides 31(11):1966–1972
Domashevskiy AV, Goss DJ (2015) Pokeweed antiviral protein, a ribosome inactivating protein: activity, inhibition and prospects. Toxins 7(2):274–298
Dutta K, Elmezayen AD, Alobaidi A, Zhu W, Morojava O, Shityakov S, Khalifa I (2020) A novel peptide analogue of spike glycoprotein shows antiviral properties against SARS-CoV-2: An in silico approach through molecular docking, molecular dynamics simulation and MM-PB/GBSA calculations. 10.21203/rs.3.rs-32796/v2
Egal M, Conrad M, MacDonald DL, Maloy WL, Motley M, Genco CA (1999) Antiviral effects of synthetic membrane-active peptides on herpes simplex virus, type 1. Int J Antimicrob Agents 13(1):57–60
El-Bitar AM, Sarhan MM, Aoki C, Takahara Y, Komoto M, Deng L, Moustafa MA, Hotta H (2015) Virocidal activity of Egyptian scorpion venoms against hepatitis C virus. Virol J 12(1):1–9
Esper F, Weibel C, Ferguson D, Landry ML, Kahn JS (2005) Evidence of a novel human coronavirus that is associated with respiratory tract disease in infants and young children. J Infect Dis 191(4):492–498
Galdiero S, Falanga A, Tarallo R, Russo L, Galdiero E, Cantisani M, Morelli G, Galdiero M (2013) Peptide inhibitors against herpes simplex virus infections. J Pept Sci 19(3):148–158
Gan YR, Huang H, Huang YD, Rao CM, Zhao Y, Liu JS, Wu L, Wei DQ (2006) Synthesis and activity of an octapeptide inhibitor designed for SARS coronavirus main proteinase. Peptides 27(4):622–625
Gao J, Lu G, Qi J, Li Y, Wu Y, Deng Y, Geng H, Li H, Wang Q, Xiao H, Tan W (2013) Structure of the fusion core and inhibition of fusion by a heptad repeat peptide derived from the S protein of Middle East respiratory syndrome coronavirus. J Virol 87(24):13134–13140
Gordon YJ, Huang LC, Romanowski EG, Yates KA, Proske RJ, McDermott AM (2005) Human cathelicidin (LL-37), a multifunctional peptide, is expressed by ocular surface epithelia and has potent antibacterial and antiviral activity. Curr Eye Res 30(5):385–394
Gustafson KR, Sowder RC, Henderson LE, Parsons IC, Kashman Y, Cardellina JH, McMahon JB, Buckheit RW Jr, Pannell LK, Boyd MR (1994) Circulins A and B. Novel human immunodeficiency virus (HIV)-inhibitory macrocyclic peptides from the tropical tree Chassalia parvifolia. J American Chem Soc 116(20):9337–9338
Hamming I, Timens W, Bulthuis ML, Lely AT, Navis GV, van Goor H (2004) Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. The Journal of Pathology: A J Pathol Soc Great Brit Ire 203(2):631–637
He M, Zhang H, Li Y, Wang G, Tang B, Zhao J, Huang Y, Zheng J (2018) Cathelicidin-derived antimicrobial peptides inhibit Zika virus through direct inactivation and interferon pathway. Front Immunol 9:722
Henriques ST, Craik DJ (2010) Cyclotides as templates in drug design. Drug Discov Today 15(1-2):57–64
Ho TY, Wu SL, Chen JC, Wei YC, Cheng SE, Chang YH, Liu HJ, Hsiang CY (2006) Design and biological activities of novel inhibitory peptides for SARS-CoV spike protein and angiotensin-converting enzyme 2 interaction. Antivir Res 69(2):70–76
Holly MK, Diaz K, Smith JG (2017) Defensins in viral infection and pathogenesis. Annu Rev Virol 4:369–391
Holthausen DJ, Lee SH, Kumar VT, Bouvier NM, Krammer F, Ellebedy AH, Wrammert J, Lowen AC, George S, Pillai MR, Jacob J (2017) An amphibian host defense peptide is virucidal for human H1 hemagglutinin-bearing influenza viruses. Imm 46(4):587–595
