Abstract
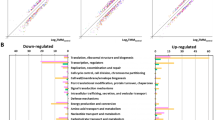
Bacteria belonging to the Staphylococcus genus reside in various natural environments; however, only disease-associated Staphylococcus strains have received attention while ecological function and physiologies of non-pathogenic strains were often neglected. Because high level of tolerance against NaCl is a common trait of Staphylococcus, we investigated the characteristics of halotolerance in Staphylococcus sp. OJ82 isolated from fermented seafood containing a high concentration of NaCl. Among the 292 isolates screened, OJ82 showed the highest β-galactosidase and extracellular protease activities under high-salt conditions. Comparative genomic analysis with other Staphylococcus strains showed that (a) replication origins are highly conserved, (b) the OJ82 strain has a high number of amino acid transport- and metabolism-related genes, and (c) OJ82 has many unique proteins (15 %) and 12 prophage-related genomic islands. RNA-seq analysis under high-salt conditions showed that genes involved in cell membranes, transport, osmotic stress, ATP synthesis, and translation are highly expressed. OJ82 may use the ribulose monophosphate pathway to detoxify some toxic intermediates under high-salt conditions. Six new and three known non-coding small RNAs of the OJ82 strain were also found in the RNA-seq analysis. Genomic and transcriptomic analyses identified target β-galactosidase and extracellular protease. Interestingly, the OJ82 strain became resistant to bacteriocin produced by the Bacillus strain only under high-salt conditions. Our data showed that the OJ82 strain adapted to high-salt conditions by expressing core cellular processes (translation, ATP production) and defense genes (membrane synthesis, compatible solute transports, ribulose monophosphate pathway) could survive bacteriocin exposure under high-salt conditions.






Similar content being viewed by others
References
Bonis M, Ecobichon C, Guadagnini S, Prévost MC, Boneca IG (2010) A M23B family metallopeptidase of Helicobacter pylori required for cell shape, pole formation and virulence. Mol Microbiol 78:809–819
Carver TJ, Rutherford KM, Berriman M, Rajandream MA, Barrell BG, Parkhill J (2005) ACT: the Artemis Comparison Tool. Bioinformatics 21:3422–3423
Castro SL, Nelman-Gonzalez M, Nickerson CA, Ott CM (2011) Induction of attachment-independent biofilm formation and repression of hfq expression by low-fluid-shear culture of Staphylococcus aureus. Appl Environ Microbiol 77:6368–6378
Chan CY, Carmack CS, Long DD, Maliyekkel A, Shao Y, Roninson IB, Ding Y (2009) A structural interpretation of the effect of GC-content on efficiency of RNA interference. BMC Bioinforma 30:10
Corbière Morot-Bizot S, Leroy S, Talon R (2006) Staphylococcal community of a small unit manufacturing traditional dry fermented sausages. Int J Food Microbiol 108:210–217
Dartigalongue C, Raina S (1998) A new heat-shock gene, ppiD, encodes a peptidyl-prolyl isomerase required for folding of outer membrane proteins in Escherichia coli. EMBO J 17:3968–3980
Delaune A, Poupel O, Mallet A, Coic YM, Msadek T, Dubrac S (2011) Peptidoglycan crosslinking relaxation plays an important role in Staphylococcus aureus WalKR-dependent cell viability. PLoS ONE 28:17054
Diep DB, Nes IF (2002) Ribosomally synthesized antibacterial peptides in gram positive bacteria. Curr Drug Targets 3:107–122
