Abstract
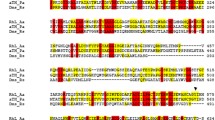
The reductase component (MhpP) of the Sulfobacillus acidophilus TPY multicomponent phenol hydroxylase exhibits only 40 % similarity to Pseudomonas sp. strain CF600 phenol hydroxylase reductase. Amino acid sequence alignment analysis revealed that four cysteine residues (Cys-X 4 -Cys-X 2 -Cys-X 29-35 -Cys) are conserved in the N terminus of MhpP for [2Fe-2S] cluster binding, and two other motifs (RXYS and GXXS/T) are conserved in the C terminus for binding the isoalloxazine and phosphate groups of flavin adenine dinucleotide (FAD). Two motifs (S/T-R and yXCGp) responsible for binding to reduce nicotinamide adenine dinucleotide phosphate (NADPH) are also conserved in MhpP, although some residues differ. To confirm the function of this reductase, MhpP was heterologously expressed in Escherichia coli BL21(DE3) and purified. UV-visible spectroscopy and electron paramagnetic resonance spectroscopy revealed that MhpP contains a [2Fe-2S] cluster. MhpP mutants in which the four cysteine residues were substituted via site-directed mutagenesis lost the ability to bind the [2Fe-2S] cluster, resulting in a decrease in enzyme-specific oxidation of NADPH. Thin-layer chromatography revealed that MhpP contains FAD. Substrate specificity analyses confirmed that MhpP uses NADPH rather than NADH as an electron donor. MhpP oxidizes NADPH using cytochrome c, potassium ferricyanide, or nitro blue tetrazolium as an electron acceptor, with a specific activity of 1.7 ± 0.36, 0.78 ± 0.13, and 0.16 ± 0.06 U/mg, respectively. Thus, S. acidophilus TPY MhpP is a novel NADPH-dependent reductase component of phenol hydroxylase that utilizes FAD and a [2Fe-2S] cluster as cofactors.







Similar content being viewed by others
References
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72(1–2):248–254
Cafaro V, Scognamiglio R, Viggiani A, Izzo V, Passaro I, Notomista E, Piaz FD, Amoresano A, Casbarra A, Pucci P, Di Donato A (2002) Expression and purification of the recombinant subunits of toluene/o-xylene monooxygenase and reconstitution of the active complex. Eur J Biochem 269(22):5689–5699
Ceccarelli EA, Arakaki AK, Cortez N, Carrillo N (2004) Functional plasticity and catalytic efficiency in plant and bacterial ferredoxin-NADP (H) reductases. BBA-Proteins Proteom 1698(2):155–165
Chatwood LL, Muller J, Gross JD, Wagner G, Lippard SJ (2004) NMR structure of the flavin domain from soluble methane monooxygenase reductase from Methylococcus capsulatus (Bath). Biochemistry 43(38):11983–11991
Correll CC, Batie CJ, Ballou DP, Ludwig ML (1992) Phthalate dioxygenase reductase: a modular structure for electron transfer from pyridine nucleotides to [2Fe-2S]. Science 258(5088):1604–1610
Correll CC, Ludwig ML, Bruns CM, Karplus PA (1993) Structural prototypes for an extended family of flavoprotein reductases: comparison of phthalate dioxygenase reductase with ferredoxin reductase and ferredoxin. Protein Sci 2(12):2112–2133
Donadio G, Sarcinelli C, Pizzo E, Notomista E, Pezzella A, Di Cristo C, De Lise F, Di Donato A, Izzo V (2015) The toluene o-xylene monooxygenase enzymatic activity for the biosynthesis of aromatic antioxidants. PLoS One 10(4):e0124427
García LL, Rivas-Marín E, Floriano B, Bernhardt R, Ewen KM, Reyes-Ramírez F, Santero E (2011) ThnY is a ferredoxin reductase-like iron-sulfur flavoprotein that has evolved to function as a regulator of tetralin biodegradation gene expression. J Biol Chem 286(3):1709–1718
Guigliarelli B, Bertrand P (1999) Application of EPR spectroscopy to the structural and functional study of iron-sulfur proteins. Adv Inorg Chem 47(08):421–497
Haigler BE, Gibson DT (1990) Purification and properties of NADH-ferredoxinNAP reductase, a component of naphthalene dioxygenase from Pseudomonas sp. strain NCIB 9816. J Bacteriol 172(1):457–464
Jia Z, Xia H, Yuandong L, Jianshe L, Guanzhou Q (2007) Expression, purification, and characterization of a [Fe2S2] cluster containing ferredoxin from Acidithiobacillus ferrooxidans. Curr Microbiol 55(6):518–523
Karlsson A, Beharry ZM, Eby DM, Coulter ED, Neidle EL, Kurtz DM, Eklund H, Ramaswamy S (2002) X-ray crystal structure of benzoate 1, 2-dioxygenase reductase from Acinetobacter sp. strain ADP1. J Mol Biol 318(2):261–272
Karplus PA, Bruns CM (1994) Structure-function relations for ferredoxin reductase. J Bioenerg Biomembr 26(1):89–99
Karplus PA, Daniels MJ (1991) Atomic structure of ferredoxin-NADP+ reductase: prototype for a structurally novel flavoenzyme family. Science 251(4989):60–66
Kukor JJ, Olsen RH (1992) Complete nucleotide sequence of tbuD, the gene encoding phenol/cresol hydroxylase from Pseudomonas pickettii PKO1, and functional analysis of the encoded enzyme. J Bacteriol 174(20):6518–6526