Hong W, Li T, Song Y, Zhang R, Zeng Z, Han S, Zhang X, Wu Y, Li W, Cao Z (2014) Inhibitory activity and mechanism of two scorpion venom peptides against herpes simplex virus type 1. Antivir Res 102:1–10
Hood JL, Jallouk AP, Campbell N, Ratner L, Wickline SA (2013) Cytolytic nanoparticles attenuate HIV-1 infectivity. Antivir Ther 18(1):95–103
Howell MD, Streib JE, Leung DY (2007) Antiviral activity of human β-defensin 3 against vaccinia virus. J Allergy Clin Immunol 119(4):1022–1025
Huang HN, Pan CY, Rajanbabu V, Chan YL, Wu CJ, Chen JY (2011) Modulation of immune responses by the antimicrobial peptide, epinecidin (Epi)-1, and establishment of an Epi-1-based inactivated vaccine. Biomat 32(14):3627–3636
Hultmark D, Steiner H, Rasmuson T, Boman HG (2005) Insect immunity. Purification and properties of three inducible bactericidal proteins from hemolymph of immunized pupae of Hyalophora cecropia. Eur J Biochem 106:7–16
Imai M, Okada N, Okada H (2000) Inhibition of HIV-1 infection by an intramolecular antisense peptide to T20 in gp160. Microbiol Immunol 44(3):205–212
Ireland DC, Wang CK, Wilson JA, Gustafson KR, Craik DJ (2008) Cyclotides as natural anti-HIV agents. Pept Sci 90(1):51–60
Ishag HZ, Li C, Huang L, Sun MX, Ni B, Guo CX, Mao X (2013) Inhibition of Japanese encephalitis virus infection in vitro and in vivo by pokeweed antiviral protein. Virus Res 171(1):89–96
Jack HW, Tzi BN (2005) Sesquin, a potent defensin-like antimicrobial peptide from ground beans with inhibitory activities toward tumor cells and HIV-1 reverse transcriptase. Peptides 26:1120–1126
Jang S, Hyun S, Kim S, Lee S, Lee IS, Baba M, Lee Y, Yu J (2014) Cell-penetrating, dimeric α-helical peptides: nanomolar inhibitors of HIV-1 transcription. Angew Chem 126(38):10250–10253
Kaspar AA, Reichert JM (2013) Future directions for peptide therapeutics development. Drug Discov Today 18(17-18):807–817
Keogan S, Passic S, Krebs FC (2012) Infection by CXCR4-tropic human immunodeficiency virus type 1 is inhibited by the cationic cell-penetrating peptide derived from HIV-1 tat. Int J Pept. https://doi.org/10.1155/2012/349427
Khalifa I, Li M, Mamet T, Li C (2019) Maltodextrin or gum Arabic with whey proteins as wall-material blends increased the stability and physiochemical characteristics of mulberry microparticles. Food Biosci 31:100445
Khalifa I, Zhu W, Mohammed HHH, Dutta K, Li C (2020a) Tannins inhibit SARS-CoV-2 through binding with catalytic dyad residues of 3CLpro: An in silico approach with 19 structural different hydrolysable tannins. J Food Biochem 44:e13432. https://doi.org/10.1111/jfbc.13432
Khalifa I, Nawaz A, Sobhy R, Althwab SA, Barakat H (2020b) Polyacylated anthocyanins constructively network with catalytic dyad residues of 3CLpro of 2019-nCoV than monomeric anthocyanins: a structural-relationship activity study with 10 anthocyanins using in-silico approaches. J Mol Graph Model 100:107690
Khalifa I, Xia D, Dutta K, Peng J, Jia Y, Li C (2020c) Mulberry anthocyanins exert anti-AGEs effects by selectively trapping glyoxal and structural-dependently blocking the lysyl residues of β-lactoglobulins. Bioorg Chem 96:103615
Kilby JM, Hopkins S, Venetta TM, DiMassimo B, Cloud GA, Lee JY, Alldredge L, Hunter E, Lambert D, Bolognesi D, Matthews T (1998) Potent suppression of HIV-1 replication in humans by T-20, a peptide inhibitor of gp41-mediated virus entry. Nat Med 4(11):1302–1307
