Abstract
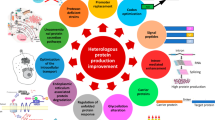
Filamentous fungi, particularly Aspergillus species, have recently attracted attention as host organisms for recombinant protein production. Because the secretory yields of heterologous proteins are generally low compared with those of homologous proteins or proteins from closely related fungal species, several strategies to produce substantial amounts of recombinant proteins have been conducted. Codon optimization is a powerful tool for improving the production levels of heterologous proteins. Although codon optimization is generally believed to improve the translation efficiency of heterologous genes without affecting their mRNA levels, several studies have indicated that codon optimization causes an increase in the steady-state mRNA levels of heterologous genes in filamentous fungi. However, the mechanism that determines the low mRNA levels when native heterologous genes are expressed was poorly understood. We recently showed that the transcripts of heterologous genes are polyadenylated prematurely within the coding region and that the heterologous gene transcripts can be stabilized significantly by codon optimization, which is probably attributable to the prevention of premature polyadenylation in Aspergillus oryzae. In this review, we describe the detailed mechanism of premature polyadenylation and the rapid degradation of mRNA transcripts derived from heterologous genes in filamentous fungi.



Similar content being viewed by others
References
Akashi H (2001) Gene expression and molecular evolution. Curr Opin Genet Dev 11:660–666
Bagar T, Altenbach K, Read ND, Bencina M (2009) Live-cell imaging and measurement of intracellular pH in filamentous fungi using a genetically encoded ratiometric probe. Eukaryot Cell 8:703–712
Beaudoing E, Freier S, Wyatt JR, Claverie JM, Gautheret D (2000) Patterns of variant polyadenylation signal usage in human genes. Genome Res 10:1001–1010
Berg MG, Singh LN, Younis I, Liu Q, Pinto AM, Kaida D, Zhang Z, Cho S, Sherrill-Mix S, Wan L, Dreyfuss G (2012) U1 snRNP determines mRNA length and regulates isoform expression. Cell 150:53–64
Chen W, Xie T, Shao Y, Chen F (2012) Genomic characteristics comparisons of 12 food-related filamentous fungi in tRNA gene set, codon usage and amino acid composition. Gene 497:116–124
Chiu W, Niwa Y, Zeng W, Hirano T, Kobayashi H, Sheen J (1996) Engineered GFP as a vital reporter in plants. Curr Biol 6:325–330
Christensen T, Woeldike H, Boel F, Mortensen SB, Hjortshoej K, Thim L, Hansen M (1988) High level expression of recombinant genes in Aspergillus oryzae. Bio/Technology 6:1419–1422
Cormack BP, Bertram G, Egerton M, Gow NA, Falkow S, Brown AJ (1997) Yeast-enhanced green fluorescent protein (yEGFP): a reporter of gene expression in Candida albicans. Microbiology 143:303–311
De Rocher EJ, Vargo-Gogola TC, Diehn SH, Green PJ (1998) Direct evidence for rapid degradation of Bacillus thuringiensis toxin mRNA as a cause of poor expression in plants. Plant Physiol 117:1445–1461
Diehn SH, Chiu WL, De Rocher EJ, Green PJ (1998) Premature polyadenylation at multiple sites within a Bacillus thuringiensis toxin gene-coding region. Plant Physiol 117:1433–1443
Doma MK, Parker R (2006) Endonucleolytic cleavage of eukaryotic mRNAs with stalls in translation elongation. Nature 440:561–564
Doma MK, Parker R (2007) RNA quality control in eukaryotes. Cell 131:660–668
