Abstract
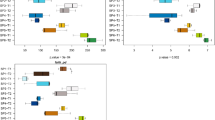
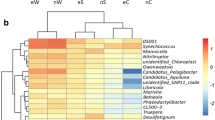
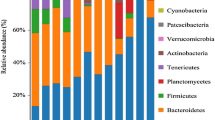
Although increasing levels of attention have been targeted towards aquaculture-associated bacteria, the bacterial community of animal intestines and its relationship with the aquaculture environment need to be further investigated. In this study, we used high-throughput sequencing to analyze the bacterial community of pond water, sediment, and the intestines of diseased and healthy animals. Our data showed that Proteobacteria, Firmicutes, Cyanobacteria, and Bacteroidetes were the dominant taxa of bacteria across all samples and accounted for more than 90% of the total sequence. Difference analysis and Venn diagrams showed that most of the intestinal bacterial OTUs (operational taxonomic units) of diseased and healthy animals were the same as those of sediment and water, indicating that the aquaculture environment was the main source of intestinal bacteria. Compared with healthy animals, a considerable reduction of OTUs was evident in diseased animals. Welch’s t test showed that the dominant bacterial taxa in sediment, water, and animal intestine were significantly different (p < 0.05) and each had its own unique dominant microorganisms. In addition, differences between the intestinal bacteria of healthy and diseased animals were represented by potential probiotics and pathogens, such as Bacillus, Vibrio, Oceanobacillus, and Lactococcus. Principal component analysis (PcoA) showed that a similar environment shaped a similar microbial structure. There was a large difference in the spectrum of intestinal bacteria in diseased animals; furthermore, the spectrum of intestinal bacteria in diseased animals was very different from the environment than in healthy animals. This study provides a theoretical basis for a relationship between the intestinal bacteria of healthy and diseased animals and the environment and provides guidance for environmental regulation and disease prevention in aquaculture areas.




Similar content being viewed by others
References
Akhter N, Wu B, Memon AM, Mohsin M (2015) Probiotics and prebiotics associated with aquaculture: A review. Fish Shellfish Immunol 45(2):733–741. https://doi.org/10.1016/j.fsi.2015.05.038
Austin B, Zhang X-H (2006) Vibrio harveyi: a significant pathogen of marine vertebrates and invertebrates. Lett Appl Microbiol 43(2):119–124. https://doi.org/10.1111/j.1472-765X.2006.01989.x
Balcázar JL, Id B, Ruiz-Zarzuela I, Cunningham D, Vendrell D, Múzquiz JL (2006) The role of probiotics in aquaculture. Vet Microbiol 114(3):173–186. https://doi.org/10.1016/j.vetmic.2006.01.009
Chaiyapechara S, Rungrassamee W, Suriyachay I, Kuncharin Y, Klanchui A, Karoonuthaisiri N, Jiravanichpaisal P (2012) Bacterial community associated with the intestinal tract of P. monodon in commercial farms. Microb Ecol 63(4):938–953
Chróst RJ (1990) Microbial ectoenzymes in aquatic environments. In: Overbeck J, Chróst RJ (eds) Aquatic Microbial Ecology: Biochemical and Molecular Approaches. Springer New York, New York, NY, pp 47–78
De Schryver P, Vadstein O (2014) Ecological theory as a foundation to control pathogenic invasion in aquaculture. ISME J 8:2360–2368. https://doi.org/10.1038/ismej.2014.84
Garcia T, Otto K, Kjelleberg S, Nelson DR (1997) Growth of Vibrio anguillarum in salmon intestinal mucus. Appl Environ Microbiol 63(3):1034–1039
Giatsis C, Sipkema D, Smidt H, Heilig H, Benvenuti G, Verreth J, Verdegem M (2015) The impact of rearing environment on the development of gut microbiota in tilapia larvae. Sci Rep 5:18206. https://doi.org/10.1038/srep18206
Gobi N, Malaikozhundan B, Sekar V, Shanthi S, Vaseeharan B, Jayakumar R, Khudus Nazar A (2016) GFP tagged Vibrio parahaemolyticus Dahv2 infection and the protective effects of the probiotic Bacillus licheniformis Dahb1 on the growth, immune and antioxidant responses in Pangasius hypophthalmus. Fish Shellfish Immunol 52:230–238. https://doi.org/10.1016/j.fsi.2016.03.006
Leser TD, Mølbak L (2009) Better living through microbial action: the benefits of the mammalian gastrointestinal microbiota on the host. Environ Microbiol 11(9):2194–2206. https://doi.org/10.1111/j.1462-2920.2009.01941.x
Li T, Li H, Gatesoupe F-J, She R, Lin Q, Yan X, Li J, Li X (2017) Bacterial signatures of “Red-Operculum” disease in the gut of crucian carp (Carassius auratus). Microbial Ecol 74(3):510–521. https://doi.org/10.1007/s00248-017-0967-1
LoGiudice K, Ostfeld RS, Schmidt KA, Keesing F (2003) The ecology of infectious disease: Effects of host diversity and community composition on Lyme disease risk. P Natl Acad Sci 100(2):567–571. https://doi.org/10.1073/pnas.0233733100
Mack DR, Michail S, Wei S, McDougall L, Hollingsworth MA (1999) Probiotics inhibit enteropathogenic E. coli adherence in vitro by inducing intestinal mucin gene expression. Am J Physiol 276(4):G941–G950. https://doi.org/10.1152/ajpgi.1999.276.4.g941
