Abstract
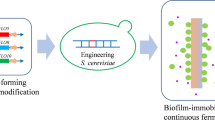
Biofilm-based fermentation, as a new immobilisation strategy, is beneficial for industrial fermentation due to its excellent environmental resistance, high productivity and continuous fermentation relative to calcium alginate-immobilised fermentation. These two techniques differ mainly regarding cell stages. Here, we describe the cell phenotype of Saccharomyces cerevisiae biofilm-based fermentation and compare cell cycle stages with those during immobilisation in calcium alginate. Most cells in the biofilm-based fermentation adhered to the cotton-fibre carrier of the biofilm and were in the G2/M phase whereas alginate-embedded cells were in the G1/G0 phase. Deletion of the RIM15 gene, which regulates cell cycle progression according to nutritional status, hampered the cell cycle arrest observed in alginate-embedded cells, enhanced biofilm formation and improved fermentation ability. The improved biofilm formation shown by the rim15△ strain could be attributed to an increase in the expression level of the adhesion protein FLO11 and synthesis of trehalose. These findings suggest that the extracellular environment is mainly responsible for the difference between biofilm-based fermentation and alginate-embedded fermentation, and that RIM15 plays an essential role in cell cycle progression.
Key points
• In the biofilm, S. cerevisiae cell populations were mostly in the G2/M phase while alginate-embedded cells were arrested in the G1/G0 phase.
• The RIM15 gene partially influenced the cell cycle progression observed during ethanol fermentation.
• Biofilm-based cells were actively adsorbed on the physical carrier.
• Biofilm immobilisation could maintain cell division activity explaining its fermentation efficiency.




Similar content being viewed by others
Data availability
The authors promise the availability of supporting data.
References
Basak S, Rajurkar MN, Attal RO, Mallick SK (2013) Biofilms: a challenge to medical fraternity in infection control. Intech. 3:57–74
Blankenship JR, Mitchell AP (2006) How to build a biofilm: a fungal perspective. Curr Opin Microbiol 9:588–594
Castro A, Lorca T (2018) Greatwall kinase at a glance. J Cell Sci 131:1–6
Chen Y, Liu Q, Zhou T, Li B, Yao S, Li A, Wu J, Ying H (2013) Ethanol production by repeated batch and continuous fermentations by Saccharomyces cerevisiae immobilized in a fibrous bed bioreactor. J Microbiol Biotechnol 23:511–517
Chen T, Liu N, Ren P, Xi X, Yang L, Sun W, Yu B, Ying H, Ouyang P, Liu D, Chen Y (2019) Efficient biofilm-based fermentation strategies for L-threonine production by Escherichia coli. Front Microbiol 10:1–11
Choi JW, Zhou W, Nie ZW, Niu YJ, Shin KT, Cui XS (2019) Spindlin1 alters the metaphase to anaphase transition in meiosis I through regulation of BUB3 expression in porcine oocytes. J Cell Physiol 234:8963–8974
Costanzo M, Nishikawa JL, Tang X, Millman JS, Schub O, Breitkreuz K, Dewar D, Rupes I, Andrews B, Tyers M (2004) CDK activity antagonizes Whi5, an inhibitor of G1/S transcription in yeast. Cell 117:899–913
Dahmann C, Futcher B (1995) Specialization of B-type cyclins for mitosis or meiosis in S. cerevisiae. Genetics 140:957–963
Daisuke W, Satoru N, Yoshikazu O, Yoichiro K, Yan Z, Takeshi A, Hitoshi S (2011) Ethanol fermentation driven by elevated expression of the G1 cyclin gene CLN3 in sake yeast. J Biosci Bioeng 112:577–582
Flemming HC, Wingender J (2010) The biofilm matrix. Nat Rev Microbiol 8:623–633
Galazzo JL, Bailey JE (1990) Growing Saccharomyces cerevisiae in calcium-alginate beads induces cell alterations which accelerate glucose conversion to ethanol. Biotechnol Bioeng 36:417–426
Gòdia F, Casas C, Castellano B, Solà C (1987) Immobilized cells: behaviour of carrageenan entrapped yeast during continuous ethanol fermentation. Appl Microbiol Biotechnol 26:342–346
Hinfray C, Jouenne T, Mignot L, Junter GA (1995) Influence of the oxygenation level on d-xylose fermentation by free and agar-entrapped cultures of Candida shehatae. Appl Microbiol Biotechnol 42:682–687
Hoyer LL (2001) The ALS gene family of Candida albicans. Trends Microbiol 185:176–180
Izmirlioglu G, Demirci A (2016) Ethanol production in biofilm reactors from potato waste hydrolysate and optimization of growth parameters for Saccharomyces cerevisiae. Fuel 181:643–651