Kliger Y, Gallo SA, Peisajovich SG, Muñoz-Barroso I, Avkin S, Blumenthal R, Shai Y (2001) Mode of action of an antiviral peptide from HIV-1 inhibition at a post-lipid mixing stage. J Biol Chem 276(2):1391–1397
Kuczer M, Midak-Siewirska A, Zahorska R, Łuczak M, Konopińska D (2011) Further studies on the antiviral activity of alloferon and its analogues. J Pept Sci 17(11):715–719
Kuczer M, Majewska A, Zahorska R (2013) New alloferon analogues: synthesis and antiviral properties. Chem Biol Drug Des 81:302–309
Lai CC, Shih TP, Ko WC, Tang HJ, Hsueh PR (2020) Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and corona virus disease-2019 (COVID-19): the epidemic and the challenges. Int J Antimicrob Agents 17:105924
Li F (2016) Structure, function, and evolution of coronavirus spike proteins. Ann Rev Virol 3:237–261
Li S, Zhou H, Huang G, Liu N (2009) Inhibition of HBV infection by bovine lactoferrin and iron-, zinc-saturated lactoferrin. Med Microbiol Immunol 198:19–25
Li Q, Zhao Z, Zhou D, Chen Y, Hong W, Cao L, Yang J, Zhang Y, Shi W, Cao Z, Wu Y (2011) Virucidal activity of a scorpion venom peptide variant mucroporin-M1 against measles, SARS-CoV and influenza H5N1 viruses. Peptides 32(7):1518–1525
Li W, Feng Y, Kuang Y, Zeng W, Yang Y, Li H, Jiang Z, Li M (2014) Construction of eukaryotic expression vector with mBD1-mBD3 fusion genes and exploring its activity against influenza A virus. Viruses 6(3):1237–1252
Liu IJ, Kao CL, Hsieh SC, Wey MT, Kan LS, Wang WK (2009) Identification of a minimal peptide derived from heptad repeat (HR) 2 of spike protein of SARS-CoV and combination of HR1-derived peptides as fusion inhibitors. Antivir Res 81(1):82–87
Liya G, Yuguang W, Jian L, Huaiping Y, Xue H, Jianwei H, Jiaju M, Youran L, Chen M, Yiqing J (2020) Studies on viral pneumonia related to novel coronavirus SARS-CoV-2, SARS-CoV, and MERS-CoV: a literature review. APMIS 128:423–432. https://doi.org/10.1111/apm.13047
Lorin C, Saidi H, Belaid A, Zairi A, Baleux F, Hocini H, Bélec L, Hani K, Tangy F (2005) The antimicrobial peptide dermaseptin S4 inhibits HIV-1 infectivity in vitro. Virol 334(2):264–275
Lou Z, Sun Y, Rao Z (2014) Current progress in antiviral strategies. Trends Pharmacol Sci 35(2):86–102
Lu XT, Sims AC, Denison MR (1998) Mouse hepatitis virus 3C-like protease cleaves a 22-kilodalton protein from the open reading frame 1a polyprotein in virus-infected cells and in vitro. J Virol 72(3):2265–2271
Lu Z, Van Wagoner RM, Harper MK, Baker HL, Hooper JN, Bewley CA, Ireland CM (2011) Mirabamides E-H, HIV-inhibitory depsipeptides from the sponge Stelletta clavosa. J Nat Prod 74(2):185–193
Lu G, Hu Y, Wang Q, Qi J, Gao F, Li Y, Zhang Y, Zhang W, Yuan Y, Bao J, Zhang B (2013) Molecular basis of binding between novel human coronavirus MERS-CoV and its receptor CD26. Nature 500(7461):227–231
Marcocci ME, Amatore D, Villa S, Casciaro B, Aimola P, Franci G, Grieco P, Galdiero M, Palamara AT, Mangoni ML, Nencioni L (2018) The amphibian antimicrobial peptide temporin b inhibits in vitro herpes simplex virus 1 infection. Antimicrob Agents Chemother 62(5)
Marr AK, Jenssen H, Moniri MR, Hancock RE, Panté N (2009) Bovine lactoferrin and lactoferricin interfere with intracellular trafficking of Herpes simplex virus-1. Biochimie 91(1):160–164
