Abstract
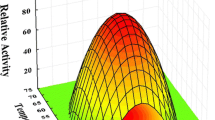
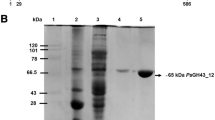
Arabinanases from glycoside hydrolase family GH93 are enzymes with exo-activity that hydrolyze the α-1,5 bonds between arabinose residues present on arabinan. Currently, several initiatives aiming to use byproducts rich in arabinan such as pectin and sugar beet pulp as raw material to produce various compounds of interest are being developed. However, it is necessary to use robust enzymes that have an optimal performance under pH and temperature conditions used in the industrial processes. In this work, the first GH93 from the thermophilic fungus Thermothielavioides terrestris (Abn93T) was heterologously expressed in Aspergillus nidulans, purified and biochemically characterized. The enzyme is a thermophilic glycoprotein (optimum activity at 70 °C) with prolonged stability in acid pHs (4.0 to 6.5). The presence of glycosylation affected slightly the hydrolytic capacity of the enzyme, which was further increased by 34% in the presence of 1 mM CoCl2. Small-angle X-ray scattering results show that Abn93T is a globular-like-shaped protein with a slight bulge at one end. The hydrolytic mechanism of the enzyme was elucidated using capillary zone electrophoresis and molecular docking calculations. Abn93T has an ability to produce (in synergism with arabinofuranosidases) arabinose and arabinobiose from sugar beet arabinan, which can be explored as fermentable sugars and prebiotics.
Key points
• Thermophilic exo-arabinanase from family GH93
• Molecular basis of arabinan depolymerization








Similar content being viewed by others
References
Alhassid A, Ben-David A, Tabachnikov O, Libster D, Naveh E, Zolotnitsky G, Shoham Y, Shoham G (2009) Crystal structure of an inverting GH 43 1,5-alpha-L-arabinanase from Geobacillus stearothermophilus complexed with its substrate. Biochem J 422(1):73–82. https://doi.org/10.1042/BJ20090180
Al-Tamimi MA, Palframan RJ, Cooper JM, Gibson GR, Rastall RA (2006) In vitro fermentation of sugar beet arabinan and arabino-oligosaccharides by the human gut microflora. J Appl Microbiol 100(2):407–414. https://doi.org/10.1111/j.1365-2672.2005.02780.x
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:410
Ardevol A, Rovira C (2015) Reaction mechanisms in carbohydrate-active enzymes: glycoside hydrolases and glycosyltransferases. Insights from ab Initio Quantum Mechanics/Molecular Mechanics Dynamic Simulations. J Am Chem Soc 137(24):7528–7547. https://doi.org/10.1021/jacs.5b01156
Arnold K, Bordoli L, Kopp J, Schwede T (2006) The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinform 22(2):195–201. https://doi.org/10.1093/bioinformatics/bti770
Berka RM, Grigoriev IV, Otillar R, Salamov A, Grimwood J, Reid I, Ishmael N, John T, Darmond C, Moisan MC, Henrissat B, Coutinho PM, Lombard V, Natvig DO, Lindquist E, Schmutz J, Lucas S, Harris P, Powlowski J, Bellemare A, Taylor D, Butler G, de Vries RP, Allijn IE, van den Brink J, Ushinsky S, Storms R, Powell AJ, Paulsen IT, Elbourne LD, Baker SE, Magnuson J, Laboissiere S, Clutterbuck AJ, Martinez D, Wogulis M, de Leon AL, Rey MW, Tsang A (2011) Comparative genomic analysis of the thermophilic biomass-degrading fungi Myceliophthora thermophila and Thielavia terrestris. Nat Biotechnol 29(10):922–927. https://doi.org/10.1038/nbt.1976
Berlowska J, Pielech-Przybylska K, Balcerek M, Dziekonska-Kubczak U, Patelski P, Dziugan P, Kregiel D (2016) Simultaneous saccharification and fermentation of sugar beet pulp for efficient bioethanol production. Biomed Res Int 2016:3154929–3154910. https://doi.org/10.1155/2016/3154929
Berto GL, Velasco J, Tasso Cabos Ribeiro C, Zanphorlin LM, Noronha Domingues M, Tyago Murakami M, Polikarpov I, de Oliveira LC, Ferraz A, Segato F (2019) Functional characterization and comparative analysis of two heterologous endoglucanases from diverging subfamilies of glycosyl hydrolase family 45. Enzym Microb Technol 120:23–35. https://doi.org/10.1016/j.enzmictec.2018.09.005