Dong H, Nilsson L, Kurland CG (1996) Co-variation of tRNA abundance and codon usage in Escherichia coli at different growth rates. J Mol Biol 260:649–663
Drummond DA, Wilke CO (2008) Mistranslation-induced protein misfolding as a dominant constraint on coding-sequence evolution. Cell 134:341–352
Fernández-Abalos JM, Fox H, Pitt C, Wells B, Doonan JH (1998) Plant-adapted green fluorescent protein is a versatile vital reporter for gene expression, protein localization and mitosis in the filamentous fungus, Aspergillus nidulans. Mol Microbiol 27:121–130
Fleissner A, Dersch P (2010) Expression and export: recombinant protein production system for Aspergillus. Appl Microbiol Biotechnol 87:1255–1270
Frischmeyer PA, van Hoof A, O’Donnell K, Guerrerio AL, Parker R, Dietz HC (2002) An mRNA surveillance mechanism that eliminates transcripts lacking termination codon. Science 295:2258–2261
Garneau NL, Wilusz J, Wilusz CJ (2007) The highways and byways of mRNA decay. Nat Rev Mol Cell Biol 8:113–126
Gooch VD, Mehra A, Larrondo LF, Fox J, Touroutoutoudis M, Loros JJ, Dunlap JC (2008) Fully codon-optimized luciferase uncovers novel temperature characteristics of the Neurospora clock. Eukaryot Cell 7:28–37
Gouka RJ, Punt PJ, Hessing JG, van den Hondel CA (1996) Analysis of heterologous protein production in defined recombinant Aspergillus awamori strains. Appl Environ Microbiol 62:1951–1957
Gouka RJ, Punt PJ, van den Hondel CAMJJ (1997) Efficient production of secreted proteins by Aspergillus: progress, limitations and prospects. Appl Microbiol Biotechnol 47:1–11
Graber JH, Cantor CR, Mohr SC, Smith TF (1999a) Genomic detection of new yeast pre-mRNA 3′-end-processing signals. Nucleic Acids Res 27:888–894
Graber JH, Cantor CR, Mohr SC, Smith TF (1999b) In silico detection of control signals: mRNA 3′-end processing sequences in diverse species. Proc Natl Acad Sci U S A 96:14055–14060
Graber JH, McAllister GD, Smith TF (2002) Probabilistic prediction of Saccharomyces cerevisiae mRNA 3′-processing sites. Nucleic Acids Res 30:1851–1858
Graille M, Séraphin B (2012) Surveillance pathways rescuing eukaryotic ribosomes lost in translation. Nat Rev Mol Cell Biol 13:727–735
Grec S, Wang Y, Le Guen L, Negrouk V, Boutry M (2000) Cryptic polyadenylation sites within the coding sequence of three yeast genes expressed in tobacco. Gene 242:87–95
Gustafsson C, Govindarajan S, Minshull J (2004) Codon bias and heterologous protein expression. Trends Biotechnol 22:346–353
Haas J, Park EC, Seed B (1996) Codon usage limitation in the expression of HIV-1 envelope glycoprotein. Curr Biol 6:315–324
Haffani YZ, Overney S, Yelle S, Bellemare G, Belzile FJ (2000) Premature polyadenylation contributes to the poor expression of the Bacillus thuringiensis cry3Ca1 gene in transgenic potato plants. Mol Gen Genet 264:82–88
Hong S-B, Lee M, Kim D-H, Varga J, Frisvad JC, Perrone G, Gomi K, Yamada O, Machida M, Houbraken J, Samson RA (2013) Aspergillus luchuensis, an industrially important black Aspergillus in East Asia. PLoS ONE 8:e63769
Hong S-B, Yamada O, Samson RA (2014) Taxonomic re-evaluation of black koji molds. Appl Microbiol Biotechnol (in press) doi 10.1007/s00253-013-5332-9
Hu J, Lutz CS, Wilusz J, Tian B (2005) Bioinformatic identification of candidate cis-regulatory elements involved in human mRNA polyadenylation. RNA 11:1485–1493
Hung F, Deng L, Ravnikar P, Condon R, Li B, Do L, Saha D, Tsao YS, Merchant A, Liu Z, Shi S (2010) mRNA stability and antibody production in CHO cells: improvement through gene optimization. Biotechnol J 5:393–401