Matanic VCA, Castilla V (2004) Antiviral activity of antimicrobial cationic peptides against Junin virus and herpes simplex virus. Int J Antimicrob Agents 23(4):382–389
Matsumura T, Sugiyama N, Murayama A, Yamada N, Shiina M, Asabe S, Wakita T, Imawari M, Kato T (2016) Antimicrobial peptide LL-37 attenuates infection of hepatitis C virus. Hepatol Res 46(9):924–932
Mechlia MB, Belaid A, Castel G, Jallet C, Mansfield KL, Fooks AR, Hani K, Tordo N (2019) Dermaseptins as potential antirabies compounds. Vacc 37(33):4694–4700
Mehany T, Khalifa I, Barakat H, Althwab SA, Alharbi YM, El-Sohaimy S (2021) Polyphenols as promising biologically active substances for preventing SARS-CoV-2: A review with research evidence and underlying mechanisms. Food Biosci 100891
Meyer-Hoffert U, Schwarz T, Schröder J-M, Gläser R (2008) Expression of human beta-defensin-2 and -3 in Verruca vulgaris and Condylomata acuminata. J Eur Acad Dermatol Venereol 22:1050–1054
Migliolo L, Silva ON, Silva PA, Costa MP, Costa CR, Nolasco DO, Barbosa JA, Silva MR, Bemquerer MP, Lima LM, Romanos MT (2012) Structural and functional characterization of a multifunctional alanine-rich peptide analogue from Pleuronectes americanus. PLoS One 7(10):e47047
Mistry N, Drobni P, Näslund J, Sunkari VG, Jenssen H, Evander M (2007) The anti-papillomavirus activity of human and bovine lactoferricin. Antivir Res 5(3):258–265
Monajjemi M, Shojaei S, Mollaamin F (2020) An overview on Coronaviruses family from past to Covid-19: introduce some inhibitors as antiviruses from Gillan’s plants. Biointerf Res Applied Chem 10(3):5575–5585
Monteiro JM, Oliveira MD, Dias RS, Nacif-Marçal L, Feio RN, Ferreira SO, Oliveira LL, Silva CC, Paula SO (2018) The antimicrobial peptide HS-1 inhibits dengue virus infection. Virol 514:79–87
Mooney C, Haslam NJ, Pollastri G, Shields DC (2012) Towards the improved discovery and design of functional peptides: common features of diverse classes permit generalized prediction of bioactivity. PLoS One 7(10):e45012
Mora L, Escudero E, Arihara K, Toldrá F (2015) Antihypertensive effect of peptides naturally generated during Iberian dry-cured ham processing. Food Res Int 78:71–78
Ngai PH, Ng TB (2005) Phaseococcin, an antifungal protein with antiproliferative and anti-HIV-1 reverse transcriptase activities from small scarlet runner beans. Biochem Cell Biol 83:212–220
NHFPC (National Health and Family Planning Commission of People’s Republic of China)(2015) Guideline on diagnosis and treatment of Middle East respiratory syndrome. Chin J Viral Dis 5:37–39
Otwinowski ZWM (1997) Processing of X-ray diffraction data collected in oscillation mode macromolecular crystallography. Methods Enzymol 276:306–315
Pärn K, Eriste E, Langel Ü (2015) The antimicrobial and antiviral applications of cell-penetrating peptides. In Cell-penetrating peptides (pp. 223-245). Humana Press, New York, NY
Peterson JJ, Beecher GR, Bhagwat SA, Dwyer JT, Gebhardt SE, Haytowitz DB, Holden JM (2006) Flavanones in grapefruit, lemons, and limes: a compilation and review of the data from the analytical literature. J Food Compos Anal 19:74–80
Prompetchara E, Ketloy C, Palaga T (2020) Immune responses in COVID-19 and potential vaccines: lessons learned from SARS and MERS epidemic. Asian Pac J Allergy Immunol 38(1):1–9