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Carapito R, Imberty A, Jeltsch JM, Byrns SC, Tam PH, Lowary TL, Varrot A, Phalip V (2009) Molecular basis of arabinobio-hydrolase activity in phytopathogenic fungi: crystal structure and catalytic mechanism of Fusarium graminearum GH93 exo-alpha-L-arabinanase. J Biol Chem 284(18):12285–12296. https://doi.org/10.1074/jbc.M900439200
Chakiath C, Lyons MJ, Kozak RE, Laufer CS (2009) Thermal Stabilization of Erwinia chrysanthemi pectin methylesterase a for application in a sugar beet pulp biorefinery. Appl Environ Microbiol 75(23):7343–7349. https://doi.org/10.1128/AEM.01010-09
Cheenpracha S, Jitonnom J, Komek M, Ritthiwigrom T, Laphookhieo S (2016) Acetylcholinesterase inhibitory activity and molecular docking study of steroidal alkaloids from Holarrhena pubescens barks. Steroids 108:92–98. https://doi.org/10.1016/j.steroids.2016.01.018
Cho C, Choi SY, Luo ZW, Lee SY (2015) Recent advances in microbial production of fuels and chemicals using tools and strategies of systems metabolic engineering. Biotechnol Adv 33(7):1455–1466. https://doi.org/10.1016/j.biotechadv.2014.11.006
Copley RR, Russel RB, Ponting CP (2001) Sialidase-like Asp-boxes: sequence-similar structures within different protein folds. Protein Sci 10:285–292. https://doi.org/10.1110/ps.31901
de Vries RP, Visser J (2001) Aspergillus enzymes involved in degradation of plant cell wall polysaccharides. Microbiol Mol Biol Rev 65(4):497–522, table of contents. https://doi.org/10.1128/MMBR.65.4.497-522.2001
Dey PM (1973) β-L-arabinosidase from Cajanus indicus: a new enzyme. Biochim Biophys Acta 302:393–398
Dey PM (1983) Further characterization of β-L-arabinosidase from Cajanus indicus. Biochim Biophys Acta 746:8–13
Endalur Gopinarayanan V, Nair NU (2019) Pentose metabolism in Saccharomyces cerevisiae: the need to engineer global regulatory systems. Biotechnol J 14(1):e1800364. https://doi.org/10.1002/biot.201800364
Evangelista RA, Liu MS, Chen FTA (1995) Characterization of 9-aminopyrene-1,4,6-trisulfonate-derivatized sugars by capillary electrophoresis with laser-induced fluorescence detection. Anal Chem 67:2239–2245
Franke D, Svergun DI (2009) DAMMIF, a program for rapid ab-initio shape determination in small-angle scattering. J Appl Crystallogr 42(Pt 2):342–346. https://doi.org/10.1107/S0021889809000338
Franke D, Petoukhov MV, Konarev PV, Panjkovich A, Tuukkanen A, Mertens HDT, Kikhney AG, Hajizadeh NR, Franklin JM, Jeffries CM, Svergun DI (2017) ATSAS 2.8: a comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J Appl Crystallogr 50(Pt 4):1212–1225. https://doi.org/10.1107/S1600576717007786
Gaskell A, Crennell S, Taylor GG (1995) The three domains of a bacterial sialidase: a beta-propeller, animmunoglobulin module and a galactose-binding jelly-roll. Struc 3(11):1197–1205
Gibson DG, Young L, Chuang RY, Venter JC, Hutchison CA 3rd, Smith HO (2009) Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6(5):343–345. https://doi.org/10.1038/nmeth.1318
Glusker JP (1980) Citrate conformation and chelation: enzymatic implications. Acc Chem Res 13:345–352
Goncalves TA, Damasio AR, Segato F, Alvarez TM, Bragatto J, Brenelli LB, Citadini AP, Murakami MT, Ruller R, Paes Leme AF, Prade RA, Squina FM (2012) Functional characterization and synergic action of fungal xylanase and arabinofuranosidase for production of xylooligosaccharides. Bioresour Technol 119:293–299. https://doi.org/10.1016/j.biortech.2012.05.062
Hanwell MD, Curtis DE, Lonie DC, Vandermeersch T, Zurek E, Hutchinson GR (2012) Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J Chem Inf 4:1–17
Henrissat B, Davies G (1997) Structural and sequence-based classification of glycoside hydrolases. Curr Opin Struct Biol 7(5):644