Martins P, Cleary DFR, Pires ACC, Rodrigues AM, Quintino V, Calado R, Gomes NCM (2013) Molecular analysis of bacterial communities and detection of potential pathogens in a recirculating aquaculture system for Scophthalmus maximus and Solea senegalensis. PloS One 8(11):e80847. https://doi.org/10.1371/journal.pone.0080847
Millet YA, Alvarez D, Ringgaard S, von Andrian UH, Davis BM, Waldor MK (2014) Insights into Vibriocholerae intestinal colonization from monitoring fluorescently labeled bacteria. PLoS Pathog 10(10):e1004405. https://doi.org/10.1371/journal.ppat.1004405
Moriarty DJW (1997) The role of microorganisms in aquaculture ponds. Aquaculture 151(1):333–349. https://doi.org/10.1016/S0044-8486(96)01487-1
Nerstedt A, Nilsson EC, Ohlson K, Håkansson J, Thomas Svensson L, Löwenadler B, Svensson UK, Mahlapuu M (2007) Administration of Lactobacillus evokes coordinated changes in the intestinal expression profile of genes regulating energy homeostasis and immune phenotype in mice. Brit J Nutr 97(6):1117–1127. https://doi.org/10.1017/S0007114507682907
Qi C, Wang L, Liu M, Jiang K, Wang M, Zhao W, Wang B (2017) Transcriptomic and morphological analyses of Litopenaeus vannamei intestinal barrier in response to Vibrio paraheamolyticus infection reveals immune response signatures and structural disruption. Fish Shellfish Immunol 70:437–450. https://doi.org/10.1016/j.fsi.2017.09.004
Rungrassamee W, Klanchui A, Chaiyapechara S, Maibunkaew S, Tangphatsornruang S, Jiravanichpaisal P, Karoonuthaisiri N (2013) Bacterial population in intestines of the black tiger shrimp (Penaeus monodon) under different growth stages. PloS One 8(4):e60802. https://doi.org/10.1371/journal.pone.0060802
Rungrassamee W, Klanchui A, Maibunkaew S, Chaiyapechara S, Jiravanichpaisal P, Karoonuthaisiri N (2014) Characterization of intestinal bacteria in wild and domesticated adult black tiger shrimp (Penaeus monodon). PloS One 9(3):e91853
Rungrassamee W, Klanchui A, Maibunkaew S, Karoonuthaisiri N (2016) Bacterial dynamics in intestines of the black tiger shrimp and the Pacific white shrimp during Vibrio harveyi exposure. J Invertebr Pathol 133:12–19. https://doi.org/10.1016/j.jip.2015.11.004
Rurangwa E, Verdegem MCJ (2015) Microorganisms in recirculating aquaculture systems and their management. Rev Aquacult 7(2):117–130. https://doi.org/10.1111/raq.12057
Sun F, Wang Y, Wang C, Zhang L, Tu K, Zheng Z (2019) Insights into the intestinal microbiota of several aquatic organisms and association with the surrounding environment. Aquaculture 507:196–202. https://doi.org/10.1016/j.aquaculture.2019.04.026
Tang Y, Tao P, Tan J, Mu H, Peng L, Yang D, Tong S, Chen L (2014) Identification of bacterial community composition in freshwater aquaculture system farming of Litopenaeus vannamei reveals distinct temperature-driven patterns. Int J Mol Sci 15(8):13663–13680
Wang C, Lin G, Yan T, Zheng Z, Chen B, Sun F (2014) The cellular community in the intestine of the shrimp Penaeus penicillatus and its culture environments. Fisheries Sci 80:1001–1007. https://doi.org/10.1007/s12562-014-0765-3
Wu S, Wang G, Angert ER, Wang W, Li W, Zou H (2012) Composition, diversity, and origin of the bacterial community in grass carp intestine. PloS One 7(2):e30440. https://doi.org/10.1371/journal.pone.0030440
Xiong J, Wang K, Wu J, Qiuqian L, Yang K, Qian Y, Zhang D (2015) Changes in intestinal bacterial communities are closely associated with shrimp disease severity. Appl Microbiol Biot 99(16):6911–6919. https://doi.org/10.1007/s00253-015-6632-z
Yang H, Zou S, Zhai L, Wang Y, Zhang F, An L, Yang G (2017) Pathogen invasion changes the intestinal microbiota composition and induces innate immune responses in the zebrafish intestine. Fish Shellfish Immunol 71:35–42. https://doi.org/10.1016/j.fsi.2017.09.075
Zhou Q, Li K, Jun X, Bo L (2009) Role and functions of beneficial microorganisms in sustainable aquaculture. Bioresource Technol 100(16):3780–3786. https://doi.org/10.1016/j.biortech.2008.12.037
Funding
This research was supported by the Project of Guangdong Science and Technology Department (2017A020216008), the Project of Fujian Science and Technology Department (2016I1002 and 2017T3010), the National Natural Science Foundation of China (41506127).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This paper does not contain any studies with human participants or vertebrate animals performed by any of the authors.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sun, F., Wang, C., Chen, L. et al. The intestinal bacterial community of healthy and diseased animals and its association with the aquaculture environment. Appl Microbiol Biotechnol 104, 775–783 (2020). https://doi.org/10.1007/s00253-019-10236-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-10236-z