Quiñones-Mateu ME, Lederman MM, Feng Z, Chakraborty B, Weber J, Rangel HR, Marotta ML, Mirza M, Jiang B, Kiser P, Medvik K (2003) Human epithelial β-defensins 2 and 3 inhibit HIV-1 replication. Aids 17(16):39–48
Rabaan AA, Al-Ahmed SH, Haque S, Sah R, Tiwari R, Malik YS, Dhama K, Yatoo MI, Bonilla-Aldana DK, Rodriguez-Morales AJ (2020) SARS-CoV-2, SARS-CoV, and MERS-CoV: a comparative overview. Infez Med 28(2):174–184
Rajamohan F, Kurinov IV, Venkatachalam TK, Uckun FM (1999) Deguanylation of human immunodeficiency virus (HIV-1) RNA by recombinant pokeweed antiviral protein. Biochem Biophys Res Commun 263:419–424
Ren W, Qu X, Li W, Han Z, Yu M, Zhou P, Zhang SY, Wang LF, Deng H, Shi Z (2008) Difference in receptor usage between severe acute respiratory syndrome (SARS) coronavirus and SARS-like coronavirus of bat origin. J Virol 82(4):1899–1907
Rosa SGV, Santos WC (2020) Clinical trials on drug repositioning for COVID-19 treatment. Rev Panam Salud Publica 44:1
Rothan HA, Bahrani H, Rahman NA, Yusof R (2014) Identification of natural antimicrobial agents to treat dengue infection: in vitro analysis of latarcin peptide activity against dengue virus. BMC Microbiol 14(1):40
Salvatore M, García-Sastre A, Ruchala P, Lehrer RI, Chang T, Klotman ME (2007) β-defensin inhibits influenza virus replication by cell-mediated mechanism (s). J Infect Dis 196(6):835–843
Sample CJ, Hudak KE, Barefoot BE, Koci MD, Wanyonyi MS, Abraham S, Staats HF, Ramsburg EA (2013) A mastoparan-derived peptide has broad-spectrum antiviral activity against enveloped viruses. Peptides 48:96–105
Sencanski M, Radosevic D, Perovic V, Gemovic B, Stanojevic M, Veljkovic N, Glisic S (2015) Natural products as promising therapeutics for treatment of influenza disease. Curr Pharmaceut Des 21(38):5573–5588
Sharma A, Singla D, Rashid M, Raghava GPS (2014) Designing of peptides with desired half-life in intestine-like environment. BMC Bioinf 15(1):282
Shartouny JR, Jacob J (2019) Mining the tree of life: host defense peptides as antiviral therapeutics. In Seminars in cell & developmental biology (Vol. 88, pp. 147-155). Academic Press
Shereen MA, Khan S, Kazmi A, Bashir N, Siddique R (2020) COVID-19 infection: origin, transmission, and characteristics of human coronaviruses. J Adv Res 24:91–98
Shestakov A, Jenssen H, Nordström I, Eriksson K (2012) Lactoferricin but not lactoferrin inhibit herpes simplex virus type 2 infection in mice. Antivir Res 93:340–345
Simmons G, Gosalia DN, Rennekamp AJ, Reeves JD, Diamond SL, Bates P (2005) Inhibitors of cathepsin L prevent severe acute respiratory syndrome coronavirus entry. Proc Natl Acad Sci 102(33):11876–11881
Sitaram N, Nagaraj R (1999) Interaction of antimicrobial peptides with biological and model membranes: structural and charge requirements for activity. Biochim Biophys Acta (BBA)-Biomemb 1462(1-2):29–54
Sousa FH, Casanova V, Findlay F, Stevens C, Svoboda P, Pohl J, Proudfoot L, Barlow PG (2017) Cathelicidins display conserved direct antiviral activity towards rhinovirus. Peptides 95:76–83
Steffen I, Pohlmann S (2010) Peptide-based inhibitors of the HIV envelope protein and other class I viral fusion proteins. Curr Pharm Design 16(9):1143–1158
Sun Y, Zhang H, Shi J, Zhang Z, Gong R (2017) Identification of a novel inhibitor against Middle East respiratory syndrome coronavirus. Viruses 9(9):255
