Abstract
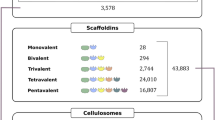
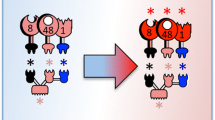
In this study, extended artificial scaffoldins possessing multiple cohesin modules were created in vivo by employing split-intein-mediated protein ligation. Artificial scaffoldins having one Clostridium thermocellum cohesin (Coht), one carbohydrate binding module (CBM) from Clostridium cellulolyticum scaffolding protein CipC, and one to five cohesins (Cohc) derived from CipC, were assembled. These scaffoldins were used to assemble cellulosomal enzyme complexes for investigating the interplay among endoglucanase, exoglucanase, and scaffoldin-borne CBM, on the hydrolysis of a model microcrystalline cellulose substrate, Avicel. The cellulosomal complexes were assembled in vitro by incubating recombinant C. thermocellum endoglucanase (At) and C. cellulolyticum exoglucanase (Ec), with the various artificial scaffoldins. Under a fixed total cellulase concentration, improved hydrolysis is noted by recruiting both Ec and At on the same scaffoldin, for all scaffoldins tested, compared with free cellulases. The improvement is more profound with scaffoldins having a higher Cohc/Coht ratio (i.e., increased Ec/At ratio). Furthermore, among scaffoldins having the same Cohc/Coht ratio, highest rates of Avicel hydrolysis are noted when Coht, and hence an endoglucanase, is situated next to the CBM and not flanked by Cohc. These results point to the importance of using scaffoldins with sufficiently high numbers of cohesin units to achieve an optimal exo-/endo-glucanase ratio to create efficient designer cellulosomes. Furthermore, intein-trans-splicing is proven here to be an effective method for assembling complex scaffoldins and more intricate cellulosomes.







Similar content being viewed by others
References
Aranko AS, Zuger S, Buchinger E, Iwai H (2009) In vivo and in vitro protein ligation by naturally occurring and engineered split DnaE inteins. PLoS One 4(4):e5185. https://doi.org/10.1371/journal.pone.0005185
Bayer EA, Lamed R, Himmel ME (2007) The potential of cellulases and cellulosomes for cellulosic waste management. Curr Opin Biotechnol 18(3):237–245. https://doi.org/10.1016/j.copbio.2007.04.004
Boder ET, Wittrup KD (2000) Yeast surface display for directed evolution of protein expression, affinity, and stability. Methods Enzymol 328:430–444. https://doi.org/10.1016/S0076-6879(00)28410-3
Boisset C, Fraschini C, Schülein M, Henrissat B, Chanzy H (2000) Imaging the enzymatic digestion of bacterial cellulose ribbons reveals the endo character of the cellobiohydrolase Cel6A from Humicola insolens and its mode of synergy with cellobiohydrolase Cel7A. Appl Environ Microbiol 66(4):1444–1452. https://doi.org/10.1128/AEM.66.4.1444-1452.2000
Cha J, Matsuoka S, Chan H, Yukawa H, Inui M, Doi RH (2007) Effect of multiple copies of cohesins on cellulase and hemicellulase activities of Clostridium cellulovorans mini-cellulosomes. J Microbiol Biotechnol 17(11):1782–1788
Chundawat SPS, Paavola CD, Raman B, Nouailler M, Chan SL, Mielenz JR, Receveur-Brechot V, Trent JD, Dale BE (2016) Saccharification of thermochemically pretreated cellulosic biomass using native and engineered cellulosomal enzyme systems. React Chem Eng 1(6):616–628. https://doi.org/10.1039/C6RE00172F
Fan LH, Zhang ZJ, XY Y, Xue YX, Tan TW (2012) Self-surface assembly of cellulosomes with two miniscaffoldins on Saccharomyces cerevisiae for cellulosic ethanol production. Proc Natl Acad Sci U S A 109(33):13260–13265. https://doi.org/10.1073/pnas.1209856109