Eijsink VG, Axelsson L, Diep DB, Havarstein LS, Holo H, Nes IF (2002) Production of class II bacteriocins by lactic acid bacteria; an example of biological warfare and communication. Antonie Van Leeuwenhoek 81:639–654
El Farran CA, Sekar A, Balakrishnan A, Shanmugam S, Arumugam P, Gopalswamy J (2013) Prevalence of biofilm-producing Staphylococcus epidermidis in the healthy skin of individuals in Tamil Nadu, India. Indian J Med Microbiol 31:19–23
Epstein W (1986) Osmoregulation by potassium transport in Escherichia coli. FEMS Microbiol Rev 39:73–78
Fredericq P (1957) Colicins. Ann Rev Microbiol 11:7–22
Geissmann T, Chevalier C, Cros MJ, Boisset S, Fechter P, Noirot C, Schrenzel J, François P, Vandenesch F, Gaspin C, Romby P (2009) A search for small noncoding RNAs in Staphylococcus aureus reveals a conserved sequence motif for regulation. Nucleic Acids Res 37:7239–7257
Göthel SF, Scholz C, Schmid FX, Marahiel MA (1998) Cyclophilin and trigger factor from Bacillus subtilis catalyze in vitro protein folding and are necessary for viability under starvation conditions. Biochemistry 37:13392–13399
Gottesman S (2005) Micros for microbes: non-coding regulatory RNAs in bacteria. Trends Genet 21:399–404
Guan L, Cho KH, Lee JH (2011) Analysis of the cultivable bacterial community in jeotgal, a Korean salted and fermented seafood, and identification of its dominant bacteria. Food Microbiol 28:101–113
Gunn BA, Colwell RR (1983) Numerical taxonomy of Staphylococci isolated from the marine environment. Int J Syst Evol Microbiol 33:751–759
Hahne H, Mäder U, Otto A, Bonn F, Steil L, Bremer E, Hecker M, Becher D (2010) A comprehensive proteomics and transcriptomics analysis of Bacillus subtilis salt stress adaptation. J Bacteriol 192:870–882
Hájek V, Ludwig W, Schleifer KH, Springer N, Zitzelsberger W, Kroppenstedt RM, Kocur M (1992) Staphylococcus muscae, a new species isolated from flies. Int J Syst Evol Microbiol 42:97–101
Hempel RJ, Morton DJ, Seale TW, Whitby PW, Stull TL (2013) The role of the RNA chaperone Hfq in Haemophilus influenzae pathogenesis. BMC Microbiol 13:134
Holtmann G, Bakker EP, Uozumi N, Bremer E (2003) KtrAB and KtrCD: two K+ uptake systems in Bacillus subtilis and their role in adaptation to hypertonicity. J Bacteriol 185:1289–1298
Ito M, Guffanti AA, Oudega B, Krulwich TA (1999) mrp, a multigene, multifunctional locus in Bacillus subtilis with roles in resistance to cholate and to Na+ and in pH homeostasis. J Bacteriol 181:2394–2402
Jung J, Park W (2013) Comparative genomic and transcriptomic analysis reveal habitat differentiation and different transcriptional responses during pectin metabolism in Alishewanella species. Appl Environ Microbiol 79:6351–6361
Jung J, Chun J, Park W (2012) Genome sequence of extracellular-protease-producing Alishewanella jeotgali isolated from traditional Korean fermented seafood. J Bacteriol 194:2097
Kandror O, Sherman M, Rhode M, Goldberg AL (1995) Trigger factor is involved in GroEL-dependent protein degradation in Escherichia coli and promotes binding of GroEL to unfolded proteins. EMBO J 14:6021–6027
Kato N, Yurimoto H, Thauer RK (2006) The physiological role of the ribulose monophosphate pathway in bacteria and archaea. Biosci Biotech Biochem 70:10–21
Lambert LH, Cox T, Mitchell K, Rosselló-Mora RA, Del Cueto C, Dodge DE, Orkand P, Cano RJ (1998) Staphylococcus succinus sp. nov., isolated from Dominican amber. Int J Syst Evol Microbiol 48:511–518