Tiwari V, Liu J, Valyi-Nagy T, Shukla D (2011) Anti-heparan sulfate peptides that block herpes simplex virus infection in vivo. J Biol Chem 286(28):25406–25415
Tomlinson JA, Walker VM, Flewett TH, Barclay GR (1974) The inhibition of infection by cucumber mosaic virus and influenza virus by extracts from Phytolacca americana. J Gen Virol 22:225–232
Uckun FM, Rajamohan F, Pendergrass S, Ozer Z, Waurzyniak B, Mao C (2003) Structure-based design and engineering of a nontoxic recombinant pokeweed antiviral protein with potent anti-human immunodeficiency virus activity. Antimicrob Agents Chemother 47(3):1052–1061
Uckun FM, Rustamova L, Vassilev AO, Tibbles HE, Petkevich AS (2005) CNS activity of pokeweed anti-viral protein (PAP) in mice infected with lymphocytic choriomeningitis virus (LCMV). BMC Infect Dis 5(1):9
Van der Strate BW, Beljaars L, Molema G, Harmsen MC, Meijer DK (2001 Dec 1) Antiviral activities of lactoferrin. Antivir Res 52(3):225–239
VanCompernolle SE, Taylor RJ, Oswald-Richter K, Jiang J, Youree BE, Bowie JH, Tyler MJ, Conlon JM, Wade D, Aiken C, Dermody TS (2005) Antimicrobial peptides from amphibian skin potently inhibit human immunodeficiency virus infection and transfer of virus from dendritic cells to T cells. J Virol 79(18):11598–11606
Vilas Boas LC, de Lima LM, Migliolo L, Mendes GD, de Jesus MG, Franco OL, Silva PA (2017) Linear antimicrobial peptides with activity against herpes simplex virus 1 and Aichi virus. Pept Sci 108(2):e22871
Vilas Boas LC, Campos ML, Berlanda RL, de Carvalho NN, Franco OL (2019) Antiviral peptides as promising therapeutic drugs. Cell Mol Life Sci 76(18):3525–3542
Wachinger M, Kleinschmidt A, Winder D, von Pechmann N, Ludvigsen A, Neumann M, Holle R, Salmons B, Erfle V, Brack-Werner R (1998) Antimicrobial peptides melittin and cecropin inhibit replication of human immunodeficiency virus 1 by suppressing viral gene expression. J Gen Virol 79(4):731–740
Wang W, Owen SM, Rudolph DL, Cole AM, Hong T, Waring AJ, Lal RB, Lehrer RI (2004) Activity of α-and θ-defensins against primary isolates of HIV-1. The J Immunol 173(1):515–520
Wang G, Watson KM, Peterkofsky A, Buckheit RW (2010) Identification of novel human immunodeficiency virus type 1-inhibitory peptides based on the antimicrobial peptide database. Antimicrob Agents Chemother 54:1343–1346
Wang WY, Wong JH, Ip DT, Wan DC, Cheung RC, Ng TB (2016) Bovine lactoferrampin, human lactoferricin, and lactoferrin 1-11 inhibit nuclear translocation of HIV integrase. Appl Biochem Biotechnol 79(7):1202–1212
Watanabe S, Takada A, Watanabe T, Ito H, Kida H, Kawaoka Y (2000) Functional importance of the coiled-coil of the Ebola virus glycoprotein. J Virol 74(21):10194–10201
Weidmann J, Craik DJ (2016) Discovery, structure, function, and applications of cyclotides: circular proteins from plants. J Exp Bot 67(16):4801–4812
Wild C, Greenwell T, Matthews T (1993) A synthetic peptide from HIV-1 gp41 is a potent inhibitor of virus-mediated cell-cell fusion. AIDS Res Human Retrovir 9(11):1051–1053
Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, Graham BS, McLellan JS (2020) Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Sci 367(6483):1260–1263
Xia S, Xu W, Wang Q, Wang C, Hua C, Li W, Lu L, Jiang S (2018) Peptide-based membrane fusion inhibitors targeting HCoV-229E spike protein HR1 and HR2 domains. Int J Mol Sci 19(2):487
