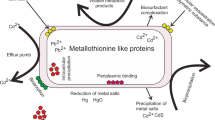
Abstract
Despite current remediation efforts, arsenic contamination in water sources is still a major health problem, highlighting the need for new approaches. In this work, strains of the nonpathogenic and highly arsenic-resistant bacterium Corynebacterium glutamicum were used as inexpensive tools to accumulate inorganic arsenic, either as arsenate (AsV) or arsenite (AsIII) species. The assays made use of “resting cells” from these strains, which were assessed under well-established conditions and compared with C. glutamicum background controls. The two mutant AsV-accumulating strains were those used in a previously published study: (i) ArsC1/C2, in which the gene/s encoding the mycothiol-dependent arsenate reductases is/are disrupted, and (ii) MshA/C mutants unable to produce mycothiol, the low molecular weight thiol essential for arsenate reduction. The AsIII-accumulating strains were either those lacking the arsenite permease activities (Acr3-1 and Acr3-2) needed in AsIII release or recombinant strains overexpressing the aquaglyceroporin genes (glpF) from Corynebacterium diphtheriae or Streptomyces coelicolor, to improve AsIII uptake. Both genetically modified strains accumulated 30-fold more AsV and 15-fold more AsIII than the controls. The arsenic resistance of the modified strains was inversely proportional to their metal accumulation ability. Our results provide the basis for investigations into the use of these modified C. glutamicum strains as a new bio-tool in arsenic remediation efforts.




Similar content being viewed by others
References
Alcántara C, Blasco A, Zúñiga M, Monedero V (2014) Accumulation of Polyphosphate in Lactobacillus spp. and its involvement in stress resistance. Appl Environ Microbiol 80:1650–1659
Argos M, Kalra T, Rathouz PJ, Chen Y, Pierce B, Parvez F, Islam T, Ahmed A, Rakibuz-Zaman M, Hasan R, Sarwar G, Slavkovich V, Van Geen A, Graziano J, Ahsan H (2010) Arsenic exposure from drinking water, and all-cause and chronic-disease mortalities in Bangladesh (HEALS): a prospective cohort study. Lancet 24:252–258
Branco R, Francisco R, Chung AP, Vasconcelos-Morais P (2009) Identification of an Aox system that requires cytochrome c in the highly arsenic-resistant bacterium Ochrobactrum tritici SCII24. Appl Environ Microbiol 75:5141–5147
Chwirka JD, ThompsonIII BM, Stomp JM (2000) Removing arsenic from groundwater. J Am Water Works Assoc 92:79–88
Dhankher OP, Rosen A, McKinney EC, Meagher RB (2006) Hyperaccumulation of arsenic in the shoots of Arabidopsis silenced for arsenate reductase (ACR2). Proc Natl Acad Sci 103:5413–5418
Drewniak L, Sklodowska A (2013) Arsenic-transforming microbes and their role in biomining processes. Environ Sci Pollut Res Int 20:7728–7739
Feng J, Che Y, Milse J, Yin YJ, Liu L, Rückert C, Shen XH, Qi SW, Kalinowski J, Liu SJ (2006) The gene Ncgl2918 encodes a novel maleylpyruvate isomerase that needs mycothiol as cofactor and links mycothiol biosynthesis and gentisate assimilation in Corynebacterium glutamicum. J Biol Chem 281:10778–10785
Feo JC, Ordoñez E, Letek M, Castro MA, Muñoz MI, Gil JA, Mateos LM, Aller AJ (2007) Retention of inorganic arsenic by coryneform mutant strains. Water Res 41:531–542
Kostal J, Yang R, Wu CH, Mulchandani A, Chen W (2004) Enhanced arsenic accumulation in engineered bacterial cells expressing ArsR. Appl Environ Microbiol 70:4582–4587
Kruger MC, Bertin PN, Heipieper HJ, Arsène-Ploetze F (2013) Bacterial metabolism of environmental arsenic mechanisms and biotechnological applications. Appl Microbiol Biotechnol 97:3827–3841
Lindner SN, Vidaurre D, Willbold S, Schoberth SM, Wendisch VF (2007) NCgl2620 encodes a class II polyphosphate kinase in Corynebacterium glutamicum. Appl Environ Microbiol 73:5026–5033
Liu SX, Athar M, Lippai I, Waldren C, Hei TK (2001) Induction of oxyradicals by arsenic: implication for mechanism of genotoxicity. Proc Natl Acad Sci 98:1643–1648
Liu Z, Shen J, Carbrey JM, Mukhopadhyay R, Agre P, Rosen BP (2002) Arsenite transport by mammalian aquaglyceroporins AQP7 and AQP9. Proc Natl Acad Sci 99:6053–6058
Mateos LM, Pisabarro A, Pátek M, Malumbres M, Guerrero C, Eikmanns BJ, Sahm H, Martín JF (1994) Transcriptional analysis and regulatory signals of the hom-thrB cluster of Brevibacterium lactofermentum. J Bacteriol 176:7362-7371
Meiswinkel TM, Rittman D, Lindner SN, Wendisch VF (2013) Crude glycerol-based production of amino acids and putrescine by Corynebacterium glutamicum. Bioresour Technol 145:254–258