Fernandes R, Roy V, Wu HC, Bentley WE (2010) Engineered biological nanofactories trigger quorum sensing response in targeted bacteria. Nat Nanotechnol 5(3):213–217. https://doi.org/10.1038/nnano.2009.457
Fierobe HP, Mingardon F, Mechaly A, Belaich A, Rincon MT, Pages S, Lamed R, Tardif C, Belaich JP, Bayer EA (2005) Action of designer cellulosomes on homogeneous versus complex substrates: controlled incorporation of three distinct enzymes into a defined trifunctional scaffoldin. J Biol Chem 280(16):16325–16334. https://doi.org/10.1074/jbc.M414449200
Gaudin C, Belaich A, Champ S, Belaich JP (2000) CelE, a multidomain cellulase from Clostridium cellulolyticum: a key enzyme in the cellulosome? J Bacteriol 182(7):1910–1915. https://doi.org/10.1128/JB.182.7.1910-1915.2000
Goffin P, Dehottay P (2017) Complete genome sequence of Escherichia coli BLR(DE3), a recA-deficient derivative of E. coli BL21(DE3). Genome Announc 5(22):e00441–e00417. https://doi.org/10.1128/genomeA.00441-17
Gräslund S, Nordlund P, Weigelt J, Bray J, Gileadi O, Knapp S (2008) Protein production and purification. Nat Methods 5(2):135–146. https://doi.org/10.1038/nmeth.f.202
Han ZL, Zhang B, Wang YE, Zuo YY, WW S (2012) Self-assembled amyloid-like oligomeric-cohesin scaffoldin for augmented protein display on the Saccharomyces cerevisiae cell surface. Appl Environ Microbiol 78(9):3249–3255. https://doi.org/10.1128/AEM.07745-11
Han Z, Madzak C, WW S (2013) Tunable nano-oleosomes derived from engineered Yarrowia lipolytica. Biotechnol Bioeng 110(3):702–710. https://doi.org/10.1002/bit.24761
Iwai H, Zuger S, Jin J, Tam PH (2006) Highly efficient protein trans-splicing by a naturally split DnaE intein from Nostoc punctiforme. FEBS Lett 580(7):1853–1858. https://doi.org/10.1016/j.febslet.2006.02.045
Jiang W, Boder ET (2010) High-throughput engineering and analysis of peptide binding to class II MHC. Proc Natl Acad Sci U S A 107(30):13258–13263. https://doi.org/10.1073/pnas.1006344107
Krauss J, Zverlov VV, Schwarz WH (2012) In vitro reconstitution of the complete Clostridium thermocellum cellulosome and synergistic activity on crystalline cellulose. Appl Environ Microbiol 78(12):4301–4307. https://doi.org/10.1128/AEM.07959-11
Kruus K, Lua AC, Demain AL, JH W (1995) The anchorage function of CipA (CelL), a scaffolding protein of the Clostridium thermocellum cellulosome. Proc Natl Acad Sci U S A 92(20):9254–9258. https://doi.org/10.1073/pnas.92.20.9254
Kubala MH, Kovtun O, Alexandrov K, Collins BM (2010) Structural and thermodynamic analysis of the GFP:GFP-nanobody complex. Protein Sci 19(12):2389–2401. https://doi.org/10.1002/pro.519
Morais S, Heyman A, Barak Y, Caspi J, Wilson DB, Lamed R, Shoseyov O, Bayer EA (2010) Enhanced cellulose degradation by nano-complexed enzymes: synergism between a scaffold-linked exoglucanase and a free endoglucanase. J Biotechnol 147(3–4):205–211. https://doi.org/10.1016/j.jbiotec.2010.04.012
Mosbah A, Belaı̈ch A, Bornet O, Belaı̈ch J-P, Henrissat B, Darbon H (2000) Solution structure of the module X2_1 of unknown function of the cellulosomal scaffolding protein CipC of Clostridium cellulolyticum. J Mol Biol 304(2):201–217. https://doi.org/10.1006/jmbi.2000.4192
Pages S, Gal L, Belaich A, Gaudin C, Tardif C, Belaich JP (1997) Role of scaffolding protein CipC of Clostridium cellulolyticum in cellulose degradation. J Bacteriol 179(9):2810–2816. https://doi.org/10.1128/jb.179.9.2810-2816.1997