Hess B (2008) GROMACS 4: Algorithms for highly efficient, load-balanced, and scalable molecular simulation. J Chem Theory Comput 4:435–447
Hess B, Bekker H, Berendsen HJC, Fraaije JGEM (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18(12):1463–1472
Ichinose H, Fujimoto Z, Honda M, Harazono K, Nishimoto Y, Uzura A, Kaneko S (2009) A beta-l-Arabinopyranosidase from Streptomyces avermitilis is a novel member of glycoside hydrolase family 27. J Biol Chem 284(37):25097–25106. https://doi.org/10.1074/jbc.M109.022723
Jitonnom J, Hannongbua S (2018) Theoretical study of the arabinan hydrolysis by an inverting GH43 arabinanase. Mol Simul 44(8):631–637. https://doi.org/10.1080/08927022.2017.1422212
Jitonnom J, Lee VS, Nimmanpipug P, Rowlands HA, Mulholland AJ (2011) Quantum mechanics/molecular mechanics modeling of substrate-assisted catalysis in family 18 chitinases: conformational changes and the role of Asp142 in catalysis in ChiB. Biochem 50(21):4697–4711. https://doi.org/10.1021/bi101362g
Jitonnom J, Lomthaisong K, Lee VS (2012) Computational design of peptide inhibitor based on modifications of proregion from Plutella xylostella midgut trypsin. Chem Biol Drug Des 79(4):583–593. https://doi.org/10.1111/j.1747-0285.2011.01312.x
Jitonnom J, Limb MA, Mulholland AJ (2014) QM/MM free-energy simulations of reaction in Serratia marcescens Chitinase B reveal the protonation state of Asp142 and the critical role of Tyr214. J Phys Chem B 118(18):4771–4783. https://doi.org/10.1021/jp500652x
Jitonnom J, Mujika JI, van der Kamp MW, Mulholland AJ (2017) Quantum mechanics/molecular mechanics simulations identify the ring-opening mechanism of creatininase. Biochem 56(48):6377–6388. https://doi.org/10.1021/acs.biochem.7b01032
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79(2):926–935. https://doi.org/10.1063/1.445869
Kozin MB, Svergun DI (2000) Automated matching of high- and low-resolution structural models. J Appl Crystallogr 34:33–41
Kuhnel S, Hinz SW, Pouvreau L, Wery J, Schols HA, Gruppen H (2010) Chrysosporium lucknowense arabinohydrolases effectively degrade sugar beet arabinan. Bioresour Technol 101(21):8300–8307. https://doi.org/10.1016/j.biortech.2010.05.070
Kühnel S, Schols HA, Gruppen H (2011) Aiming for the complete utilization of sugar-beet pulp: examination of the effects of mild acid and hydrothermal pretreatment followed by enzymatic digestion. Biotech Biofuels 4:14
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Lang PT, Brozell SR, Mukherjee S, Pettersen EF, Meng EC, Thomas V, Rizzo RC, Case DA, James TL, Kuntz ID (2009) DOCK 6: combining techniques to model RNA-small molecule complexes. RNA 15(6):1219–1230. https://doi.org/10.1261/rna.1563609
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinf 23(21):2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Lees JG, Smith BR, Wien F, Miles AJ, Wallace BA (2004) CDtool-an integrated software package for circular dichroism spectroscopic data processing, analysis, and archiving. Anal Biochem 332(2):285–289. https://doi.org/10.1016/j.ab.2004.06.002
Mandels M, Sternberg D (1976) Recent advances in cellulose technology. J Ferment Technol 54:286
Mardones W, Callegari E, Eyzaguirre J (2015) Heterologous expression of a Penicillium purpurogenum exo-arabinanase in Pichia pastoris and its biochemical characterization. Fungal Biol 119(12):1267–1278. https://doi.org/10.1016/j.funbio.2015.09.009
Margolles A, de los Reyes-Gavilan CG (2003) Purification and functional characterization of a novel -L-arabinofuranosidase from Bifidobacterium longum B667. Appl Environ Microbiol 69(9):5096–5103. https://doi.org/10.1128/aem.69.9.5096-5103.2003
Matsuo N, Kaneko S, Kuno A, Kobayashi H, Kusakabe I (2000) Purification, characterization and gene cloning of two α-L-arabinofuranosidases from Streptomyces chartreusis GS901. Biochem J 346:9–15
Miller GL (1959) Use of dinitrosaIicyIic acid reagent for determination of reducing sugar. Anal Chem 31(3):426–428