Jorgensen P, Nishikawa JL, Breitkreutz BJ, Tyers M (2002) Systematic identification of pathways that couple cell growth and division in yeast. Science 297:395–400
Joseph R, Naugonly A, Feldman M, Herzog IM, Fridman M, Cohen Y (2016) Cationic pillararenes potently inhibit biofilm formation without affecting bacterial growth and viability. J Am Chem Soc 138:754–757
Kaplan JB, Ragunath C, Ramasubbu N, Fine DH (2003) Detachment of Actinobacillus actinomycetemcomitans biofilm cells by an endogenous β-hexosaminidase activity. J Bacteriol 185:4693–4698
Karatan E, Watnick P (2009) Signals, regulatory networks, and materials that build and break bacterial biofilms. Microbiol Mol Biol Rev 73:310–347
Kishimoto T (2015) Entry into mitosis: a solution to the decades-long enigma of MPF. Chromosoma:124417–124428
Li Z, Chen Y, Liu D, Zhao N, Cheng H, Ren H, Guo T, Niu H, Zhuang W, Wu J, Ying H (2015) Involvement of glycolysis/gluconeogenesis and signaling regulatory pathways in Saccharomyces cerevisiae biofilms during fermentation. Front Microbiol 6:1–12
Liu D, Chen Y, Ding FY, Zhao T, Wu JL, Guo T, Ren HF, Li BB, Niu HQ, Cao Z, Lin XQ, Xie JJ, He XJ, Ying HJ (2014) Biobutanol production in a Clostridium acetobutylicum biofilm reactor integrated with simultaneous product recovery by adsorption. Biotechnol Biofuels 7:1–13
Liu Q, Cheng H, Wu J, Chen X, Ying H, Zhou P, Chen Y (2015) Long-term production of fuel ethanol by immobilized yeast in repeated-batch simultaneous saccharification and fermentation of cassava. Energy Fuel 29:185–190
Love G, Gough S, Brady D, Barron N, Nigam P, Singh D, Marchant R, McHale AP (1998) Continuous ethanol fermentation at 45°C using Kluyveromyces marxianus IMB3 immobilized in calcium alginate and kissiris. Bioprocess Eng 18:187–189
Manitpisitkul P, Chiou WL (1993) Intravenous verapamil kinetics in rats: marked arteriovenous concentration difference and comparison with humans. Biopharm Drug Dispos 14:555–566
Moreno-García J, García-Martínez T, Mauricio JC, Moreno J (2018) Yeast immobilization systems for alcoholic wine fermentations: actual trends and future perspectives. Front Microbiol 9:241–253
Nagarajan S, Kruckeberg AL, Schmidt KH, Kroll E, Hamilton M, McInnerney K, Summers R, Taylor T, Rosenzweig F (2014) Uncoupling reproduction from metabolism extends chronological lifespan in yeast. PNAS 111:1538–1547
Nagashima M, Azuma M, Noguchi S, Inuzuka K, Samejima H (1984) Continuous ethanol fermentation using immobilized yeast cells. Biotechnol Bioeng 26:992–997
Nikawa JN, Kawabata M (2000) PCR- and ligation-mediated synthesis of marker cassettes with long flanking homology regions for gene disruption in Saccharomyces cerevisiae. Biotechniques 28:1112–1116
Oehlen LJWM, Jeoung DI, Cross FR (1998) Cyclin-specific START events and the G1-phase specificity of arrest by mating factor in budding yeast. Mol Gen Genet 258:183–198
Oomuro M, Kato T, Zhou Y, Watanabe D, Motoyama Y, Yamagishi H, Akao T, Aizawa M (2016) Defective quiescence entry promotes the fermentation performance of bottom-fermenting brewer’s yeast. J Biosci Bioeng 122:577–582
Pedruzzi I, Dubouloz F, Cameroni E, Wanke V, Roosen J, Winderickx J, Virgilio CD (2003) TOR and PKA signaling pathways converge on the protein kinase Rim15 to control entry into G0. Mol Cell 12:1607–1613
Radosavljević M, Lević S, Belović M, Pejin J, Djukić-Vuković A, Mojović L, Nedović V (2019) Immobilization of lactobacillus rhamnosus in polyvinyl alcohol/calcium alginate matrix for production of lactic acid. Bioprocess Biosyst Eng 43:315–322
Rattanapan A, Limtong S, Phisalaphong M (2011) Ethanol production by repeated batch and continuous fermentations of blackstrap molasses using immobilized yeast cells on thin-shell silk cocoons. Appl Energy 88:4400–4404
Sarkar S, Dalgaard JZ, Millar JBA, Arumugam P (2014) The Rim15-Endosulfine-PP2ACdc55 signalling module regulates entry into gametogenesis and quiescence via distinct mechanisms in budding yeast. PLoS Genet 10:1607–1613
Schwob E, Nasmyth K (1993) CLB5 and CLB6, a new pair of B cyclins involved in DNA replication in Saccharomyces cerevisiae. Genes Dev 7:1160–1175
Shi X, Zou Y, Chen Y, Ying H (2018) Overexpression of THI4 and HAP4 improves glucose metabolism and ethanol production in Saccharomyces cerevisiae. Front Microbiol 9:1–10
Simon I, Barnett J, Hannett N, Harbison CT, Rinaldi NJ, Volkert TL, Wyrick JJ, Zetlinger J, Gifford DK, Jaakkola TS, Young RA (2001) Serial regulation of transcriptional regulators in the yeast cell cycle. Cell 26:697–708