Lambert ML, Suetens C, Savey A, Palomar M, Hiesmayr M, Morales I, Agodi A, Frank U, Mertens K, Schumacher M, Wolkewitz M (2011) Clinical outcomes of health-care-associated infections and antimicrobial resistance in patients admitted to European intensive-care units: a cohort study. Lancet Infect Dis 11:30–38
Leroy S, Giammarinaro P, Chacornac JP, Lebert I, Talon R (2010) Biodiversity of indigenous staphylococci of naturally fermented dry sausages and manufacturing environments of small-scale processing units. Food Microbiol 27:294–301
MacLean MJ, Ness LS, Ferguson GP, Booth IR (1998) The role of glyoxalase I in the detoxification of methylglyoxal and in the activation of the KefB K+ efflux system in Escherichia coli. Mol Microbiol 27:563–571
Miller JH (1992) A short course in bacterial genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Miller HK, Carroll RK, Burda WN, Krute CN, Davenport JE, Shaw LN (2012) The extracytoplasmic function sigma factor σS protects against both intracellular and extracytoplasmic stresses in Staphylococcus aureus. J Bacteriol 194:4342–4354
Nam YD, Lee SY, Lim SI (2012) Microbial community analysis of Korean soybean pastes by next-generation sequencing. Int J Food Microbiol 155:36–42
Nguyenet TT, Eiamphungporn W, Mäder U, Liebeke M, Lalk M, Hecker M, Helmann JD, Antelmann H (2009) Genome-wide responses to carbonyl electrophiles in Bacillus subtilis: control of the thiol-dependent formaldehyde dehydrogenase AdhA and cysteine proteinase YraA by the MerR-family regulator YraB (AdhR). Mol Microbiol 71:876–894
Padan E, Maisler N, Taglicht D, Karpel R, Schuldiner S (1989) Deletion of ant in Escherichia coli reveals its function in adaptation to high salinity and an alternative Na+/H+ antiporter systems(s). J Biol Chem 264:20297–20302
Pane-Farre J, Jonas B, Förstner K, Engelmann S, Hecker M (2006) The σB regulon in Staphylococcus aureus and its regulation. Int J Med Microbiol 296:237–258
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786
Pomeranz Y (1964) Lactase occurrence and properties. Food Tech 88:682
Probst AJ, Hertel C, Richter L, Wassill L, Ludwig W, Hammes WP (1998) Staphylococcus condimenti sp. nov., from soy sauce mash, and Staphylococcus carnosus subsp. utilis subsp. nov. Int J Syst Evol Microbiol 48:651–658
Pruitt KD, Tatusova T, Klimke W, Maglott DR (2009) NCBI reference sequences: current status, policy and new initiatives. Nucleic Acids Res 37:32–36
Ringus DL, Ivy RA, Wiedmann M, Boor KJ (2012) Salt stress-induced transcription of σB- and CtsR-regulated genes in persistent and non-persistent Listeria monocytogenes strains from food processing plants. Foodborne Pathog Dis 9:198–206
Schleifer KH, Kloos WE (1975) Isolation and characterization of staphylococci from human skin I. Amended description of Staphylococcus epidermidis and Staphylococcus saprophyticus and descriptions of three new species: Staphylococcus cohnii, Staphylococcus haemolyticus, and Staphylococcus xylous. Int J Syst Evol Microbiol 25:50–61
Schwan WR, Lehmann L, McCormick J (2006) Transcriptional activation of the Staphylococcus aureus putP gene by low-proline-high osmotic conditions and during infection of murine and human tissues. Infect Immun 74:399–409
Sklar JG, Wu T, Kahne D, Silhavy TJ (2007) Defining the roles of the periplasmic chaperones SurA, Skp, and DegP in Escherichia coli. Genes Dev 21:2473–2484
Sleator RD, Hill C (2001) Bacterial osmoadaptation: the role of osmolytes in bacterial stress and virulence. FEMS Microbiol Rev 26:49–71
