Abstract
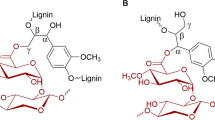
Glucuronoyl esterases (GEs) are recently discovered enzymes that are suggested to cleave the ester bond between lignin alcohols and xylan-bound 4-O-methyl-d-glucuronic acid. Although their potential use for enhanced enzymatic biomass degradation and synthesis of valuable chemicals renders them attractive research targets for biotechnological applications, the difficulty to purify natural fractions of lignin-carbohydrate complexes hampers the characterization of fungal GEs. In this work, we report the synthesis of three aryl alkyl or alkenyl d-glucuronate esters using lipase B from Candida antarctica (CALB) and their use to determine the kinetic parameters of two GEs, StGE2 from the thermophilic fungus Myceliophthora thermophila (syn. Sporotrichum thermophile) and PaGE1 from the coprophilous fungus Podospora anserina. PaGE1 was functionally expressed in the methylotrophic yeast Pichia pastoris under the transcriptional control of the alcohol oxidase (AOX1) promoter and purified to its homogeneity (63 kDa). The three d-glucuronate esters contain an aromatic UV-absorbing phenol group that facilitates the quantification of their enzymatic hydrolysis by HPLC. Both enzymes were able to hydrolyze the synthetic esters with a pronounced preference towards the cinnamyl-d-glucuronate ester. The experimental results were corroborated by computational docking of the synthesized substrate analogues. We show that the nature of the alcohol portion of the hydrolyzed ester influences the catalytic efficiency of the two GEs.





Similar content being viewed by others
References
Aita GA, Salvi DA, Walker MS (2011) Enzyme hydrolysis and ethanol fermentation of dilute ammonia pretreated energy cane. Bioresour Technol 102(6):4444–4448
Balakshin M, Capanema E, Gracz H, Chang HM, Jameel H (2011) Quantification of lignin-carbohydrate linkages with high-resolution NMR spectroscopy. Planta 233(6):1097–1110
Bey M, Berrin J-G, Poidevin L, Sigoillot J-C (2011) Heterologous expression of Pycnoporus cinnabarinus cellobiose dehydrogenase in Pichia pastoris and involvement in saccharification processes. Microb Cell Fact 10:113
Blom N, Sicheritz-Ponten T, Gupta R, Gammeltoft S, Brunak S (2004) Prediction of post-translational glycosylation and phosphorylation of proteins from the amino acid sequence. Proteomics 4:1633–1649
Bounias M (1980) N-(1-naphthyl)ethylenediamine dihydrochloride as a new reagent for nanomole quantification of sugars on thin-layer plates by a mathematical calibration process. Anal Biochem 106(2):291–295
Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, Henrissat B (2009) The carbohydrate-active enzymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res 37:D233–D238
Charavgi MD, Dimarogona M, Topakas E, Christakopoulos P, Chrysina ED (2013) The structure of a novel glucuronoyl esterase from Myceliophthora thermophila gives new insights into its role as a potential biocatalyst. Acta Crystallogr D Biol Crystallogr 69(Pt 1):63–73
Choi JW, Choi D-H, Faix O (2007) Characterization of lignin–carbohydrate linkages in the residual lignins isolated from chemical pulps of spruce (Picea abies) and beech wood (Fagus sylvatica). J Wood Sci 53:309–313
de Graaf M, Nevalainen TJ, Scheeren HW, Pinedo HM, Haisma HJ, Boven E (2004) A methylester of the glucuronide prodrug DOX-GA3 for improvement of tumor-selective chemotherapy. Biochem Pharmacol 68(11):2273–2281
Ďuranová M, Hirsch J, Kolenová K, Biely P (2009a) Fungal glucuronoyl esterases and substrate uronic acid recognition. Biosci Biotechnol Biochem 73:2483–2487
Ďuranová M, Spanikova S, Wosten HAB, Biely P, de Vries RP (2009b) Two glucuronoyl esterases of Phanerochaete chrysosporium. Arch Microbiol 191:133–140
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In: Walker JM (ed) The proteomics protocols handbook. Humana Press, New Jersey, pp 571–607
Gibson LJ (2012) The hierarchical structure and mechanics of plant materials. J R Soc Interface 9(76):2749–2766
Hirsch J, Kooš M, Kováč P (1998) Improved synthesis of an aldobiouronic acid related to hardwood xylans, and preparation of a derivative thereof suitable for linking to proteins. Carbohydr Res 310:145–149
Hori C, Gaskell J, Igarashi K, Samejima M, Hibbett DS, Henrissat B, Cullen D (2013) Genomewide analysis of polysaccharides degrading enzymes in 11 white- and brown-rot Polyporales provides insight into mechanisms of wood decay. Mycologia 105(6):1412–1427