Laura LG, Francisca RR, Eduardo S (2013) The ferredoxin ThnA3 negatively regulates tetralin biodegradation gene expression via ThnY, a ferredoxin reductase that functions as a regulator of the catabolic pathway. PLoS One 8(9):e73910
Li B, Chen Y, Liu Q, Hu S, Chen X (2011) Complete genome analysis of Sulfobacillus acidophilus strain TPY, isolated from a hydrothermal vent in the Pacific Ocean. J Bacteriol 193(19):5555–5556
Liu Y, Nesheim JC, Paulsen KE, Stankovich MT, Lipscomb JD (1997) Roles of the methane monooxygenase reductase component in the regulation of catalysis. Biochemistry 36(17):5223–5233
Mason JR, Cammack R (1992) The electron-transport proteins of hydroxylating bacterial dioxygenases. Annu Rev Microbiol 46(1):277–305
Merkx M, Kopp DA, Sazinsky MH, Blazyk JL, Mueller J, Lippard SJ (2001) Dioxygen activation and methane hydroxylation by soluble methane monooxygenase: a tale of two irons and three proteins. Angew Chem Int Ed 40(15):2782–2807
Nakajima T, Uchiyama H, Yagi O, Nakahara T (1992) Purification and properties of a soluble methane monooxygenase from Methylocystis sp. M. Biosci Biotechnol Biochem 56(5):736–740
Notomista E, Lahm A, Donato AD, Tramontano A (2003) Evolution of bacterial and archaeal multicomponent monooxygenases. J Mol Evol 56(4):435–445
Pessione E, Divari S, Griva E, Cavaletto M, Rossi GL, Gilardi G, Giunta C (1999) Phenol hydroxylase from Acinetobacter radioresistens is a multicomponent enzyme. Eur J Biochem 265(2):549–555
Powlowski J, Shingler V (1990) In vitro analysis of polypeptide requirements of multicomponent phenol hydroxylase from Pseudomonas sp. strain CF600. J Bacteriol 172(12):6834–6840
Qian H, Edlund UJ, Shingler V, Sethson I (1997) Solution structure of phenol hydroxylase protein component P2 determined by NMR spectroscopy. Biochemistry 36(3):495–504
Sazinsky MH, Lippard SJ (2006) Correlating structure with function in bacterial multicomponent monooxygenases and related diiron proteins. Acc Chem Res 37(45):558–566
Shingler V, Powlowski J, Marklund U (1992) Nucleotide sequence and functional analysis of the complete phenol/3,4-dimethylphenol catabolic pathway of Pseudomonas sp. strain CF600. J Bacteriol 174(3):711–724
Shinohara Y, Uchiyama H, Yagi O, Kusakabe I (1998) Purification and characterization of component B of a soluble methane monooxygenase from Methylocystis sp. M. J Ferment Bioeng 85(1):37–42
Shirabe K, Yubisui T, Nishino T, Takeshita M (1991) Role of cysteine residues in human NADH-cytochrome b5 reductase studied by site-directed mutagenesis. Cys-273 and Cys-283 are located close to the NADH-binding site but are not catalytically essential. J Biol Chem 266(12):7531–7536
Tinberg CE, Woon Ju S, Viviana I, Lippard SJ (2011) Multiple roles of component proteins in bacterial multicomponent monooxygenases: phenol hydroxylase and toluene/o-xylene monooxygenase from Pseudomonas sp. OX1. Biochemistry 50(11):1788–1798
Wang W, Liang AD, Lippard SJ (2015) Coupling oxygen consumption with hydrocarbon oxidation in bacterial multicomponent monooxygenases. Acc Chem Res 48(9):2632–2639
Whittington DA, Lippard SJ (2001) Crystal structures of the soluble methane monooxygenase hydroxylase from Methylococcus capsulatus (Bath) demonstrating geometrical variability at the dinuclear iron active site. J Am Chem Soc 123(5):827–838
Yamaguchi M, Fujisawa H (1978) Characterization of NADH-cytochrome c reductase, a component of benzoate 1, 2-dioxygenase system from Pseudomonas arvilla c-1. J Biol Chem 253(24):8848–8853
Zhou WG, Guo WB, Zhou HB, Chen XH (2016) Phenol degradation by Sulfobacillus acidophilus TPY via the meta-pathway. Microbiol Res 190:37–45
Acknowledgments
The study was supported by grants from the Natural Science Foundation of China (41306166), the National Key Basic Research Program of China (“973”-Program, 2015CB755903), the Natural Science Foundation of Fujian Province, China (2016J05079), the Scientific Research Project of the Marine Public Welfare Industry of China (201205020), and the Xiamen South Ocean Research Center (13GZP002NF08).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
No human participants and/or animals were involved in this work.
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Meng Li and Wenbin Guo contributed equally to this work.
Rights and permissions
About this article
Cite this article
Li, M., Guo, W. & Chen, X. A novel NADPH-dependent reductase of Sulfobacillus acidophilus TPY phenol hydroxylase: expression, characterization, and functional analysis. Appl Microbiol Biotechnol 100, 10417–10428 (2016). https://doi.org/10.1007/s00253-016-7704-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7704-4