Meng YL, Liu Z, Rosen BP (2004) As(III) and Sb(III) uptake by GlpF and efflux by ArsB in Escherichia coli. J Biol Chem 279:18334–18341
Newton GL, Buchmeier N, Fahey RC (2008) Biosynthesis and functions of mycothiol, the unique protective thiol of Actinobacteria. Microbiol Mol Biol Rev 72:471–494
Ordóñez E, Letek M, Valbuena N, Gil JA, Mateos LM (2005) Analysis of genes involved in arsenic resistance in Corynebacterium glutamicum ATCC 13032. Appl Environ Microbiol 71:6206–6215
Ordoñez E, Thiyagarajan S, Cook JD, Stemmler TL, Gil JA, Mateos LM, Rosen BP (2008) Evolution of metal(loid) binding sites in transcriptional regulators. J Biol Chem 283:25706–25714
Ordoñez E, Van Belle K, Roos G, De Galan S, Letek M, Gil JA, Wyns L, Mateos LM, Messens J (2009) Arsenate reductase, mycothiol, and mycoredoxin concert thiol/disulfide exchange. J Biol Chem 284:15107–15116
Pan-Hou H, Kiyono M, Omura T, Endo G (2002) Polyphosphate produced in recombinant Escherichia coli confers mercury resistance. FEMS Microbiol Lett 207:159–64
Sauge-Merle S, Cuiné S, Carrier P, Lecomte-Pradines C, Luu D, Peltier G (2003) Enhanced toxic metal accumulation in engineered bacterial cells expressing Arabidopsis thaliana phytochelatin synthase. Appl Environ Microbiol 69:490–494
Shah D, Shen MW, Chen W, Da Silva NA (2010) Enhanced arsenic accumulation in Saccharomyces cerevisiae overexpressing transporters Fps1p or Hxt7p. J Biotechnol 150:101–107
Shen MW, Shah D, Chen W, Da Silva N (2012) Enhanced arsenate uptake in Saccharomyces cerevisiae overexpressing the Pho84 phosphate transporter. Biotechnol Prog 28:654-661
Singh S, Lee W, Da Silva NA, Mulchandani A, Chen W (2008a) Enhanced arsenic accumulation by engineered yeast cells expressing Arabidopsis thaliana phytochelatin synthase. Biotechnol Bioeng 99:333–340
Singh S, Mulchandani A, Chen W (2008b) Highly selective and rapid arsenic removal by metabolically engineered Escherichia coli cells expressing Fucus vesiculosus metallothionein. Appl Environ Microbiol 74:2924–2927
Singh S, Kang SH, Lee W, Mulchandani A, Chen W (2010) Systematic engineering of phytochelatin synthesis and arsenic transport for enhanced arsenic accumulation in E. coli. Biotechnol Bioeng 105:780–785
Tsai SL, Singh S, Chen W (2009) Arsenic metabolism by microbes in nature and the impact on arsenic remediation. Curr Opin Biotechnol 20:659–667
Tsai SL, Singh S, Da Silva NA, Chen W (2012) Co-expression of Arabidopsis thaliana phytochelatin synthase and Treponema denticola cysteine desulfhydrase for enhanced arsenic accumulation. Biotecnol Bioeng 109:605–608
Villadangos AF, Ordoñez E, Muñoz MI, Pastrana IM, Fiuza M, Gil JA, Mateos LM, Aller AJ (2010) Retention of arsenate using genetically modified coryneform bacteria and determination of arsenic in solid samples by ICP-MS. Talanta 80:1421–1427
Villadangos AF, Van Belle K, Wahni K, Dufe VT, Freitas S, Nur H, De Galan S, Gil JA, Collet JF, Mateos LM, Messens J (2011) Corynebacterium glutamicum survives arsenic stress with arsenate reductases coupled to two distinct redox mechanisms. Mol Microbiol 82:998–1014
Villadangos AF, Fu HL, Gil JA, Messens J, Rosen BP, Mateos LM (2012) Efflux permease CgAcr3-1 of Corynebacterium glutamicum is an arsenite-specific antiporter. J Biol Chem 287:723–735
Wojas S, Clemens S, Skłodowska A (2010) Arsenic response of AtPCS1- and CePCS-expressing plants - effects of external As(V) concentration on As-accumulation pattern and NPT metabolism. J Plant Physiol 167:169–175
Yang J, Salam AA, Rosen BP (2011) Genetic mapping of the interface between the ArsD metallochaperone and the ArsA ATPase. Mol Microbiol 79:872–881
Acknowledgements
AFV and EO were recipients of fellowships from the Junta de Castilla y León (Spain). This work was financially supported by grants from Junta de Castilla y León, Spain (ref. 2014/211330), and the HOA project of the Vrije Universiteit Brussel (Brussels). We thank Professor De Xoyse (H.P.A., London, UK), for providing C. diphtheriae DNA.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 362 kb)
Rights and permissions
About this article
Cite this article
Villadangos, A.F., Ordóñez, E., Pedre, B. et al. Engineered coryneform bacteria as a bio-tool for arsenic remediation. Appl Microbiol Biotechnol 98, 10143–10152 (2014). https://doi.org/10.1007/s00253-014-6055-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-6055-2