Moon JS, Shin SY, Choi HS, Joo W, Cho SK, Li L, Kang JH, Kim TJ, Han NS (2015) In vitro digestion and fermentation properties of linear sugar-beet arabinan and its oligosaccharides. Carbohydr Polym 131:50–56. https://doi.org/10.1016/j.carbpol.2015.05.022
Mukherjee S, Balius TE, Rizzo RC (2010) Docking validation resources: protein family and ligand flexibility experiments. J Chem Inf Model 50:1986–2000
Petoukhov MV, Konarev PV, Kikhney AG, Svergun DI (2007) ATSAS 2.1 – towards automated and web-supported small-angle scattering data analysis. J Appl Crystallogr 40:223–228
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera--a visualization system for exploratory research and analysis. J Comput Chem 25(13):1605–1612. https://doi.org/10.1002/jcc.20084
Piiadov V, Ares de Araujo E, Oliveira Neto M, Craievich AF, Polikarpov I (2019) SAXSMoW 2.0: Online calculator of the molecular weight of proteins in dilute solution from experimental SAXS data measured on a relative scale. Protein Sci 28(2):454–463. https://doi.org/10.1002/pro.3528
Quistgaard EM, Thirup SS (2009) Sequence and structural analysis of the Asp-box motif and Asp-box beta-propellers; a widespread propeller-type characteristic of the Vps10 domain family and several glycoside hydrolase families. BMC Struct Biol 9:46. https://doi.org/10.1186/1472-6807-9-46
Sakamoto T, Thibault JF (2001) Exo-arabinanase of Penicillium chrysogenum able to release arabinobiose from alpha-1,5-L-arabinan. Appl Environ Microbiol 67(7):3319–3321. https://doi.org/10.1128/AEM.67.7.3319-3321.2001
Sakamoto T, Fujita T, Kawasaki H (2004) Transglycosylation catalyzed by a Penicillium chrysogenum exo-1,5-alpha-L-arabinanase. Biochim Biophys Acta 1674(1):85–90. https://doi.org/10.1016/j.bbagen.2004.06.001
Schneidman-Duhovny D, Hammel M, Sali A (2010) FoXS: a web server for rapid computation and fitting of SAXS profiles. Nucleic Acids Res 38(Web Server issue):W540–W544. https://doi.org/10.1093/nar/gkq461
Segato F, Damasio AR, Goncalves TA, de Lucas RC, Squina FM, Decker SR, Prade RA (2012) High-yield secretion of multiple client proteins in Aspergillus. Enzym Microb Technol 51(2):100–106. https://doi.org/10.1016/j.enzmictec.2012.04.008
Segato F, Dias B, Berto GL, de Oliveira DM, De Souza FHM, Citadini AP, Murakami MT, Damasio ARL, Squina FM, Polikarpov I (2017) Cloning, heterologous expression and biochemical characterization of a non-specific endoglucanase family 12 from Aspergillus terreus NIH2624. Biochim Biophys Acta, Proteins Proteomics 1865(4):395–403. https://doi.org/10.1016/j.bbapap.2017.01.003
Seiboth B, Metz B (2011) Fungal arabinan and L-arabinose metabolism. Appl Microbiol Biotechnol 89(6):1665–1673. https://doi.org/10.1007/s00253-010-3071-8
Semenova MV, Volkov PV, Rozhkova AM, Zorov IN, Sinitsyn AP (2018) Cloning, isolation, and properties of a new homologous exoarabinase from the Penicillium canescens fungus. Appl Biochem Microbiol 54(4):387–395. https://doi.org/10.1134/s0003683818040130
Shapiro AL, Viñuela E, Maizel JVJ (1967) Molecular weight estimation of polypeptide chains by electrophoresis in SDS-polyacrylamide gels. Biochem Biophys Res Commun 28(5):815–820
Shevchenko A, Wilm M, Vorm O, Mann M (1996) Mass spectrometric sequencing of proteins from silver-stained polyacrylamide gels. Anal Chem 68:850–858
Sogabe Y, Kitatani T, Yamaguchi A, Kinoshita T, Adachi H, Takano K, Inoue T, Mori Y, Matsumura H, Sakamoto T, Tada T (2011) High-resolution structure of exo-arabinanase from Penicillium chrysogenum. Acta Crystallogr D Biol Crystallogr 67(Pt 5):415–422. https://doi.org/10.1107/S0907444911006299
Steentoft C, Vakhrushev SY, Joshi HJ, Kong Y, Vester-Christensen MB, Schjoldager KT, Lavrsen K, Dabelsteen S, Pedersen NB, Marcos-Silva L, Gupta R, Bennett EP, Mandel U, Brunak S, Wandall HH, Levery SB, Clausen H (2013) Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology. EMBO J 32(10):1478–1488. https://doi.org/10.1038/emboj.2013.79