Pakarinen A, Haven MØ, Djajadi DT, Várnai A, Puranen T, Viikari L (2014) Cellulases without carbohydrate-binding modules in high consistency ethanol production process. Biotechnol Biofuels 7(1):27. https://doi.org/10.1186/1754-6834-7-27
Peckham GD, Bugos RC, Su WW (2006) Purification of GFP fusion proteins from transgenic plant cell cultures. Protein Expr Purif 49(2):183–189. https://doi.org/10.1016/j.pep.2006.03.011
Sajjad M, Khan MI, Zafar R, Ahmad S, Niazi UH, Akhtar MW (2012) Influence of positioning of carbohydrate binding module on the activity of endoglucanase CelA of Clostridium thermocellum. J Biotechnol 161(3):206–212. https://doi.org/10.1016/j.jbiotec.2012.05.023
Shi J, Muir TW (2005) Development of a tandem protein trans-splicing system based on native and engineered split inteins. J Am Chem Soc 127(17):6198–6206. https://doi.org/10.1021/ja042287w
Srikrishnan S, Chen W, Da Silva NA (2013) Functional assembly and characterization of a modular xylanosome for hemicellulose hydrolysis in yeast. Biotechnol Bioeng 110(1):275–285. https://doi.org/10.1002/bit.24609
Stern J, Morais S, Lamed R, Bayer EA (2016) Adaptor scaffoldins: an original strategy for extended designer cellulosomes, inspired from nature. MBio 7(2):e00083. https://doi.org/10.1128/mBio.00083-16
Sun Q, Chen W (2016) HaloTag mediated artificial cellulosome assembly on a rolling circle amplification DNA template for efficient cellulose hydrolysis. Chem Commun 52(40):6701–6704. https://doi.org/10.1039/C6CC02035F
Tsai SL, Oh J, Singh S, Chen RZ, Chen W (2009) Functional assembly of minicellulosomes on the Saccharomyces cerevisiae cell surface for cellulose hydrolysis and ethanol production. Appl Environ Microbiol 75(19):6087–6093. https://doi.org/10.1128/AEM.01538-09
Walker JM, Vierstra RD (2007) A ubiquitin-based vector for the co-ordinated synthesis of multiple proteins in plants. Plant Biotechnol J 5(3):413–421. https://doi.org/10.1111/j.1467-7652.2007.00250.x
Wen F, Sun J, Zhao H (2010) Yeast surface display of trifunctional minicellulosomes for simultaneous saccharification and fermentation of cellulose to ethanol. Appl Environ Microbiol 76(4):1251–1260. https://doi.org/10.1128/AEM.01687-09
You C, Zhang XZ, Sathitsuksanoh N, Lynd LR, Zhang YH (2012) Enhanced microbial utilization of recalcitrant cellulose by an ex vivo cellulosome-microbe complex. Appl Environ Microbiol 78(5):1437–1444. https://doi.org/10.1128/AEM.07138-11
Zhang B, Rapolu M, Liang Z, Han Z, Williams PG, WW S (2015) A dual-intein autoprocessing domain that directs synchronized protein co-expression in both prokaryotes and eukaryotes. Sci Rep 5(1):8541. https://doi.org/10.1038/srep08541
Funding
This work was supported in part by the USDA NIFA AFRI grant (2010-65504-20349), NIFA multi-state project HAW05034-R, the Hawaii Community Foundation (44272, 11ADVC-49237), and by a Specific Cooperative Agreement (SCA no.58-5320-3-022) with the Daniel K. Inouye U.S. Pacific Agricultural Research Center.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This work does not contain any study with human participants or animals.
Electronic supplementary material
ESM 1
(PDF 211 kb)
Rights and permissions
About this article
Cite this article
Han, Z., Su, W.W. Intein-mediated assembly of tunable scaffoldins for facile synthesis of designer cellulosomes. Appl Microbiol Biotechnol 102, 1331–1342 (2018). https://doi.org/10.1007/s00253-017-8701-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8701-y