Talarek N, Gueydon E, Schwob E (2017) Homeostatic control of start through negative feedback between Cln3-Cdk1 and Rim15/greatwall kinase in budding yeast. ELife 6:1–21
Tyson JJ, Novak B (2008) Temporal organization of the cell cycle. Curr Biol 18:759–768
Tzeng YW, Huang JN, Schuyler SC, Wu CH, Juang YL (2011) Functions of the mitotic B-type cyclins CLB1, CLB2, and CLB3 at mitotic exit antagonized by the CDC14 phosphatase. Fungal Genet Biol 48:966–978
Wang Z, Wang Y, Yang ST, Wang R, Ren H (2010) A novel honeycomb matrix for cell immobilization to enhance lactic acid production by Rhizopus oryzae. Bioresour Technol 101:5557–5564
Watanabe D, Araki Y, Zhou Y, Maeya N, Akao T, Shimoi H (2012) A loss-of-function mutation in the PAS kinase Rim15p is related to defective quiescence entry and high fermentation rates of Saccharomyces cerevisiae sake yeast strains. Appl Environ Microbiol 78:4008–4016
Watanabe D, Zhou Y, Hirata A, Sugimoto Y, Takagi K, Akao T, Ohya Y, Takagi H, Shimoi H (2016) Inhibitory role of greatwall-like protein kinase Rim15p in alcoholic fermentation via upregulating the UDP-glucose synthesis pathway in Saccharomyces cerevisiae. Appl Environ Microbiol:82340–82351
Watanabe D, Kaneko A, Sugimoto Y, Ohnuki S, Takagi H, Ohya Y (2017) Promoter engineering of the Saccharomyces cerevisiae RIM15 gene for improvement of alcoholic fermentation rates under stress conditions. J Biosci Bioeng 123:183–189
Wei M, Fabrizio P, Hu J, Ge H, Cheng C, Li L, Longo VD (2008) Life span extension by calorie restriction depends on Rim15 and transcription factors downstream of Ras/PKA, Tor, and Sch9. PLoS Genet 4:139–149
Yan S, Wang P, Zhai Z, Yao J (2011) Fuel ethanol production from concentrated food waste hydrolysates in immobilized cell reactors by Saccharomyces cerevisiae H058. J Chem Technol Biotechnol 86:731–738
Yang L, Zheng C, Chen Y, Ying H (2018) FLO genes family and transcription factor MIG1 regulate saccharomyces cerevisiae biofilm formation during immobilized fermentation. Front Microbiol 9:1–11
Yang L, Zheng C, Chen Y, Shi X, Ying Z, Ying H (2019) Nitric oxide increases biofilm formation in Saccharomyces cerevisiae by activating the transcriptional factor Mac1p and thereby regulating the transmembrane protein Ctr1. Biotechnol Biofuels 12:1–15
Zhao XQ, Li Q, He LY, Li F, Que WW, Bai FW (2012) Exploration of a natural reservoir of flocculating genes from various Saccharomyces cerevisiae strains and improved ethanol fermentation using stable genetically engineered flocculating yeast strains. Process Biochem 47:1612–1619
Zheng W, Yao M, Fang M, Pan L, Wang L, Yang J, Dong Z, Yao D (2019) Oncogenic Wnt3a: a candidate specific marker and novel molecular target for hepatocellular carcinoma. J Cancer 10:5862–5873
Funding
This work was supported by the key program of the National Natural Science Foundation of China (Grant No. 21636003); the Outstanding Youth Foundation of China (Grant No. SBK2017010373); the National Key Research and Development Program of China (Grant No. 2018YFA0902200, 2018yfb1501705); the Program for Changjiang Scholars and Innovative Research Team in University (IRT_14R28); Jiangsu National Synergetic Innovation Center for Advanced Materials (SICAM); the Technology Support Program of Jiangsu (Grant No. BE2014715); and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Contributions
CL participated in the design of the study, constructed the plasmids and strains, participated in the fermentation experiments, drafted the manuscript and revised the manuscript. SD, DZ, WS, LL and WZ participated in the fermentation experiments and biofilm characterisation experiment. YC, HY and DL participated in the design of the study. HY conceived of the study and participated in its design. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Consent for publication
Publication consented.
Ethics approval and consent to participate
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 1392 kb)
Rights and permissions
About this article
Cite this article
Liang, C., Ding, S., Sun, W. et al. Biofilm-based fermentation: a novel immobilisation strategy for Saccharomyces cerevisiae cell cycle progression during ethanol production. Appl Microbiol Biotechnol 104, 7495–7505 (2020). https://doi.org/10.1007/s00253-020-10770-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10770-1