Vanommeslaeghe K, Hatcher E, Acharya C, Kundu S, Zhong S, Shim J, Darian E, Guvench O, Lopes P, Vorobyov I, Mackerell AD Jr (2010) CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J Comput Chem 31(4):671–690. https://doi.org/10.1002/jcc.21367
Velasco J, Oliva B, Mulinari EJ, Quintero LP, da Silva LA, Gonçalves AL, Gonçalves TA, Damasio A, Squina FM, Ferreira Milagres AM, Abdella A, Wilkins MR, Segato F (2019) Heterologous expression and functional characterization of a GH10 endoxylanase from Aspergillus fumigatus var. niveus with potential biotechnological application. Biotechnol Rep 24:e00382. https://doi.org/10.1016/j.btre.2019.e00382
Volkov VV, Svergun DI (2004) Uniqueness of ab initio shape determination in small-angle scattering. J Appl Chystallogr 36(3-1):860–864
Wang XW, Bai FY, Bensch K, Meijer M, Sun BD, Han YF, Crous PW, Samson RA, Yang FY, Houbraken J (2019) Phylogenetic re-evaluation of Thielavia with the introduction of a new family Podosporaceae. Stud Mycol 93:155–252. https://doi.org/10.1016/j.simyco.2019.08.002
Warshel A, Levitt M (1976) Theoretical studies of enzymic reactions: dielectric, electrostatic and steric stabilization of the carbonium ion in the reaction of lysozyme. J Mol Biol 103:227–249
Warwas ML, Yeung JH, Indurugalla D, Mooers AO, Bennet AJ, Moore MM (2010) Cloning and characterization of a sialidase from the filamentous fungus, Aspergillus fumigatus. Glycoconj J 27(5):533–548. https://doi.org/10.1007/s10719-010-9299-9
Wong DW, Chan VJ, Batt SB (2008) Cloning and characterization of a novel exo-alpha-1,5-L-arabinanase gene and the enzyme. Appl Microbiol Biotechnol 79(6):941–949. https://doi.org/10.1007/s00253-008-1504-4
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (2015) The I-TASSER Suite: protein structure and function prediction. Nat Methods 12(1):7–8. https://doi.org/10.1038/nmeth.3213
Zieminski K, Kowalska-Wentel M (2017) Effect of different sugar beet pulp pretreatments on biogas production efficiency. Appl Biochem Biotechnol 181(3):1211–1227. https://doi.org/10.1007/s12010-016-2279-1
Funding
This work was financially supported by the São Paulo Research Foundation (FAPESP), grants #2014/18714-2, #2014/06923-6, and #2017/00525-0 to FS; #2017/16089-1 to TAG, #2019/01165-0 to JV; #2015/50590-4 to FMS and #2019/06663-8 to BO; Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), grants #442352/2014-0 to LCO; 305748/2017-3 and 428527/2018-3 to FMS and #443916/2014-4, and #302627/2018-9 to FS. This study was also financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brazil (CAPES)—Finance Code 001. The molecular dynamics simulations and molecular docking were performed at the CENAPAD-SP (Centro Nacional de Processamento de Alto Desempenho em São Paulo), project UNICAMP/FINEP-MCT.
Author information
Authors and Affiliations
Contributions
JV, BO, ALG, ASL, EVM, GF, AM, and BAF performed the experiments and data interpretation and drafted the manuscript. TAG and FMS performed experiments and data interpretation related to CZE studies. MASK, IP, and LCO performed experiments and data interpretation related to molecular docking and SAXS. ALG, JV, and FS performed the phylogenetic analyses and data interpretation. JV, LCO, and IP participated in the design of the study and data interpretation and reviewed the manuscript. FS conceived the study, participated in data interpretation, and reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 2.66 mb).
Rights and permissions
About this article
Cite this article
Velasco, J., Oliva, B., Gonçalves, A.L. et al. Functional characterization of a novel thermophilic exo-arabinanase from Thermothielavioides terrestris. Appl Microbiol Biotechnol 104, 8309–8326 (2020). https://doi.org/10.1007/s00253-020-10806-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10806-6