Abstract
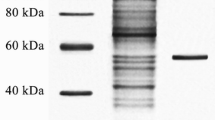
Acinetobacter sp. XMZ-26 (ACCC 05422) was isolated from soil samples obtained from glaciers in Xinjiang Province, China. The partial nucleotide sequence of a lipase gene was obtained by touchdown PCR using degenerate primers designed based on the conserved domains of cold-adapted lipases. Subsequently, a complete gene sequence encoding a 317 amino acid polypeptide was identified. Our novel lipase gene, lipA, was overexpressed in Escherichia coli. The recombinant protein (LipA) was purified by Ni-affinity chromatography, and then deeply characterised. The LipA resulted to hydrolyse pNP esters of fatty acids with acyl chain length from C2 to C16, and the preferred substrate was pNP octanoate showing a kcat = 560.52 ± 28.32 s−1, Km = 0.075 ± 0.008 mM, and a kcat/Km = 7,377.29 ± 118.88 s−1 mM−1. Maximal LipA activity was observed at a temperature of 15°C and pH 10.0 using pNP decanoate as substrate. That LipA peaked at such a low temperature and remained most activity between 5°C and 35°C indicated that it was a cold-adapted enzyme. Remarkably, this lipase retained much of its activity in the presence of commercial detergents and organic solvents, including Ninol, Triton X-100, methanol, PEG-600, and DMSO. This cold-adapted lipase may find applications in the detergent industry and organic synthesis.





Similar content being viewed by others
References
Alquati C, Gioia L, Santarossa G, Alberghina L, Fantucci P, Lotti M (2002) The cold-adapted lipase of Pseudomonas fragi: heterologous expression, biochemical characterization and molecular modeling. Eur J Biochem 269:3321–3328
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Breuil C, Kushner DJ (1975) Partial purification and characterization of the lipase of a facultatively psychrophilic bacterium (Acinetobacter O16). Can J Microbiol 21:434–441
Carriere F, Thirstrup K, Hjorth S, Boel E (1994) Cloning of the classical guinea pig pancreatic lipase and comparison with the lipase related protein. FEBS Lett 338:63–68
Choo DW, Kurihara T, Suzuki T, Soda K, Esaki N (1998) A cold-adapted lipase on an Alaskan psychrotroph, Pseudomonas sp. strain B11-1: gene cloning and enzyme purification and characterization. Appl Environ Microbiol 64:486–491
De Santi C, Tutino ML, Mandrich L, Giuliani M, Parrilli E, Del Vecchio P, de Pascale D (2010) The hormone-sensitive lipase from Psychrobacter sp. TA144: new insight in the structural/functional characterization. Biochimie 92:949–957
Dieckelmann M, Johnson LA, Beacham IR (1998) The diversity of lipases from psychrotrophic strains of Pseudomonas: a novel lipase from a highly lipolytic strain of Pseudomonas fluorescens. J Appl Microbiol 85:527–536
Feller G, Thiry M, Arpigny JL, Gerday C (1991) Cloning and expression in Escherichia coli of three lipase-coding genes from psychrotrophic Antarctic strain Moraxella TA144. Gene 102:111–115
Grochulski P, Li Y, Schrag JD, Bouthillier F, Smith P, Harrison D, Rubin B, Cygler M (1993) Insights into interfacial activation from an open structure of Candida rugosa lipase. J Biol Chem 268:12843–12847
Han SJ, Bach JH, Yoon MY, Shin PK, Cheong CS, Sung MH, Hong SP, Chung IY, Han YS (2003) Expression and characterization of a novel enantioselective lipase from Acinetobacter species SY-01. Biochimie 85:501–510
Hemilä H, Koivula TT, Palva I (1994) Hormone-sensitive lipase is closely related to several bacterial proteins, and distantly related to acetylcholine esterase and lipoprotein lipase: identification of a superfamily of esterases and lipases. Biochim Biophys Acta 1210:249–253
Hong M, Chang M (1988) Purification and characterization of an alkaline lipase from a newly isolated Acinetobacter radioresistens CMC-1. Biotechnol Lett 20:1027–1029
Ito S, Kobayashi T, Ara K, Ozaki K, Kawai S, Hatada Y (1988) Alkaline detergent enzymes from alkalophiles: enzymatic properties, genetics and structures. Extremophiles 2:185–190
Jaeger KE, Dijkstra BW, Reetz MT (1999) Bacterial biocatalysts: molecular biology, three-dimensional structures, and biotechnological applications of lipases. Annu Rev Microbiol 53:315–351