Ikemura T (1981) Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes: a proposal for a synonymous codon choice that is optimal for the E. coli translational system. J Mol Biol 151:389–409
Ikemura T (1985) Codon usage and tRNA content in unicellular and multicellular organisms. Mol Biol Evol 2:13–34
Inada T (2013) Quality control systems for aberrant mRNAs induced by aberrant translation elongation and termination. Biochim Biophys Acta 1829:634–642
Inada T, Aiba H (2005) Translation of aberrant mRNAs lacking a termination codon or with a shortened 3′-UTR is repressed after initiation in yeast. EMBO J 24:1584–1595
Iriarte A, Sanguinetti M, Fernández-Calero T, Naya H, Ramón A, Musto H (2012) Translational selection on codon usage in the genus Aspergillus. Gene 506:98–105
Isken O, Maquat LE (2007) Quality control of eukaryotic mRNA: safeguarding cells from abnormal mRNA function. Genes Dev 21:1833–1856
Ito-Harashima S, Kuroha K, Tatematsu T, Inada T (2007) Translation of the poly(A) tail plays crucial roles in nonstop mRNA surveillance via translation repression and protein destabilization by proteasome in yeast. Genes Dev 21:519–524
Ji G, Zheng J, Shen Y, Wu X, Jiang R, Lin Y, Loke JC, Davis KM, Reese GJ, Li QQ (2007) Predictive modeling of plant messenger RNA polyadenylation sites. BMC Bioinforma 8:43
Kaida D, Berg MG, Younis I, Kasim M, Singh LN, Wan L, Dreyfuss G (2010) U1 snRNP protects pre-mRNAs from premature cleavage and polyadenylation. Nature 468:664–668
Kane JF (1995) Effects of rare codon clusters on high-level expression of heterologous proteins in Escherichia coli. Curr Opin Biotechnol 6:494–500
Kinnaird JH, Burns PA, Fincham JR (1991) An apparent rare-codon effect on the rate of translation of a Neurospora gene. J Mol Biol 221:733–736
Koda A, Bogaki T, Minetoki T, Hirotsune M (2005) High expression of a synthetic gene encoding potato α-glucan phosphorylase in Aspergillus niger. J Biosci Bioeng 100:531–537
Kopke K, Hoff B, Kück U (2010) Application of the Saccharomyces cerevisiae FLP/FRT recombination system in filamentous fungi for marker recycling and construction of knockout strains devoid of heterologous genes. Appl Environ Microbiol 76:4664–4674
Kramer EB, Farabaugh PJ (2007) The frequency of translational misreading errors in E. coli is largely determined by tRNA competition. RNA 13:87–96
Kudla G, Murray AW, Tollervey D, Plotkin JB (2009) Coding-sequence determinants of gene expression in Escherichia coli. Science 324:255–258
Kurland C, Gallant J (1996) Errors of heterologous protein expression. Curr Opin Biotechnol 7:489–493
Leroch M, Mernke D, Koppenhoefer D, Schneider P, Mosbach A, Doehlemann G, Hahn M (2011) Living colors in the gray mold pathogen Botrytis cinerea: codon-optimized genes encoding green fluorescent protein and mCherry, which exhibit bright fluorescence. Appl Environ Microbiol 77:2887–2897
Li XL, Skory CD, Ximenes EA, Jordan DB, Dien BS, Hughes SR, Cotta MA (2007) Expression of an AT-rich xylanase gene from the anaerobic fungus Orpinomyces sp. strain PC-2 in and secretion of the heterologous enzyme by Hypocrea jecorina. Appl Microbiol Biotechnol 74:1264–1275
Lindsley D, Gallant J (1993) On the directional specificity of ribosome frameshifting at a “hungry” codon. Proc Natl Acad Sci U S A 90:5469–5473
Loke JC, Stahlberg EA, Strenski DG, Haas BJ, Wood PC, Li QQ (2005) Compilation of mRNA polyadenylation signals in Arabidopsis revealed a new signal element and potential secondary structures. Plant Physiol 138:1457–1468