Stapleton MR, Horsburgh MJ, Hayhurst EJ, Wright L, Jonsson IM, Tarkowski A, Kokai-Kun JF, Mond JJ, Foster SJ (2007) Characterization of IsaA and SceD, two putative lytic transglycosylases of Staphylococcus aureus. J Bacteriol 189:7316–7325
Stoller G, Rücknagel KP, Nierhaus KH, Schmid FX, Fischer G, Rahfeld JU (1995) A ribosome-associated peptidyl-prolyl cis/trans isomerase identified as the trigger factor. EMBO J 14:4939–4948
Sung J, Chun J, Choi S, Park W (2012) Genome sequence of the halotolerant Staphylococcus sp. strain OJ82, isolated from Korean traditional salt-fermented seafood. J Bacteriol 194:6353–6354
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tanasupawat S, Hashimoto Y, Ezaki T, Kozaki M, Komagata K (1992) Staphylococcus piscifermentans sp. nov., from fermented fish in Thailand. Int J Syst Evol Microbiol 42:577–581
Tang T, Wu C, Li J, Ren G, Huang D, Liu F (2012) Stress-induced HSP70 from Musca domestica plays a functionally significant role in the immune system. J Insect Physiol 58:1226–1234
Taponen S, Supré K, Piessens V, Van Coillie E, De Vliegher S, Koort JM (2012) Staphylococcus agnetis sp. nov., a coagulase-variable species from bovine subclinical and mild clinical mastitis. Int J Syst Evol Microbiol 62:61–65
Teather RM, Wood PJ (1982) Use of Congo red–polysaccharide interactions in enumeration and characterization of cellulolytic bacteria from the bovine rumen. Appl Environ Microbiol 43:777–780
Toledo-Arana A, Repoila F, Cossart P (2007) Small noncoding RNAs controlling pathogenesis. Curr Opin Microbiol 10:182–188
Valentin-Hansen P, Eriksen M, Udesen C (2004) The bacterial Sm-like protein Hfq: a key player in RNA transactions. Mol Microbiol 51:1525–1533
van Schaik W, Abee T (2005) The role of σB in the stress response of Gram-positive bacteria—targets for food preservation and safety. Curr Opin Biotechnol 16:218–224
Vernozy-Rozand C, Mazuy C, Meugnier H, Bes M, Lasne Y, Fiedler F, Etienne J, Freney J (2000) Staphylococcus fleurettii sp. nov., isolated from goat's milk cheeses. Int J Syst Evol Microbiol 50:1521–1527
Waters LS, Storz G (2009) Regulatory RNAs in bacteria. Cell 136:615–628
Whatmore AM, Chudek JA, Reed RH (1990) The effects of osmotic upshock on the intracellular solute pools of Bacillus subtilis. J Gen Microbiol 136:2527–2535
Yeom J, Imlay JA, Park W (2010) Iron homeostasis affects antibiotic-mediated cell death in Pseudomonas species. J Biol Chem 285:22689–22695
Zhang Z, Meng Q, Qiao J, Yang L, Cai X, Wang G, Chen C, Zhang L (2012) RsbV of Listeria monocytogenes contributes to regulation of environmental stress and virulence. Arch Microbiol 195:113–120
Zuleta LF, Italiani VC, Marques MV (2003) Isolation and characterization of NaCl-sensitive mutants of Caulobacter crescentus. Appl Environ Microbiol 69:3029–3035
Acknowledgments
This study was supported by a grant from the Next-Generation BioGreen 21 Program (PJ0082082013), Rural Development Administration, Korea.
Author information
Authors and Affiliations
Corresponding author
Additional information
Sungjong Choi and Jaejoon Jung contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 1753 kb)
Rights and permissions
About this article
Cite this article
Choi, S., Jung, J., Jeon, C.O. et al. Comparative genomic and transcriptomic analyses of NaCl-tolerant Staphylococcus sp. OJ82 isolated from fermented seafood. Appl Microbiol Biotechnol 98, 807–822 (2014). https://doi.org/10.1007/s00253-013-5436-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-013-5436-2