Jahandideh S, Abdolmaleki P, Jahandideh M, Barzegari Asadabadi E (2007) Sequence and structural parameters enhancing adaptation of proteins to low temperatures. J Theor Biol 246:159–166
Joseph B, Ramteke PW, Thomas G (2008) Cold adapted microbial lipases: some hot issues and recent developments. Biotechnol Adv 26:457–470
Kademi A, Fakhreddine L, Baratti J (2000) Purification and characterization of a thermostable esterase from the moderate thermophile Bacillus circulans. Appl Microbiol Biotechnol 54:173–179
Kim SH, Park IH, Lee SC, Lee YS, Zhou-Yi KCM, Ahn SC, Choi YL (2008) Discovery of three novel lipase (lipA1, lipA2, and lipA3) and lipase-specific chaperone (lipB) genes present in Acinetobacter sp. DYL129. Appl Microbiol Biotechnol 77:1041–1051
Kok RG, Thor JJ, Nugteren-Roodzant IM, Brouwer MB, Egmond MR, Nudel CB, Vosman B, Hellingwerf KJ (1995) Characterization of the extracellular lipase, LipA, of Acinetobacter calcoaceticus BD413 and sequence analysis of the cloned structural gene. Mol Microbiol 15:803–818
Kulakova L, Galkin A, Nakayama T, Nishino T, Esaki N (2004) Cold-adapted esterase from Psychrobacter sp. Ant300: gene cloning, characterization, and the effects of Gly– > Pro substitution near the active site on its catalytic activity and stability. Biochim Biophys Acta 1696:59–65
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Loreto PP, Fernando R, Juan P, Oriana S, Barbara AA, Juan AA (2008) Cloning and fusion expression of a cold-adapted lipase from marine Antarctic origin. Enzyme Microb Technol 42:371–377
Maia MM, Heasley A, Camargo de Morais MM, Melo EH, Morais MA Jr, Ledingham WM, Lima Filho JL (2001) Effect of culture conditions on lipase production by Fusarium solani in batch fermentation. Bioresour Technol 76:23–27
Martinez C, Nicolas A, van Tilbeurgh H, Egloff MP, Cudrey C, Verger R, Cambillau C (1994) Cutinase, a lipolytic enzyme with a preformed oxyanion hole. Biochemistry 33:83–89
Ogino H (2009) Organic solvent-tolerant enzymes. Seikagaku 81:1109–1118
Park IH, Kim SH, Lee YS, Lee SC, Zhou Y, Kim CM, Ahn SC, Choi YL (2009) Gene cloning, purification, and characterization of a cold-adapted lipase produced by Acinetobacter baumannii BD5. J Microbiol Biotechnol 19:128–135
Rashid N, Yuji S, Satoshi E, Haruyuki A, Tadayuki I (2001) Low-temperature lipase from psychrotrophic Pseudomonas sp. strain KB700A. Appl Environ Microbiol 67:4064–4069
Ryu HS, Kim HK, Choi WC, Kim MH, Park SY, Han NS, Oh TK, Lee JK (2001) New cold-adapted lipase from Photobacterium lipolyticum sp. nov. that is closely related to filamentous fungal lipases. Appl Microbiol Biotechnol 70:321–326
Sabate R, de Groot NS, Ventura S (2010) Protein folding and aggregation in bacteria. Cell Mol Life Sci (in press)
Snellman EA, Colwell RR (2004) Acinetobacter lipases: molecular biology, biochemical properties and biotechnological potential. J Ind Microbiol Biotechnol 31:391–400
Snellman EA, Sullivan ER, Colwell RR (2002) Purification and properties of the extracellular lipase, LipA of Acinetobacter sp. RAG-1. J Ind Microbiol Biotechnol 269:5771–5779
Suzuki T, Nakayama T, Kurihara T, Nishino T, Esaki N (2001) Cold-adapted lipolytic activity of psychrotrophic Acinetobacter sp. strain No. 6. J Biosci Bioeng 92:144–148
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Uppenberg J, Patkar S, Bergfors T, Jones TA (1994) Cold-adapted lipolytic activity of psychrotrophic Acinetobacter sp. strain No. 6. J Mol Biol 235:790–792
Yang X, Lin X, Fan T, Bian J, Huang X (2008) Cloning and expression of lipP, A Gene encoding a cold-adapted lipase from Moritella sp. 2-5-10-1. Curr Microbiol 56:194–198
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zheng, X., Chu, X., Zhang, W. et al. A novel cold-adapted lipase from Acinetobacter sp. XMZ-26: gene cloning and characterisation. Appl Microbiol Biotechnol 90, 971–980 (2011). https://doi.org/10.1007/s00253-011-3154-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-011-3154-1