Machida M, Yamada O, Gomi K (2008) Genomics of Aspergillus oryzae: learning from the history of koji mold and exploration of its future. DNA Res 15:173–183
Maertens B, Spriestersbach A, von Groll U, Roth U, Kubicek J, Gerrits M, Graf M, Liss M, Daubert D, Wagner R, Schäfer F (2010) Gene optimization mechanisms: a multi-gene study reveals a high success rate of full length human proteins expressed in Escherichia coli. Protein Sci 19:1312–1326
Mandel CR, Bai Y, Tong L (2008) Protein factors in pre-mRNA 3′-end processing. Cell Mol Life Sci 65:1099–1122
Mathew LG, Maloney B, Takeda N, Mason HS (2011) Spurious polyadenylation of Norovirus Narita 104 capsid protein mRNA in transgenic plants. Plant Mol Biol 75:263–275
McNulty DE, Claffee BA, Huddleston MJ, Porter ML, Cavnar KM, Kane JF (2003) Mistranslational errors associated with the rare arginine codon CGG in Escherichia coli. Protein Expr Purif 27:365–374
Millevoi S, Vagner S (2010) Molecular mechanisms of eukaryotic pre-mRNA 3′ end processing regulation. Nucleic Acids Res 38:2757–2774
Moqtaderi Z, Geisberg JV, Jin Y, Fan X, Struhl K (2013) Species-specific factors mediate extensive heterogeneity of mRNA 3′ ends in yeasts. Proc Natl Acad Sci U S A 110:11073–11078
Nelson G, Kozlova-Zwinderman O, Collis AJ, Knight MR, Fincham JR, Stanger CP, Renwick A, Hessing JG, Punt PJ, van den Hondel CA, Read ND (2004) Calcium measurement in living filamentous fungi expressing codon-optimized aequorin. Mol Microbiol 52:1437–1450
Nevalainen KMH, Te’o VSJ, Bergquist PL (2005) Heterologous protein expression in filamentous fungi. Trends Biotechnol 23:468–474
Nguyen KL, Ilano M, Akari H, Miyagi E, Poeschla EM, Strebel K, Bour S (2004) Codon optimization of the HIV-1 vpu and vif genes stabilizes their mRNA and allows for highly efficient Rev-independent expression. Virology 319:163–175
Novoa EM, de Pouplana LR (2012) Speeding with control: codon usage, tRNAs, and ribosomes. Trends Genet 28:574–581
Outchkourov NS, Stiekema WJ, Jongsma MA (2002) Optimization of the expression of equistatin in Pichia pastoris. Protein Expr Purif 24:18–24
Punt PJ, van Biezen N, Conesa A, Albers A, Mangnus J, van den Hondel C (2002) Filamentous fungi as cell factories for heterologous protein production. Trends Biotechnol 20:200–206
Rosenberg AH, Goldman E, Dunn JJ, Studier FW, Zubay G (1993) Effects of consecutive AGG codons on translation in Escherichia coli, demonstrated with a versatile codon test system. J Bacteriol 175:716–722
Sasaguri S, Maruyama J, Moriya S, Kudo T, Kitamoto K, Arioka M (2008) Codon optimization prevents premature polyadenylation of heterologously-expressed cellulases from termite-gut symbionts in Aspergillus oryzae. J Gen Appl Microbiol 54:343–351
Schlackow M, Marguerat S, Proudfoot NJ, Bähler J, Erban R, Gullerova M (2013) Genome-wide analysis of poly(A) site selection in Schizosaccharomyces pombe. RNA 19:1617–1631
Scholtmeijer J, Wösten HAB, Springer J, Wessels JGH (2001) Effect of introns and AT-rich sequences on expression of the bacterial hygromycin B resistance gene in the basidiomycete Schizophyllum commune. Appl Environ Microbiol 67:481–483
Schuren FHJ, Wessels JGH (1998) Expression of heterologous genes in Schizophyllum commune is often hampered by the formation of truncated transcripts. Curr Genet 33:151–156
Shen Y, Ji G, Haas BJ, Wu X, Zheng J, Reese GJ, Li QQ (2008) Genome level analysis of rice mRNA 3′-end processing signals and alternative polyadenylation. Nucleic Acids Res 36:3150–3161