Yan R, Zhao Z, He Y, Wu L, Cai D, Hong W, Wu Y, Cao Z, Zheng C, Li W (2011) A new natural α-helical peptide from the venom of the scorpion Heterometrus petersii kills HCV. Peptides 32(1):11–19
Yao C, Xi C, Hu K, Gao W, Cai X, Qin J, Lv S, Du C, Wei Y (2018) Inhibition of enterovirus 71 replication and viral 3C protease by quercetin. Virol J 15(1):116
Yasin B, Pang M, Turner JS, Cho Y, Dinh NN, Waring AJ, Lehrer RI, Wagar EA (2000) Evaluation of the inactivation of infectious Herpes simplex virus by host-defense peptides. Eur J Clin Microbiol Infect Dis 19(3):187–194
Ying T, Du L, Ju TW, Prabakaran P, Lau CC, Lu L, Liu Q, Wang L, Feng Y, Wang Y, Zheng BJ (2014) Exceptionally potent neutralization of Middle East respiratory syndrome coronavirus by human monoclonal antibodies. J Virol 88(14):7796–7805
Zapata W, Aguilar-Jiménez W, Feng Z, Weinberg A, Russo A, Potenza N, Estrada H, Rugeles MT (2016) Identification of innate immune antiretroviral factors during in vivo and in vitro exposure to HIV-1. Microbes Infect 18(3):211–219
Zeng Z, Zhang R, Hong W, Cheng Y, Wang H, Lang Y, Ji Z, Wu Y, Li W, Xie Y, Cao Z (2018) Histidine-rich modification of a scorpion-derived peptide improves bioavailability and inhibitory activity against HSV-1. Theranostics 8(1):199–211
Zhang H, Zhao Q, Bhattacharya S, Waheed AA, Tong X, Hong A, Heck S, Curreli F, Goger M, Cowburn D, Freed EO (2008) A cell-penetrating helical peptide as a potential HIV-1 inhibitor. J Mol Biol 378(3):565–580
Zhao Z, Hong W, Zeng Z, Wu Y, Hu K, Tian X, Li W, Cao Z (2012) Mucroporin-M1 inhibits hepatitis B virus replication by activating the mitogen-activated protein kinase (MAPK) pathway and down-regulating HNF4α in vitro and in vivo. J Biol Chem 287(36):30181–30190
Zhou P, Yang XL, Wang XG (2020) A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579(7798):270–273
Zhu J, Xiao G, Xu Y, Yuan F, Zheng C, Liu Y, Yan H, Cole DK, Bell JI, Rao Z, Tien P (2004) Following the rule: formation of the 6-helix bundle of the fusion core from severe acute respiratory syndrome coronavirus spike protein and identification of potent peptide inhibitors. Biochem Biophys Res Commun 319(1):283–288
Zhu W, Khalifa I, Peng J, Li C (2018) Position and orientation of gallated proanthocyanidins in lipid bilayer membranes: influence of polymerization degree and linkage type. Biomol Struct Dyn 36(11):2862–2875
Acknowledgements
The authors acknowledge database websites mentioned and Ms. Itthanan Suttikhana for assisting with the figure in this article.
Availability of data and material
Available upon request
Code availability
Available upon request
Author information
Authors and Affiliations
Contributions
TJA and IK conceptualized, wrote, and revised the manuscript. AN and NW proofread it. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval
Not applicable
Consent to participate
Not applicable
Consent for publication
Not applicable
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ashaolu, T.J., Nawaz, A., Walayat, N. et al. Potential “biopeptidal” therapeutics for severe respiratory syndrome coronaviruses: a review of antiviral peptides, viral mechanisms, and prospective needs. Appl Microbiol Biotechnol 105, 3457–3470 (2021). https://doi.org/10.1007/s00253-021-11267-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11267-1