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(1):33–38, 27-28
Karnaouri AC, Topakas E, Christakopoulos P (2014) Cloning, expression, and characterization of a thermostable GH7 endoglucanase from Myceliophthora thermophile capable of high-consistency enzymatic liquefaction. Appl Microbiol Biotechnol 98(1):231–242
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Li X-L, Špániková S, de Vries RP, Biely P (2007) Identification of genes encoding microbial glucuronoyl esterases. FEBS Lett 581:4029–4035
Moreau B, Lognay GC, Blecker C, Brohee JC, Chery F, Rollin P, Paquot M, Marlier M (2004) Synthesis of novel d-glucuronic acid fatty esters using Candida antarctica lipase in tert-butanol. Biotechnol Lett 26(5):419–424
Otto RT, Bornscheuer UT, Scheib H, Pleiss J, Syldatk C, Schmid RD (1998) Lipase-catalyzed esterification of unusual substrates: synthesis of glucuronic acid and ascorbic acid (vitamin C) esters. Biotechnol Lett 20(11):1091–1094
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786
Pokkuluri PR, Duke NE, Wood SJ, Cotta MA, Li XL, Biely P, Schiffer M (2011) Structure of the catalytic domain of glucuronoyl esterase Cip2 from Hypocrea jecorina. Proteins 79(8):2588–2592
Ralph J, Hatfield RD, Piquemal J, Yahiaoui N, Pean M, Lapierre C, Boudet AM (1998) NMR characterization of altered lignins extracted from tobacco plants down-regulated for lignification enzymes cinnamylalcohol dehydrogenase and cinnamoyl-CoA reductase. PNAS 95(22):12803–12808
Sarkanen KV (1971) In: Sarkanen KV, Ludwig CH (eds) Lignins: occurrence, formation, structure and reactions. Wiley-Interscience, New York, pp 95–163
Špániková S, Biely P (2006) Glucuronoyl esterase–novel carbohydrate esterase produced by Schizophyllum commune. FEBS Lett 580(19):4597–4601
Špániková S, Polakova M, Joniak D, Hirsch J, Biely P (2007) Synthetic esters recognized by glucuronoyl esterase from Schizophyllum commune. Arch Microbiol 188(2):185–189
Steentoft C, Vakhrushev SY, Joshi HJ, Kong Y, Vester-Christensen MB, Schjoldager KT, Lavrsen K, Dabelsteen S, Pedersen NB, Marcos-Silva L, Gupta R, Bennett EP, Mandel U, Brunak S, Wandall HH, Levery SB, Clausen H (2013) Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology. EMBO J 32(10):1478–1488
Stoscheck CM (1990) Quantification of protein. Methods Enzymol 182:50–68
Topakas E, Moukouli M, Dimarogona M, Vafiadi C, Christakopoulos P (2010) Functional expression of a thermophilic glucuronyl esterase from Sporotrichum thermophile: identification of the nucleophilic serine. Appl Microbiol Biotechnol 87(5):1765–1772
Tsai AYL, Canam T, Gorzsas A, Mellerowicz EJ, Campbell MM, Master ER (2012) Constitutive expression of a fungal glucuronoyl esterase in Arabidopsis reveals altered cell wall composition and structure. Plant Biotechnol J 10:1077–1087
Vafiadi C, Topakas E, Biely P, Christakopoulos P (2009) Purification, characterization and mass spectrometric sequencing of a thermophilic glucuronoyl esterase from Sporotrichum thermophile. FEMS Microbiol Lett 296:178–184
Wong DWS, Chan VJ, McCormack AA, Hirsch J, Biely P (2012) Functional cloning and expression of the Schizophyllum commune glucuronoyl esterase gene and characterization of the recombinant enzyme. Biotechnol Res Inter vol. 2012, article ID 951267, doi:10.1155/2012/951267
Wymelenberg AV, Gaskell J, Mozuch M, Sabat G, Ralph J, Skyba O, Mansfield SD, Blanchette RA, Martinez D, Grigoriev I, Kersten PJ, Cullen D (2010) Comparative transcriptome and secretome analysis of wood decay fungi Postia placenta and Phanerochaete chrysosporium. Appl Environ Microbiol 76:3599–3610
Acknowledgments
The financial support of General Secretariat of Research and Technology (GSRT) of Greece-ESPA 2007–2013 is gratefully acknowledged. We are also grateful to Novo Nordisk A/S for the generous gift of C. antarctica lipase CALB and to Marina Roussaki for her expertise in obtaining the NMR spectra.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 347 kb)
Rights and permissions
About this article
Cite this article
Katsimpouras, C., Bénarouche, A., Navarro, D. et al. Enzymatic synthesis of model substrates recognized by glucuronoyl esterases from Podospora anserina and Myceliophthora thermophila . Appl Microbiol Biotechnol 98, 5507–5516 (2014). https://doi.org/10.1007/s00253-014-5542-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-5542-9