Shoemaker CJ, Green R (2012) Translation drives mRNA quality control. Nat Struct Mol Biol 19:594–601
Spanjaard RA, van Duin J (1988) Translation of the sequence AGG-AGG yields 50 % ribosomal frameshift. Proc Natl Acad Sci U S A 85:7967–7971
Sparks KA, Dieckmann CL (1998) Regulation of poly(A) site choice of several yeast mRNAs. Nucleic Acids Res 26:4676–4687
Takenaka Y, Haga N, Harumoto T, Matsuura T, Mitsui Y (2002) Transformation of Paramecium caudatum with a novel expression vector harboring codon-optimized GFP gene. Gene 284:233–240
Tanaka M, Sakai Y, Yamada O, Shintani T, Gomi K (2011) In silico analysis of 3′-end-processing signals in Aspergillus oryzae using expressed sequence tags and genomic sequencing data. DNA Res 18:189–200
Tanaka M, Tokuoka M, Gomi K (2008) Nonstop mRNA formation by premature polyadenylation within the ORF of heterologous gene in Aspergillus oryzae (in Japanese). Biosci Ind 66:135–139
Tanaka M, Tokuoka M, Shintani T, Gomi K (2012) Transcripts of a heterologous gene encoding mite allergen Der f 7 are stabilized by codon optimization in Aspergillus oryzae. Appl Microbiol Biotechnol 96:1275–1282
Te’o VSJ, Cziferszky AE, Bergquist PL, Nevalainen KM (2000) Codon optimization of xylanase gene xynB from the thermophilic bacterium Dictyoglomus thermophilum for expression in the filamentous fungus Trichoderma reesei. FEMS Microbiol Lett 190:13–19
Tian B, Hu J, Zhang H, Lutz CS (2005) A large-scale analysis of mRNA polyadenylation of human and mouse genes. Nucleic Acids Res 33:201–212
Tokuoka M, Tanaka M, Ono K, Takagi S, Shintani T, Gomi K (2008) Codon optimization increases steady-state mRNA levels in Aspergillus oryzae heterologous gene expression. Appl Environ Microbiol 74:6538–6546
Tsuboi T, Kuroha K, Kudo K, Makino S, Inoue E, Kashima I, Inada T (2012) Dom34:hbs1 plays a general role in quality-control systems by dissociation of a stalled ribosome at the 3′ end of aberrant mRNA. Mol Cell 46:518–529
Tuller T, Waldman YY, Kupie M, Ruppin E (2010) Translation efficiency is determined by both codon bias and folding energy. Proc Natl Acad Sci U S A 107:3645–3650
van Hoof A, Frischmeyer PA, Dietz HC, Parker R (2002) Exosome-mediated recognition and degradation of mRNAs lacking a termination codon. Science 295:2262–2264
Ward OP (2012) Production of recombinant proteins by filamentous fungi. Biotechnol Adv 30:1119–1139
Yang TT, Cheng L, Kain SR (1996) Optimized codon usage and chromophore mutations provide enhanced sensitivity with the green fluorescent protein. Nucleic Acids Res 24:4592–4593
Zhang W, Xiao W, Wei H, Zhang J, Tian Z (2006) mRNA secondary structure at start AUG codon is a key limiting factor for human protein expression in Escherichia coli. Biochem Biophys Res Commun 349:69–78
Zhao J, Hyman L, Moore C (1999) Formation of mRNA 3′ ends in eukaryotes: mechanism, regulation, and interrelationships with other steps in mRNA synthesis. Microbiol Mol Biol Rev 63:405–445
Zolotukhin S, Potter M, Hauswirth WW, Guy J, Muzyczka N (1996) A “humanized” green fluorescent protein cDNA adapted for high-level expression in mammalian cells. J Virol 70:4646–4654
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Tanaka, M., Tokuoka, M. & Gomi, K. Effects of codon optimization on the mRNA levels of heterologous genes in filamentous fungi. Appl Microbiol Biotechnol 98, 3859–3867 (2014). https://doi.org/10.1007/s00253-014-5609-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-5609-7